Clear Sky Science · en
Molecular mapping in DNA-PAINT via modified Gaussian Mixture Modeling
Seeing the Invisible World of Molecules
Modern biology increasingly relies on microscopes that can see not just cells, but individual molecules inside them. Yet turning faint, flickering light from these molecules into a trustworthy “map” of where each one sits is surprisingly hard. This study introduces a new computational method, called G5M, that makes these molecular maps far more accurate and detailed, helping scientists understand how proteins are arranged and grouped in real cells, down to only a few billionths of a meter.
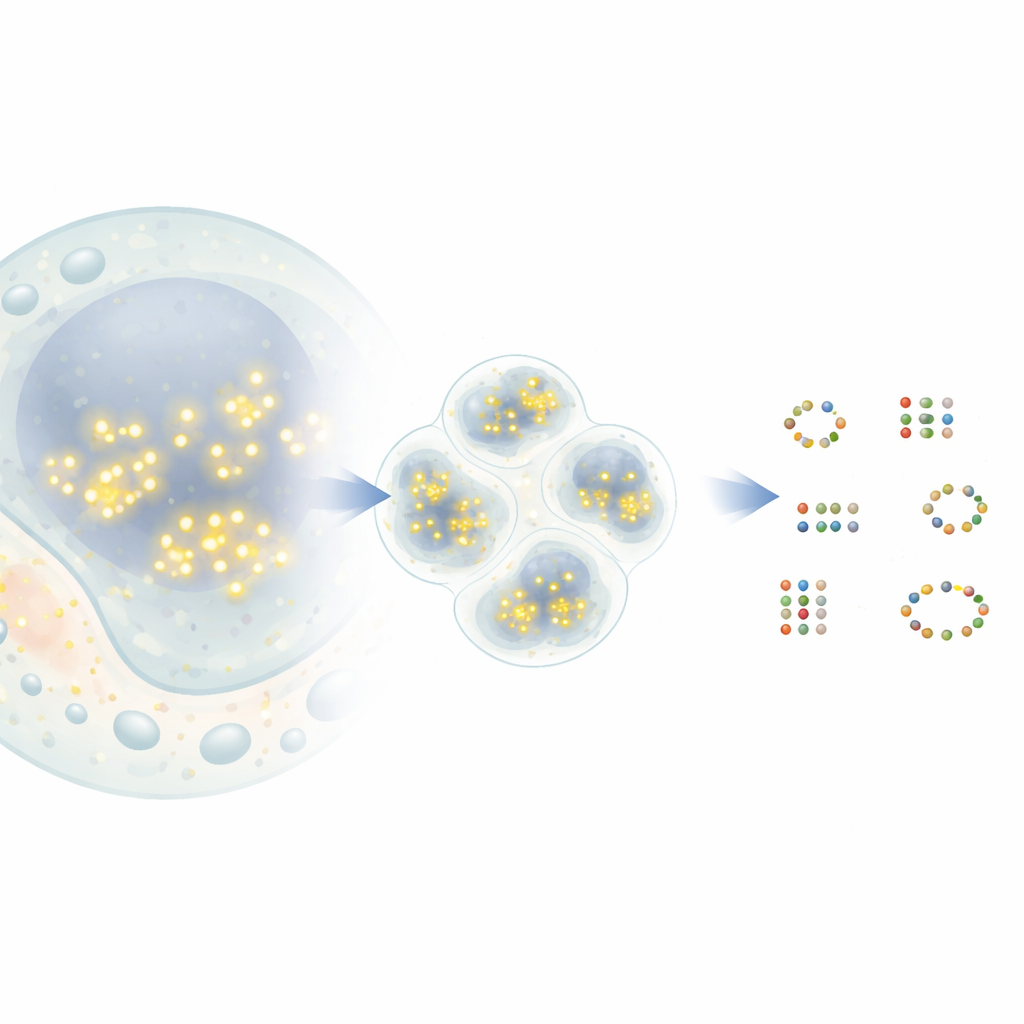
From Blinking Dots to Molecular Maps
In a popular super‑resolution technique called DNA‑PAINT, short strands of DNA carrying fluorescent dyes briefly bind and unbind to matching DNA tags attached to target proteins. Each time a dye binds, it appears as a bright dot in the microscope before vanishing again. Over time, many such events create a cloud of dots around each protein. In principle, the center of each cloud marks the true position of a protein with nanometer‑scale precision. In practice, however, dots from nearby proteins can overlap, and some dots come from random background signals. Existing analysis tools often merge close neighbors into a single protein or, conversely, invent proteins that do not exist, limiting how much biological insight can be extracted.
