Clear Sky Science · en
RPA directly stimulates Mer3 helicase processivity to ensure normal crossover formation in meiosis
How Cells Shuffle DNA to Make Healthy Eggs and Sperm
Every time a plant, animal, or human makes eggs or sperm, its cells must reshuffle chromosomes so that each offspring gets a fresh mix of parental DNA. This genetic reshuffling depends on carefully placed DNA exchanges, or crossovers, between chromosome pairs. The featured study uncovers how two key proteins, Mer3 and RPA, work together as a microscopic machine to make sure these crossovers form efficiently and safely—a process with implications for fertility, evolution, and possibly even cancer.
A Delicate Balancing Act in Sexual Reproduction
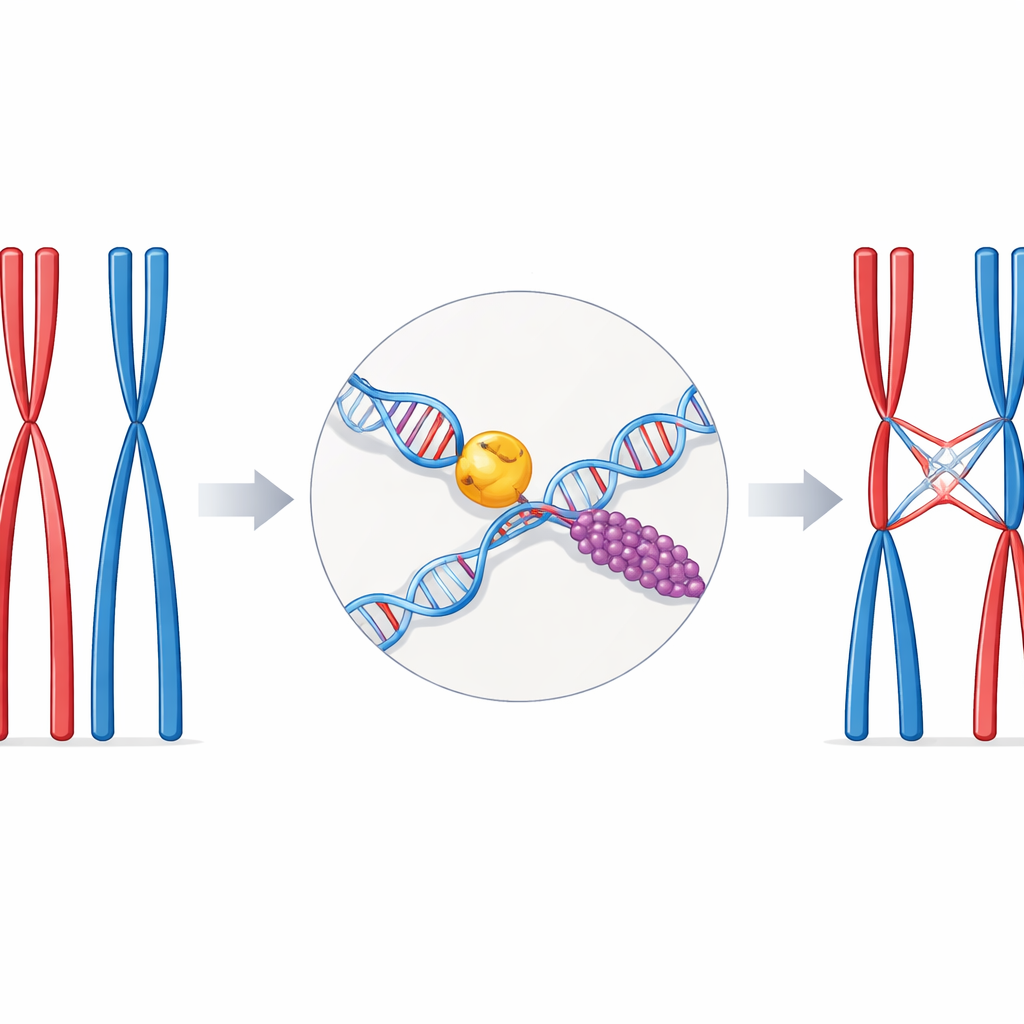
During the special cell division called meiosis, chromosomes come in matching pairs, one from each parent. For these pairs to separate correctly, they must be physically linked at a few well‑chosen spots along their length. These links are the crossovers—sections where the DNA strands from one chromosome are swapped with those of its partner. Having too few crossovers risks chromosome mis‑segregation and infertility; having too many or badly placed ones can damage the genome. Cells therefore rely on specialized repair pathways and helper proteins to turn a dangerous DNA break into a properly controlled crossover.
Meet Mer3 and RPA, a DNA‑Unwinding Team
The researchers focused on Mer3, a molecular motor that unwinds DNA and promotes crossover formation. Its human equivalent, HFM1, is known to be important for fertility. They discovered that Mer3 directly binds to RPA, a protein complex that normally coats exposed single‑stranded DNA to protect it and recruit other repair factors. Using biochemical tests, structural modeling, and interaction assays, the team mapped a specific docking site where a short tail of Mer3 fits into a groove on one RPA subunit. This interface is conserved from yeast to mammals, suggesting it is a long‑standing solution that evolution has reused to control DNA repair during meiosis.
