Clear Sky Science · en
Cross-species gene redesign leveraging ortholog information and generative modeling
Why redesigning genes across species matters
Modern biotechnology often needs to move genes from one microbe into another to make medicines, enzymes, or environmental clean-up tools. Yet a gene that works well in its original microbe can sputter in a new host, producing little protein. This article introduces a new artificial intelligence system, OrthologTransformer, that learns from evolution itself to rewrite genes so they feel “native” in a different species, boosting their performance and opening new possibilities for green technology and industry.
Limits of today’s gene tuning tricks
For decades, scientists have relied on a strategy called codon optimization to help foreign genes work in new hosts. The idea is simple: the genetic code has several three-letter “codons” that can spell the same amino acid, and different species prefer different codons. Traditional tools swap rare codons for preferred ones without changing the protein’s amino-acid sequence. This often helps, but it ignores many other features that matter for gene performance, such as RNA folding, regulatory signals, and the timing of protein production. In some cases, over-optimizing codons can even hurt protein yield. Meanwhile, nature solves cross-species adaptation in a richer way: related genes in different species, known as orthologs, commonly show amino-acid changes and small insertions or deletions as well as codon swaps, all while preserving overall function.
Learning nature’s playbook for gene rewriting
OrthologTransformer treats gene redesign as a kind of language translation: given a DNA sequence from one bacterium, it “translates” it into how that gene would likely look in another species. The model is built on the Transformer architecture used in modern language tools, but here it operates on codons instead of words. It is trained on millions of naturally paired orthologous