A Smarter Way to Find Real Molecules
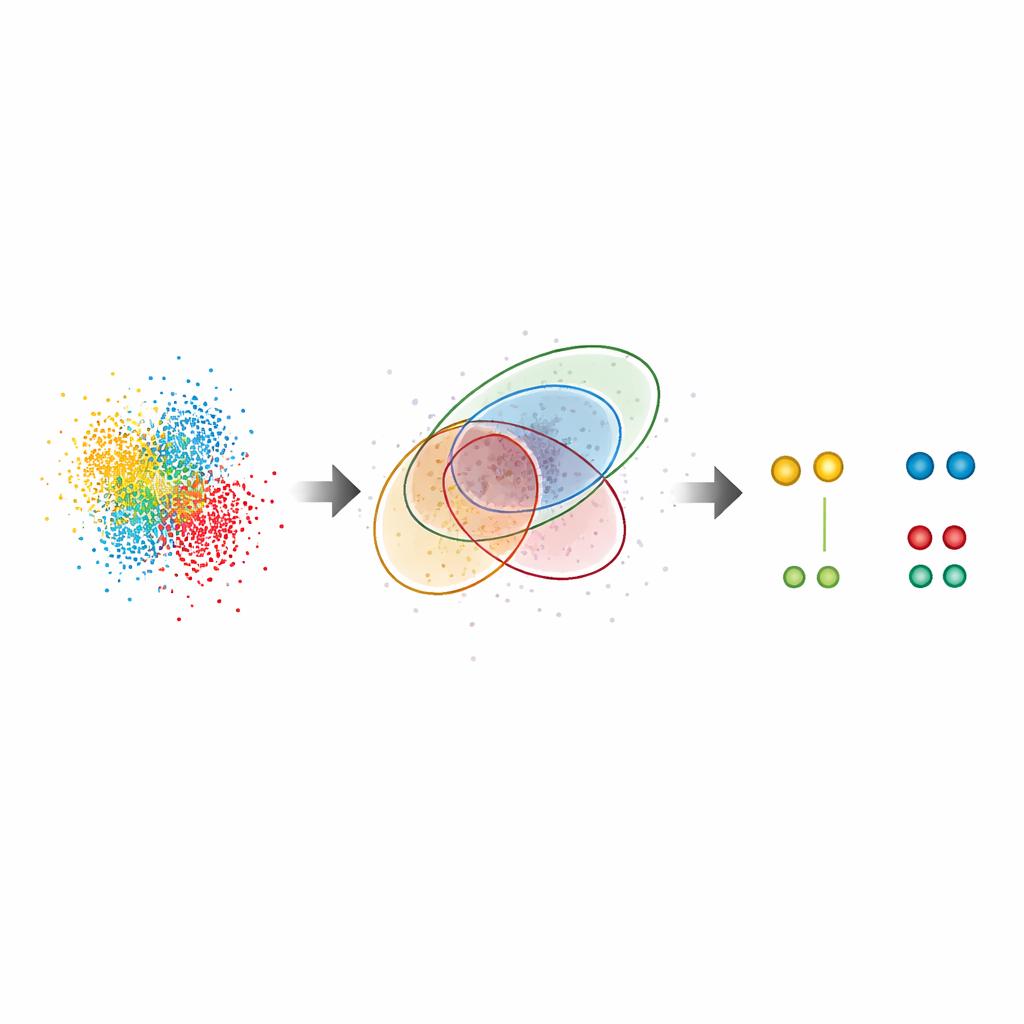
The new method, G5M, treats the swarm of dots as a mixture of simple, bell‑shaped clouds, each corresponding to one real molecule. Instead of just grouping nearby dots by density, G5M uses a probabilistic model that incorporates what is already known about the experiment: how sharply positions can be measured, how fast DNA strands bind and unbind, and how the microscope blurs light in two or three dimensions. It then tests different possible explanations—different numbers and shapes of clouds—and automatically chooses the one that best balances fit and simplicity. Additional safeguards reject suspicious solutions, such as clouds that are too narrow, too wide, based on too few dots, or not clearly separated from one another.
Proving Its Power in Simulations and DNA Nanostructures
To test G5M, the authors first used realistic computer simulations of simple scenes: pairs of molecules and small grids of twelve molecules spaced only a few nanometers apart. Compared with the current leading method, known as Gradient Ascent, G5M found many more of the molecules that should be visible at the theoretical resolution limit, while almost never reporting molecules that were not there. In key cases, it recovered closely spaced pairs twenty‑seven times more often than the older method and improved effective resolution by more than half. The team then confirmed these gains experimentally using DNA origami structures—artificial DNA shapes with docking sites placed at precisely known positions—showing that G5M could reliably count and locate nearly all expected sites over a range of imaging conditions.
Revealing Hidden Patterns in Real Cells
Beyond test samples, G5M was applied to complex biological systems. In nuclear pore complexes, giant gateways in the cell’s nucleus, the method recovered the known ring‑like arrangement of a key protein, Nup96, even where partners were separated by only about ten nanometers. It found nearly twice as many protein pairs as the standard method and reproduced independent estimates of labeling efficiency, suggesting that it neither misses many molecules nor adds spurious ones. The authors also examined CD20, a surface receptor involved in blood cancers and a target of therapeutic antibodies. Here, G5M revealed significantly more small clusters (dimers, trimers, and tetramers) of CD20 on the cell membrane, clarifying how an anti‑cancer antibody and related drug formats reorganize these receptors. It even boosted the performance of an ultra‑high‑resolution approach called RESI, which relies on separating signals across multiple imaging rounds.
What This Means for Future Microscopy
By squeezing more reliable information out of existing DNA‑PAINT data, G5M shows that better software alone can unlock new biological details, without changing microscopes or dyes. The algorithm keeps false detections extremely low while resolving molecules that are nearly touching, which is essential when asking how many proteins sit in a complex, how they are spaced, or how a drug changes their arrangement. Integrated into the open‑source Picasso platform and robust to typical settings, G5M is positioned to become a standard tool for turning blinking fluorescence into trustworthy molecular maps, helping researchers chart the nanoscale organization of life inside cells.
Citation: Kowalewski, R., Reinhardt, S.C.M., Pachmayr, I. et al. Molecular mapping in DNA-PAINT via modified Gaussian Mixture Modeling. Nat Commun 17, 2315 (2026). https://doi.org/10.1038/s41467-026-70198-5
Keywords: super-resolution microscopy, DNA-PAINT, molecular mapping, protein oligomerization, image analysis algorithms