Clear Sky Science · en
Subcellular proteomics reveals a blueprint for endosymbiont integration in trypanosomatid Angomonas deanei
Hidden partners inside single-celled life
Many single-celled organisms are not truly alone. Some harbor bacteria that live inside them as permanent tenants, trading food and services in relationships that echo the origins of our own mitochondria. This study explores such a partnership in Angomonas deanei, a microscopic parasite of insects, to reveal how a once free-living bacterium has become deeply woven into the host cell’s inner workings.
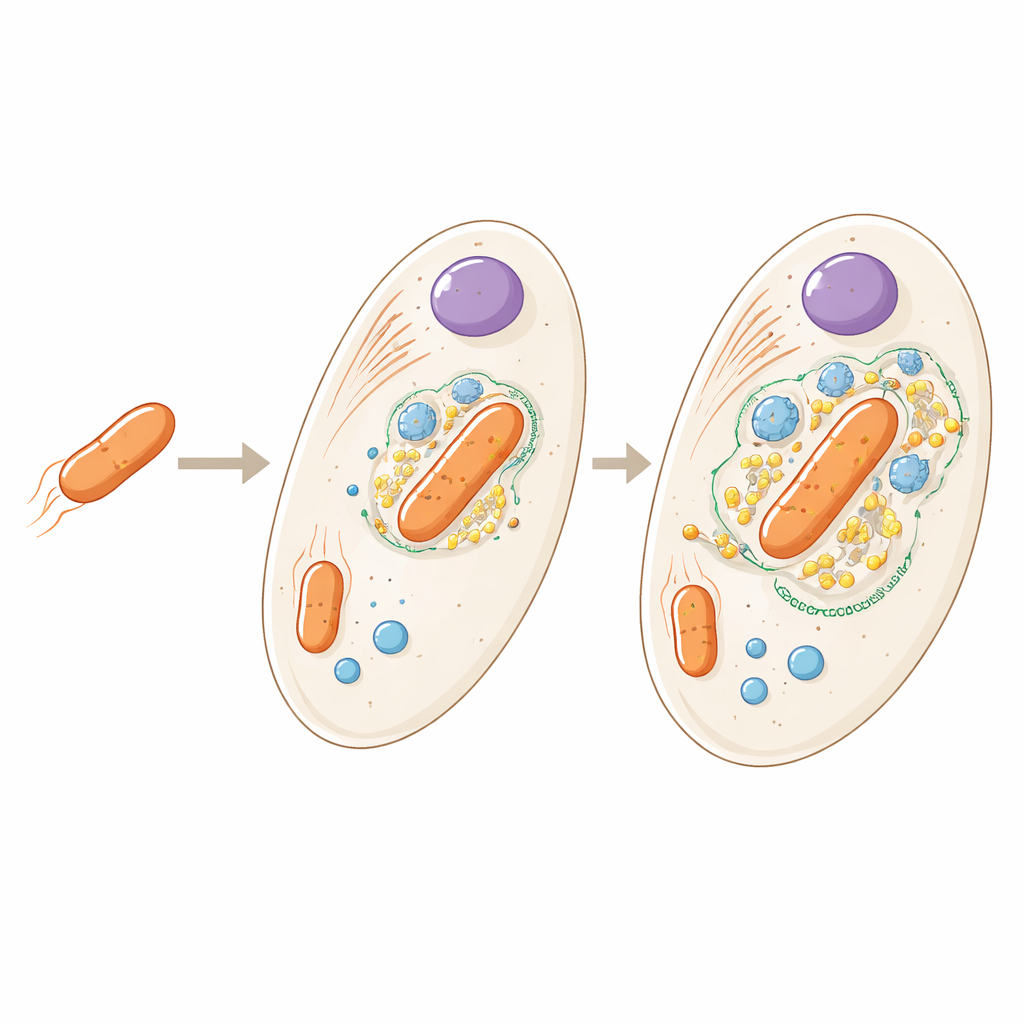
From houseguest to built-in part
The bacterium living inside Angomonas deanei is a distant relative of free-living soil microbes, but here it exists as a single copy per host cell and can no longer survive on its own. Earlier work showed that the host produces special proteins that travel into the bacterium and even help control when it divides. The new study asks a broader question: if we look across the entire cell, where exactly are all the host and bacterial proteins located, and what does that tell us about how tightly the two partners are integrated?
Mapping the cell’s inner city
To answer this, the researchers broke open large numbers of cells very gently so that internal structures stayed mostly intact. They then separated these pieces by spinning them at different speeds, and identified thousands of proteins in each fraction using high-resolution mass spectrometry. By comparing how proteins clustered across the fractions, and cross-checking with known markers and fluorescent microscopy, they created a detailed “address book” for the cell, assigning nearly 3,000 proteins to 21 distinct regions such as the nucleus, mitochondrion, specialized storage bodies, and the bacterial endosymbiont itself.
Host tools targeting the live-in bacterium
Within this atlas, the team uncovered a set of host-made proteins that consistently traveled with the endosymbiont, expanding the known collection of so‑called endosymbiont-targeted proteins. One newly confirmed example, dubbed ETP10, coats the bacterium much like an earlier protein, suggesting a scaffold of host factors wrapped around the microbial partner. Another small group of proteins shared an unusual physical behavior with a dynamin-like protein already known to pinch the bacterium during division. Together, these findings hint at a specialized, host-built machinery dedicated to positioning, shaping, and reproducing the endosymbiont inside the cell.