Watching a Single DNA Motor at Work
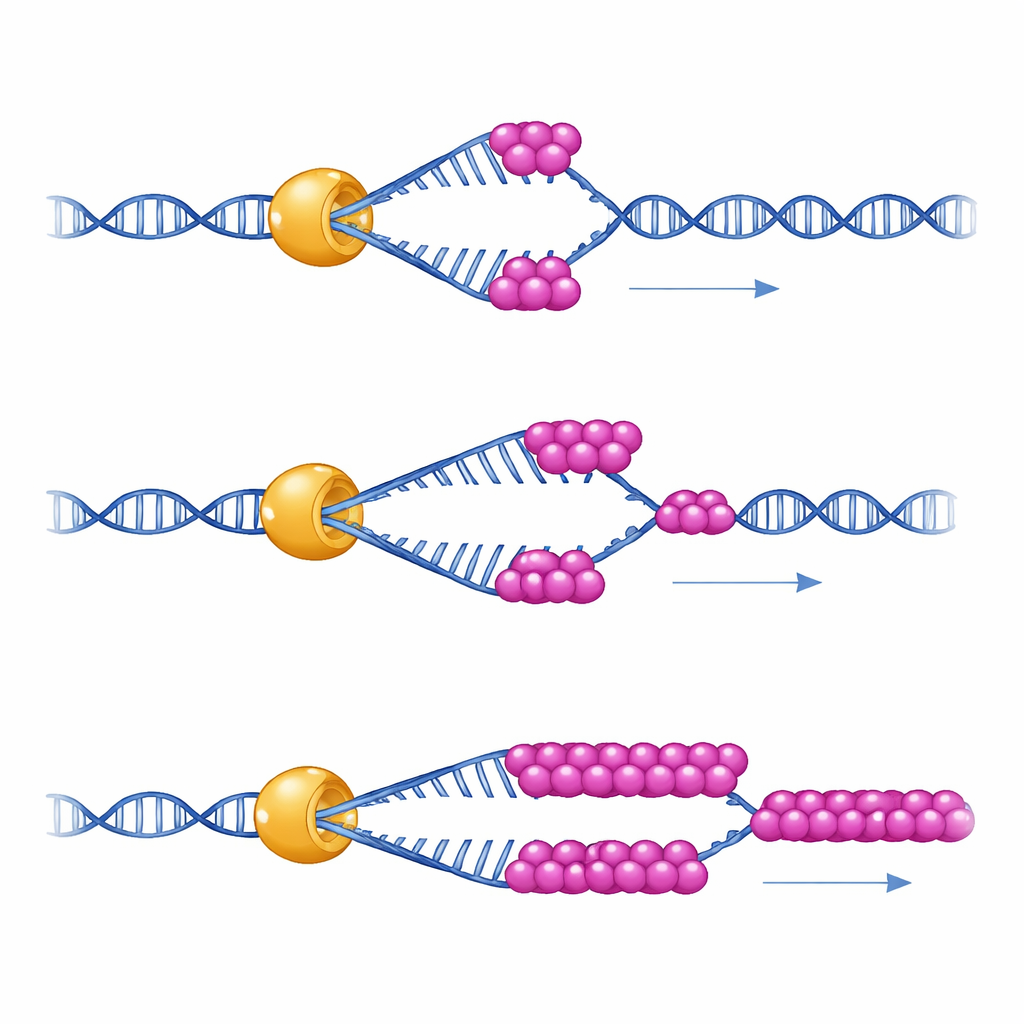
To see what this partnership actually does, the scientists used single‑molecule magnetic tweezers—an instrument that holds a single DNA hairpin like a tiny spring and measures how a single Mer3 molecule unwinds it over time. They found that Mer3 moves along DNA at a steady speed even on its own. But under conditions that mimic the relatively low physical tension inside cells, Mer3 tends to let go of the DNA after unwinding only a short stretch. When a small amount of RPA is present, Mer3 suddenly becomes far more persistent: it can unwind much longer stretches of DNA without falling off, especially when the DNA is prone to folding back on itself. A mutant version of Mer3 engineered to disrupt the RPA‑binding tail fails to gain this extra staying power, revealing that direct contact with RPA is what boosts Mer3’s processivity.
Consequences for Crossovers in Living Cells
The team then asked what happens in actual yeast cells when Mer3 cannot grip RPA properly. They replaced the normal MER3 gene with the RPA‑binding mutant and monitored meiosis. These cells could still divide and make spores, but their fertility dropped and the overall number of crossovers along certain chromosome regions decreased. Detailed DNA analysis showed more repair events ending as non‑crossovers and an accumulation of unresolved recombination intermediates. Genome‑wide mapping of Mer3’s location revealed that the mutant protein was less stably recruited to sites where programmed DNA breaks occur, even though its early association with chromosome axes looked normal. In other words, without strong RPA binding, Mer3 does not linger long enough at break sites to reliably drive them toward crossover outcomes.
What This Means for Fertility and Genome Stability
Taken together, the study shows that RPA does more than simply coat loose DNA; it directly tunes the behavior of a key meiotic helicase. By docking onto RPA, Mer3 becomes a more determined DNA unwinder, able to extend and stabilize the joint DNA structures that mature into crossovers. When this interaction is weakened, cells shift toward safer but less useful repair outcomes and leave more intermediates unresolved, subtly undermining chromosome segregation. Because the same molecular players exist in humans—and mutations in HFM1 are linked to infertility—this work provides a mechanistic framework for how tiny changes at a protein interface can ripple outward to affect reproductive health and the genetic diversity of future generations.
Citation: Altmannova, V., Orlić, L., Carrasco, C. et al. RPA directly stimulates Mer3 helicase processivity to ensure normal crossover formation in meiosis. Nat Commun 17, 2621 (2026). https://doi.org/10.1038/s41467-026-69985-x
Keywords: meiosis, genetic recombination, DNA helicase, fertility, chromosome crossover