Clear Sky Science · en
Translational regulation by oxidative desulfuration of tRNA modifications
How Cells Tune Protein Production Under Stress
Every cell must decide which proteins to make, how fast to make them, and when to slow down. This study reveals a subtle chemical switch on transfer RNA (tRNA) molecules that helps cells dial back protein production when the cell’s environment becomes more oxidizing, such as during inflammation or metabolic stress. By watching how this switch behaves in mammalian cells and in test-tube systems, the researchers uncover a new way in which damage-like changes to RNA can actually serve as a regulatory signal.
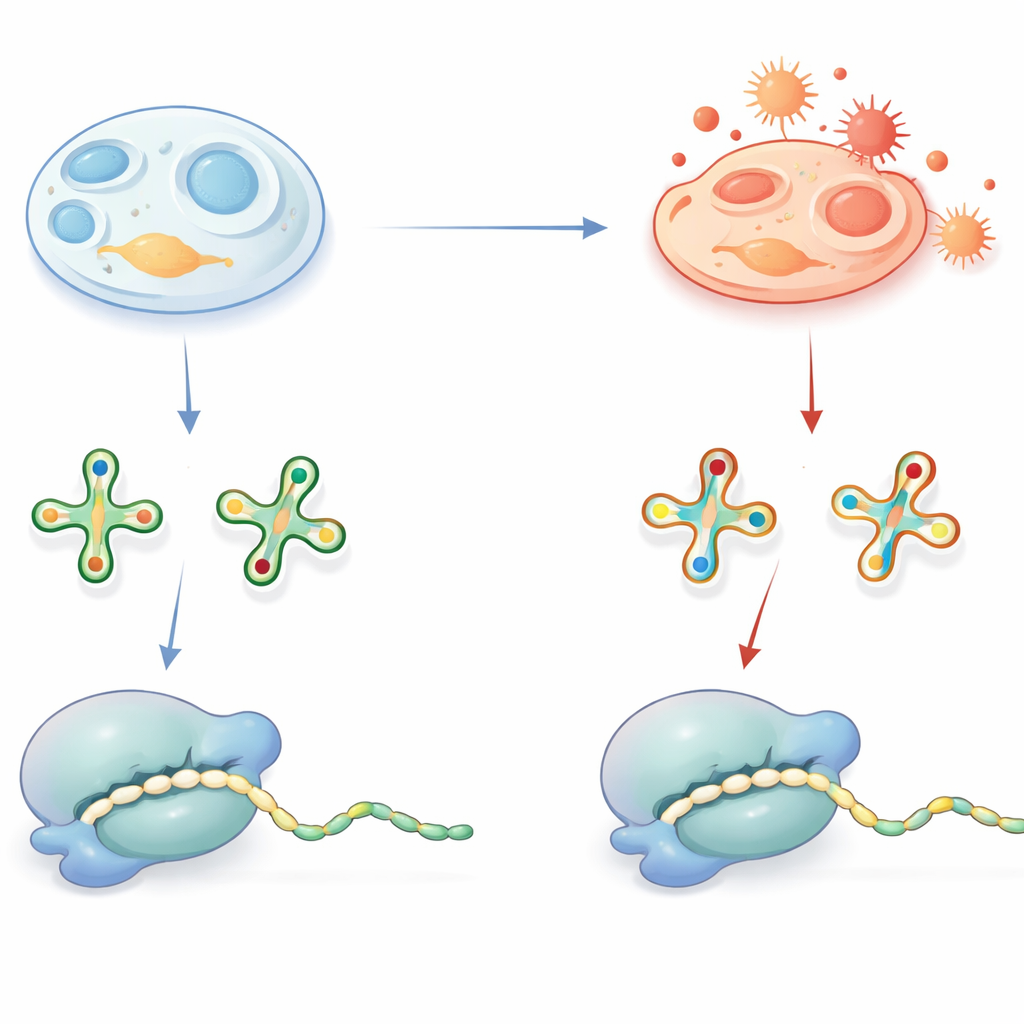
Small Marks on tRNA With Big Consequences
tRNAs act as adaptors that read three-letter “words” in messenger RNA and deliver the matching amino acids during protein synthesis. To do this job accurately and efficiently, many tRNAs carry special chemical decorations, particularly at a key position that contacts the third letter of each codon. One widespread decoration is a sulfur-containing group on a uridine base, often written as a 2-thiouridine derivative. In healthy conditions this sulfur group helps tRNA recognize the correct codons ending in the letters A or G and supports rapid, error-free translation. Defects in these decorations are already known to cause human diseases, highlighting their importance.
When Oxidative Stress Rewrites tRNA
The sulfur group that boosts decoding comes with a downside: it is chemically vulnerable to oxidation. The team asked whether, inside cells, this group is actually being stripped away under oxidative conditions, and what that would mean for protein synthesis. Using sensitive mass spectrometry, they identified oxidized, sulfur-free versions of these tRNA bases—called h2U derivatives—in mouse tissues, pig mitochondria, bacteria, yeast, and a variety of human cell lines. A clever spike-in experiment with a bacterial tRNA tracer showed that these h2U marks are not artifacts created during sample preparation but are formed inside living cells. Although the oxidized versions were present only in a fraction of tRNAs, their abundance varied across tissues and cell types, suggesting that cellular redox state and antioxidant capacity influence how often this conversion occurs.
