Clear Sky Science · en
Nonlinear genomic selection index accelerates multi-trait crop improvement
Smarter breeding for a hungrier world
As the global population grows and climates become less predictable, plant breeders must improve several crop traits at once—like yield, height, and flowering time—faster than ever before. This article presents a new mathematical tool that helps breeders do exactly that by using DNA information in a more realistic way, capturing not only the individual effects of genes but also how they interact. The approach promises to speed up the creation of better maize and wheat varieties without needing to measure every plant in the field.
Why combining many traits is so hard
Breeders rarely care about just one trait. For example, they want higher grain yield but also shorter, sturdier plants that flower at the right time. Classic “selection indices” turn several traits into a single score to rank plants. Traditionally, these indices assume that each trait contributes in a simple, straight-line way and that the effects of different traits just add up. Real biology is messier: traits influence one another, and there can be sweet spots where “more” is no longer better. Ignoring these nonlinear interactions can slow genetic progress and even push breeding in the wrong direction.
From simple lines to flexible curves
Earlier genomic tools allowed breeders to use DNA markers spread across the genome to predict how good a plant’s offspring would be, leading to so-called linear genomic selection indices. These work well when gene effects are mostly additive. The authors extend a more flexible, older idea—the quadratic phenotypic selection index, which already allowed squared terms and trait-by-trait interactions—into the DNA era. Their new tool, called the Quadratic Genomic Selection Index (QGSI), uses genomic breeding value predictions and combines them through both linear and curved (quadratic) terms. This lets the index capture complex patterns such as gene–gene interactions and optimal trait combinations, even when field measurements are not available for every cycle.
Putting the new index to the test
To see whether this added complexity pays off, the researchers compared QGSI with both linear and quadratic indices that use only field data, and with linear genomic indices that use DNA but stay simple. They ran computer simulations of maize breeding over 10 selection cycles and also analyzed two real maize and five wheat datasets from international breeding programs. Two ways of predicting genetic value from DNA were tested: a standard additive model and a more flexible Gaussian kernel model that can capture subtle gene interactions. Across these settings, QGSI consistently produced larger selection responses—that is, bigger overall improvement across traits—than the linear indices, and typically beat the quadratic phenotypic index as well.
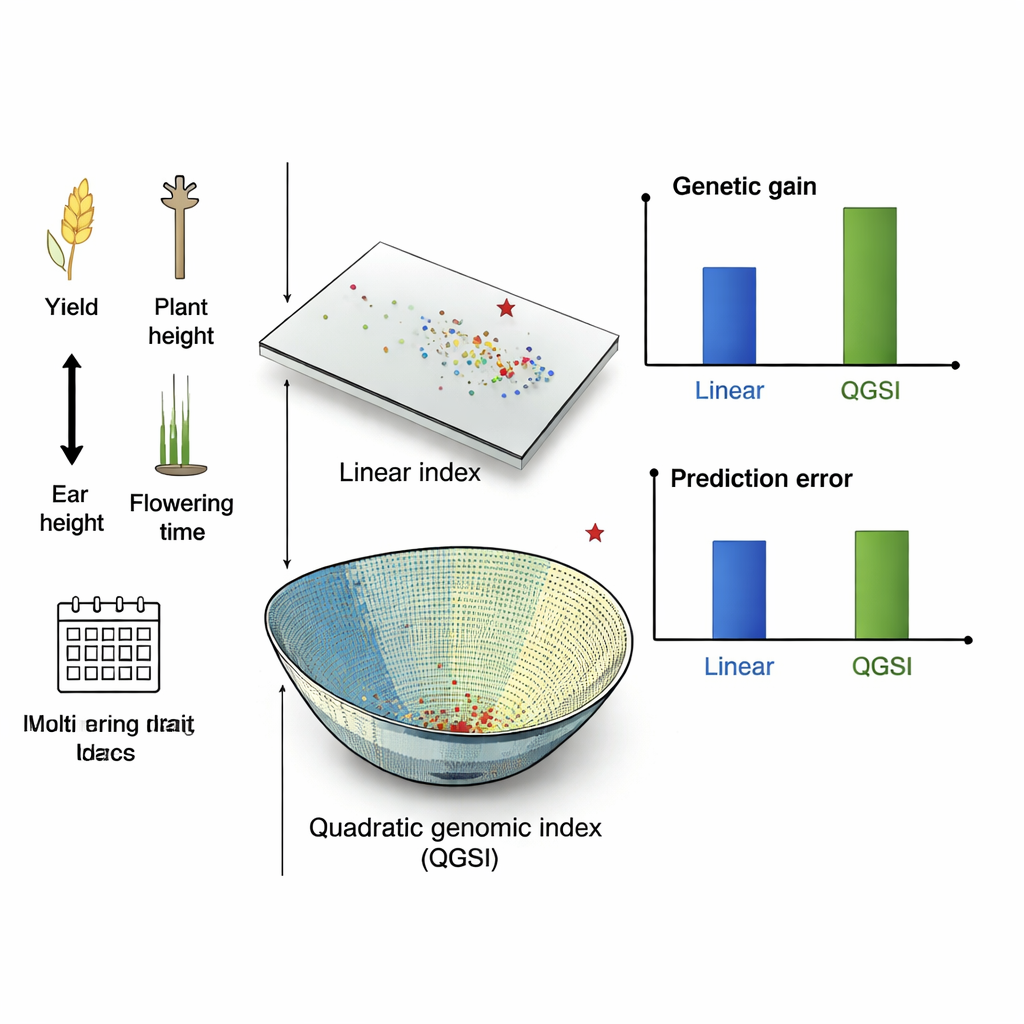
Better gains, fewer errors, more balance
In the simulated maize cycles, QGSI delivered the highest gains, outpacing both linear genomic indices and quadratic indices based only on field measurements. It also tended to have lower prediction error, meaning its scores were more reliable guides for choosing parents. In real maize populations from Mexico and Zimbabwe, QGSI achieved 80–90% higher gains than linear genomic indices when several traits were improved together. In wheat trials run under different irrigation and rainfall conditions, the pattern was similar: quadratic indices outperformed linear ones, and combining QGSI with the Gaussian kernel model gave the strongest and most stable improvements across environments, especially for grain yield while keeping plant height and flowering time in acceptable ranges.
What this means for future crops
For non-specialists, the key message is that breeders now have a more realistic scoring system that reflects how genes and traits truly interact, rather than forcing them into a straight-line model. The authors recommend using the quadratic phenotypic index when only field data are available in early stages, and switching to the QGSI once genomic data and rapid selection cycles are in place. By better capturing nonlinear genetic relationships, QGSI can accelerate multi-trait crop improvement and help deliver new maize and wheat varieties that are higher yielding, more resilient, and better adapted to challenging environments.
Citation: Jesús Cerón-Rojas, J., Montesinos-López, O.A., Montesinos-López, A. et al. Nonlinear genomic selection index accelerates multi-trait crop improvement. Nat Commun 17, 1991 (2026). https://doi.org/10.1038/s41467-026-69890-3
Keywords: genomic selection, crop breeding, maize, wheat, multi-trait improvement