genes from over two thousand bacterial species, with special tokens that tell it which species it is converting from and to. By seeing how evolution has already balanced function and host adaptation, the system learns when simple codon swaps are enough and when subtle amino-acid changes or length adjustments are tolerated. In tests spanning 45 bacterial species and hundreds of source–target combinations, the AI’s redesigned genes more closely resembled the target species’ native orthologs than both conventional codon optimization and a leading neural codon optimizer, while still keeping high similarity at the protein level. 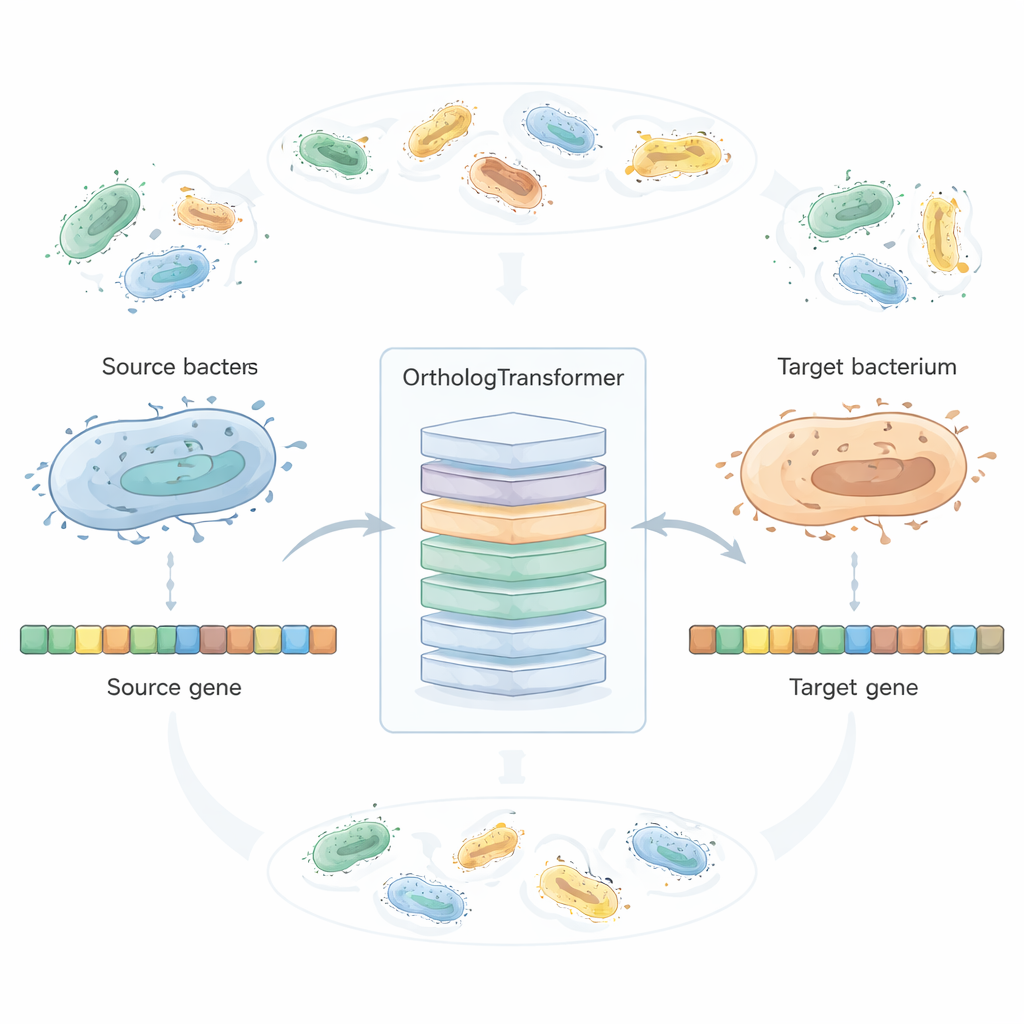
Putting AI-designed plastic-eating enzymes to work
To show that this is more than a computational trick, the team focused on PETase, an enzyme from the bacterium Ideonella sakaiensis that can break down PET plastic, the material used in most drink bottles. Ideonella grows slowly and is not ideal for industrial use, so the researchers asked OrthologTransformer to rewrite the PETase gene for a faster-growing host, Bacillus subtilis. They generated a panel of twelve redesigned gene variants, exploring different training settings and an additional search procedure that nudged sequences toward Bacillus-like DNA composition and favorable RNA structures. Despite some variants carrying many DNA changes and a few amino-acid substitutions, computer models predicted that the core 3D shape of the enzyme was preserved. When these designs were built and tested in living Bacillus cells, several produced high amounts of secreted PETase, and all showed measurable plastic-degrading activity.
An AI design that outperforms standard optimization