How Oxidized tRNAs Slow the Assembly Line
The researchers next tested how these altered tRNAs perform during protein synthesis. They chemically converted the sulfur-containing form to the oxidized h2U form in bulk tRNA and used a fully reconstituted human translation system in vitro. A reporter protein whose production strongly depends on these modified tRNAs showed sharply reduced output when the tRNAs were desulfurized, while a control reporter that avoids the affected codons translated just fine. Biochemical assays revealed why: oxidized tRNAs for lysine, glutamine, and glutamate were charged with their amino acids much less efficiently, whereas the corresponding arginine tRNAs were largely spared. Importantly, h2U-bearing tRNAs still appeared in ribosome-bound (polysomal) fractions from cells, indicating that they do participate in translation but do so poorly.
A Structural View of Weakened Decoding
To see exactly how oxidation changes decoding, the team used high-resolution cryogenic electron microscopy to image bacterial ribosomes programmed with human lysine tRNA and specific codons. In the normal, sulfur-containing state, the modified base at the tRNA “wobble” position formed either a standard base pair with an A-ending codon or a special wobble pair with a G-ending codon, both firmly nestled in the ribosome’s decoding center. After desulfuration, the same base could make only a single weak contact with the A-ending codon and formed a less stable wobble pair with the G-ending codon. Binding experiments matched these structures: oxidized lysine tRNA essentially lost the ability to recognize the AAA codon and recognized the AAG codon much less efficiently. Together, these findings show that a seemingly small chemical tweak can translate into codon-specific slowdowns in protein synthesis.
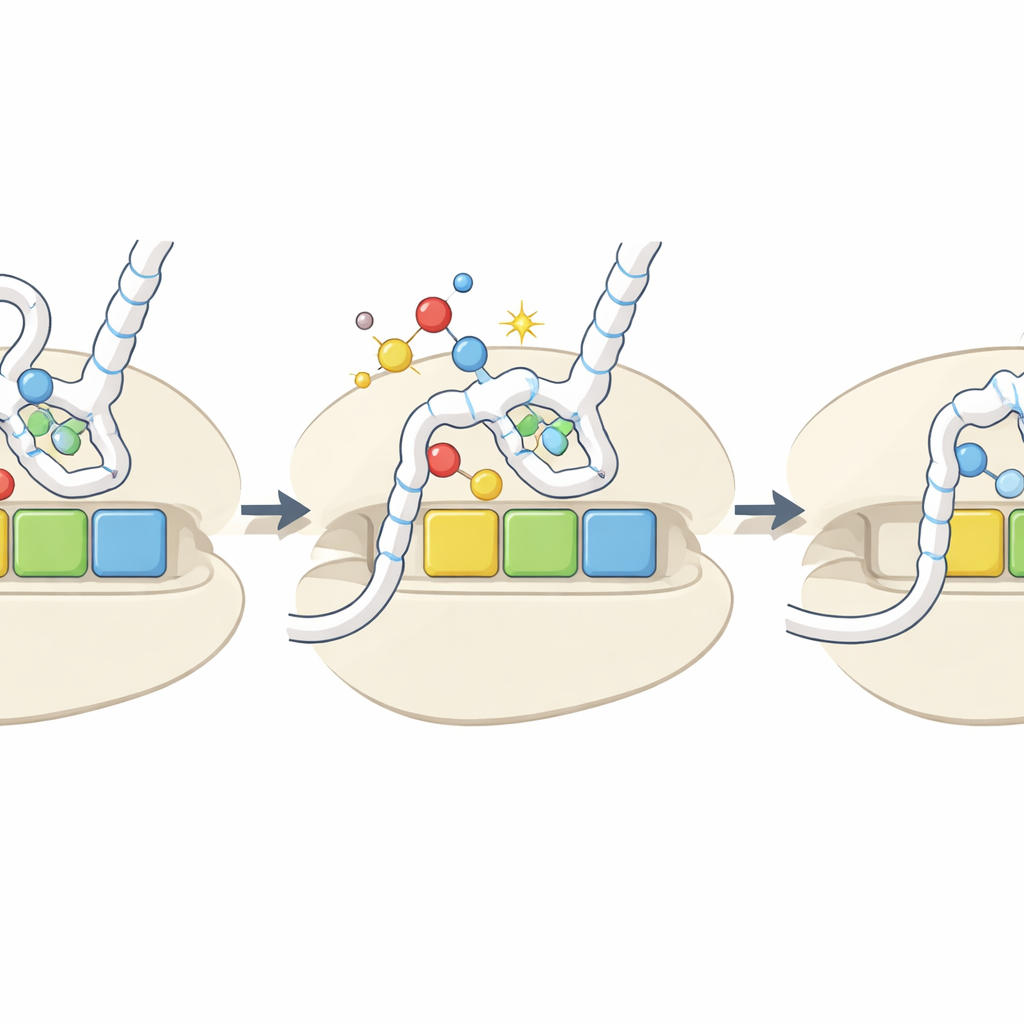
A Damage Signal That Becomes a Regulatory Knob
Overall, the study proposes that oxidative removal of sulfur from key tRNA bases acts as a built-in sensor for the cell’s redox environment. Under oxidative stress, more tRNAs are converted to the h2U form, which are poorly charged and bind weakly to certain codons. This selectively slows translation at those codons, potentially altering the stability of affected mRNAs, changing how proteins fold, and triggering stress-response pathways when ribosomes stall. Although the oxidized marks are relatively rare and their levels do not always spike under experimental stress, their codon- and amino-acid–specific effects mean that even modest changes could reshape which proteins are made, and how fast, during challenging conditions.
Citation: Mo, Y., Ishiguro, K., Miyauchi, K. et al. Translational regulation by oxidative desulfuration of tRNA modifications. Nat Commun 17, 2125 (2026). https://doi.org/10.1038/s41467-026-70126-7
Keywords: tRNA modification, oxidative stress, protein synthesis, translation regulation, RNA damage