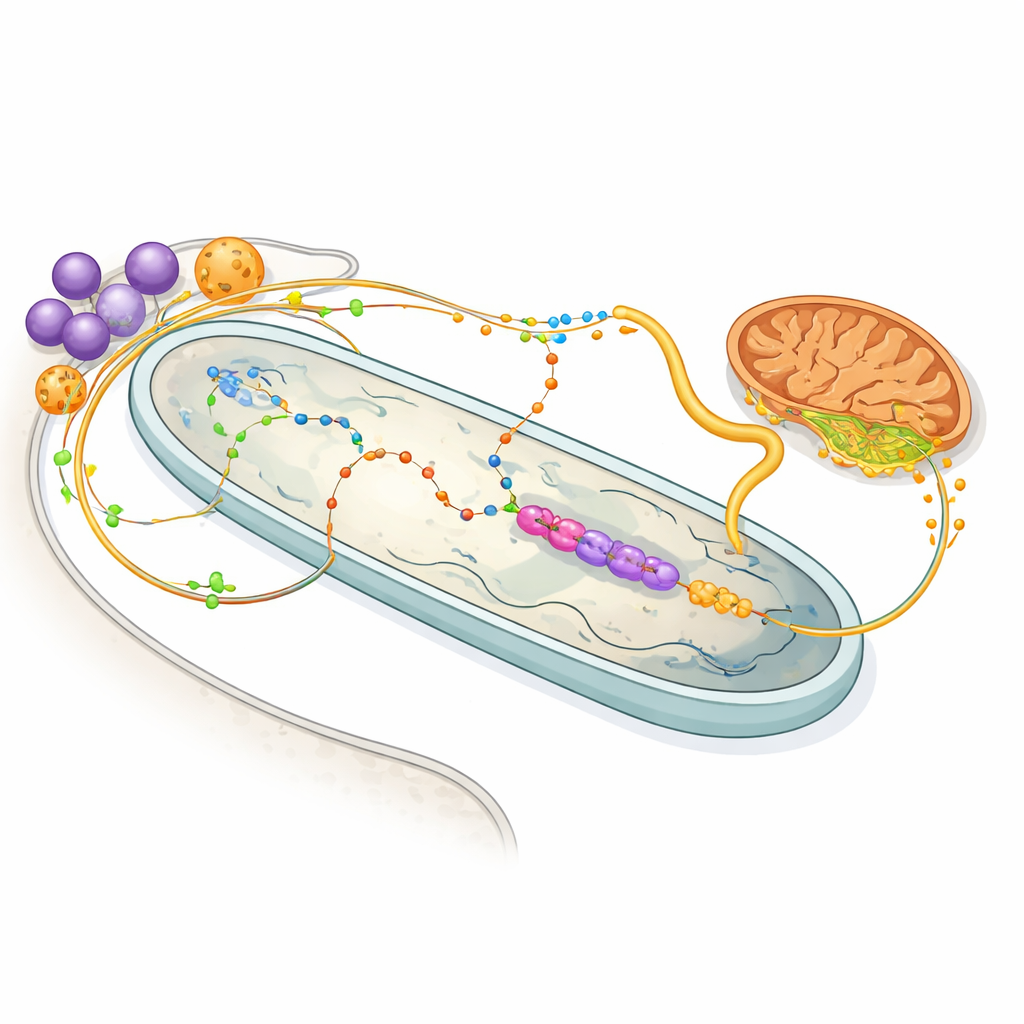
Energy sharing and chemical trade routes
The protein map also illuminates the bustling trade in molecules between host and bacterium. Enzymes in the host’s glycosomes—organelles that handle sugar and amino-acid metabolism—appear tuned to feed the bacterium with the amino acid proline and the compound 2‑oxoglutarate, both valuable energy sources. Inside the bacterium, a stripped-down set of enzymes uses 2‑oxoglutarate to generate NADH, which drives a minimal respiratory chain, then passes on succinate back to the host’s mitochondrion. Other pathways show that the bacterium performs most of the steps in making heme, a vital pigment needed by the host, while also contributing crucial building blocks for nucleotides. In effect, each partner has offloaded specific metabolic chores to the other, creating a shared, interdependent system.
New lines of communication inside the cell
Beyond metabolism, the study reveals physical and likely signaling links between the endosymbiont and other organelles. A distinct group of host proteins forms what the authors call a “contact site” cluster, enriched in nuclear envelope and endoplasmic reticulum components that physically co-sediment with the bacterium. Two types of storage organelles—glycosomes and acidocalcisomes—appear in two populations: some remain free in the cytoplasm, while others consistently travel with the bacterium. Microscopy confirms that subsets of these organelles nestle tightly against the endosymbiont. Acidocalcisomes are known calcium and phosphate reservoirs, so their close apposition suggests a two-way signaling and ion-traffic route connecting the bacterium, these stores, and the nearby endoplasmic reticulum.
A blueprint for life merged within life
Taken together, this work provides a rich, cell-wide blueprint of how a bacterium can become functionally fused with its host without yet turning into a full-fledged organelle like mitochondria. Angomonas deanei relies on its endosymbiont for key nutrients, while the bacterium, in turn, depends on host-made proteins and energy-rich molecules and is tethered to host organelles for communication and control. For non-specialists, the takeaway is that evolution can gradually hard-wire cooperation between very different forms of life, step by step, until separating them becomes impossible. This system offers a live snapshot of that process in action, helping scientists better understand how complex cells like our own once emerged from simpler beginnings.
Citation: Hammond, M., Chmelová, Ľ., van Geelen-Kuenzel, N.A. et al. Subcellular proteomics reveals a blueprint for endosymbiont integration in trypanosomatid Angomonas deanei. Nat Commun 17, 2241 (2026). https://doi.org/10.1038/s41467-026-70084-0
Keywords: endosymbiosis, cell metabolism, organelle evolution, protist biology, host microbe interactions