One AI-designed sequence, called AI-L2, stood out. Bacillus cells carrying this gene secreted especially large amounts of PETase and generated roughly three times more plastic breakdown product than any other strain in a seven-day test, and about ten times more than typical codon-optimized controls when measured by reaction products. Microscopy images of PET films exposed to AI-L2 cells revealed deep pits and holes where the plastic had been eaten away, far more dramatic than in other conditions. Detailed enzyme tests showed that the AI-L2 version of PETase not only was produced more efficiently but also processed its substrate faster, giving it higher catalytic efficiency than the original and codon-optimized enzymes. A parallel experiment in Escherichia coli showed that an OrthologTransformer-designed version of PETase, even when it changed only codon usage and not the amino-acid sequence, still outperformed a frequency-based codon-optimized gene, highlighting that the model captures subtle, host-specific preferences that traditional methods miss. 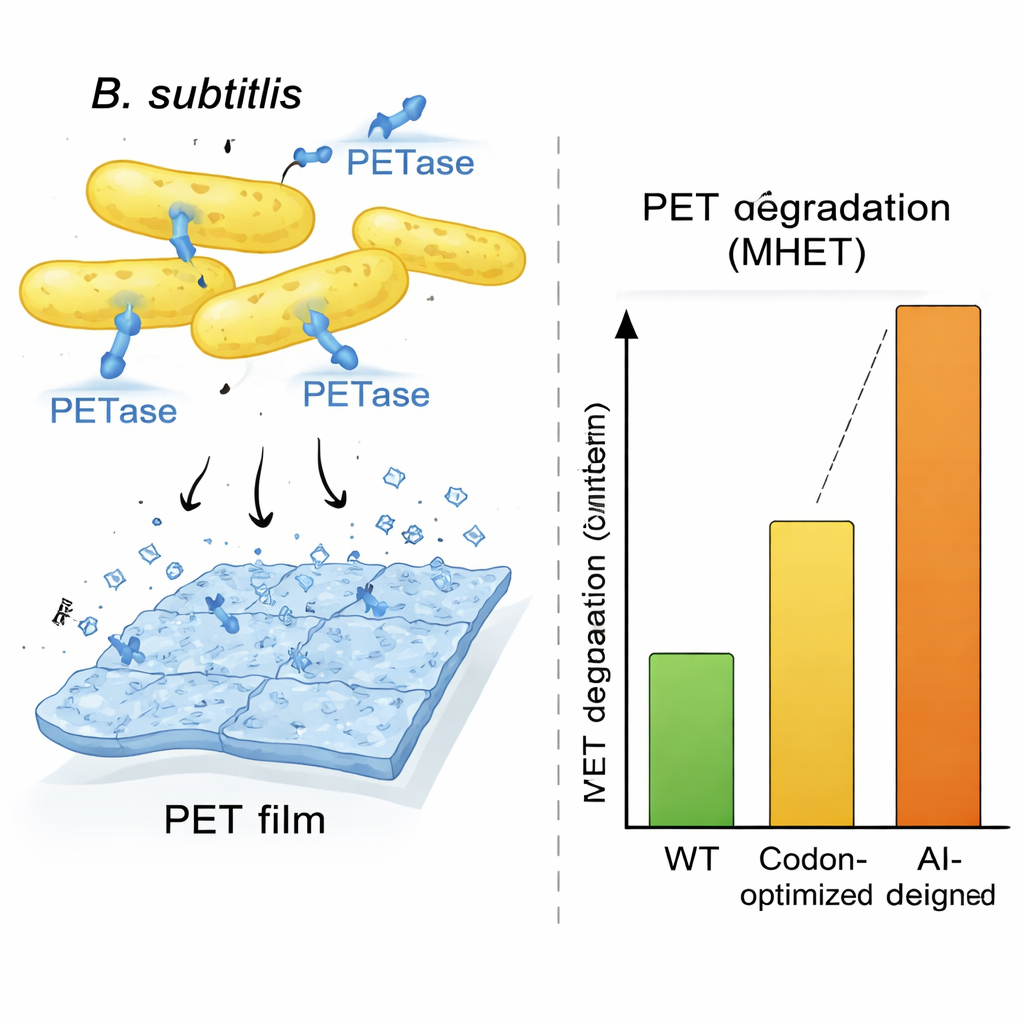
What this means for future biology and technology
In everyday terms, OrthologTransformer is like an expert translator that not only rewrites the “spelling” of a gene for a new microbe but also makes small, evolution-informed edits to the “sentence” itself when it knows they will be safe or beneficial. By learning directly from how genes have naturally adapted across thousands of bacterial species, it can propose redesigned DNA that works better in new hosts than designs limited to codon swapping. The successful creation of a more powerful, plastic-eating enzyme in Bacillus subtilis suggests that such AI-guided gene redesign could speed up the development of industrial biocatalysts, environmental clean-up microbes, and eventually even medical gene therapies, helping organisms read and use foreign genes as if they were their own.
Citation: Akiyama, M., Tashiro, M., Huang, Y. et al. Cross-species gene redesign leveraging ortholog information and generative modeling. Nat Commun 17, 2120 (2026). https://doi.org/10.1038/s41467-026-69966-0
Keywords: gene redesign, synthetic biology, orthologous genes, AI in biotechnology, plastic-degrading enzymes