Clear Sky Science · en
ERCC6L2 ensures repair fidelity for staggered-end DNA double-strand breaks
Why this discovery matters for our DNA
Every cell in your body constantly suffers damage to its DNA, especially the most dangerous kind: breaks that cut both strands of the double helix. These breaks can lead to cancer, inherited disorders, or failures of medical genome editing if they are not repaired correctly. This study reveals a previously underappreciated "guardian" protein, ERCC6L2, that specifically protects one tricky type of break from turning into large gaps and scrambled chromosomes. Understanding this guardian helps explain a rare human bone marrow disease and warns how certain gene-editing tools might backfire in vulnerable patients.
Different kinds of DNA breaks, different risks
Not all DNA breaks are created equal. Some cuts are clean, with both strands snapped at the same point, like slicing a rope straight across. Others are stepped, leaving short overhanging ends that do not line up perfectly. Modern genome editors such as Cas9 usually create clean, blunt cuts, while tools like Cas12a, TALENs, and some paired nickase systems create staggered, overhanging breaks. The researchers systematically compared how human cells cope with these two break types by knocking down thousands of genes and then asking which ones were especially important for repairing each kind of cut.
A hidden guardian for staggered breaks
From these genome-wide screens, ERCC6L2 emerged as a crucial protector for staggered breaks but largely dispensable for blunt ones. Cells lacking ERCC6L2 could still repair blunt Cas9 cuts with only small insertions or deletions, the typical fingerprints of routine DNA repair. In stark contrast, when staggered breaks were made by Cas12a, TALENs, or dual-nickase Cas9, ERCC6L2-deficient cells produced many more large deletions reaching thousands of DNA letters and widely spaced chromosomal rearrangements called translocations. These effects were seen not only in engineered cell lines but also in bone marrow cells from patients carrying inherited ERCC6L2 mutations, underscoring the clinical relevance of the finding. 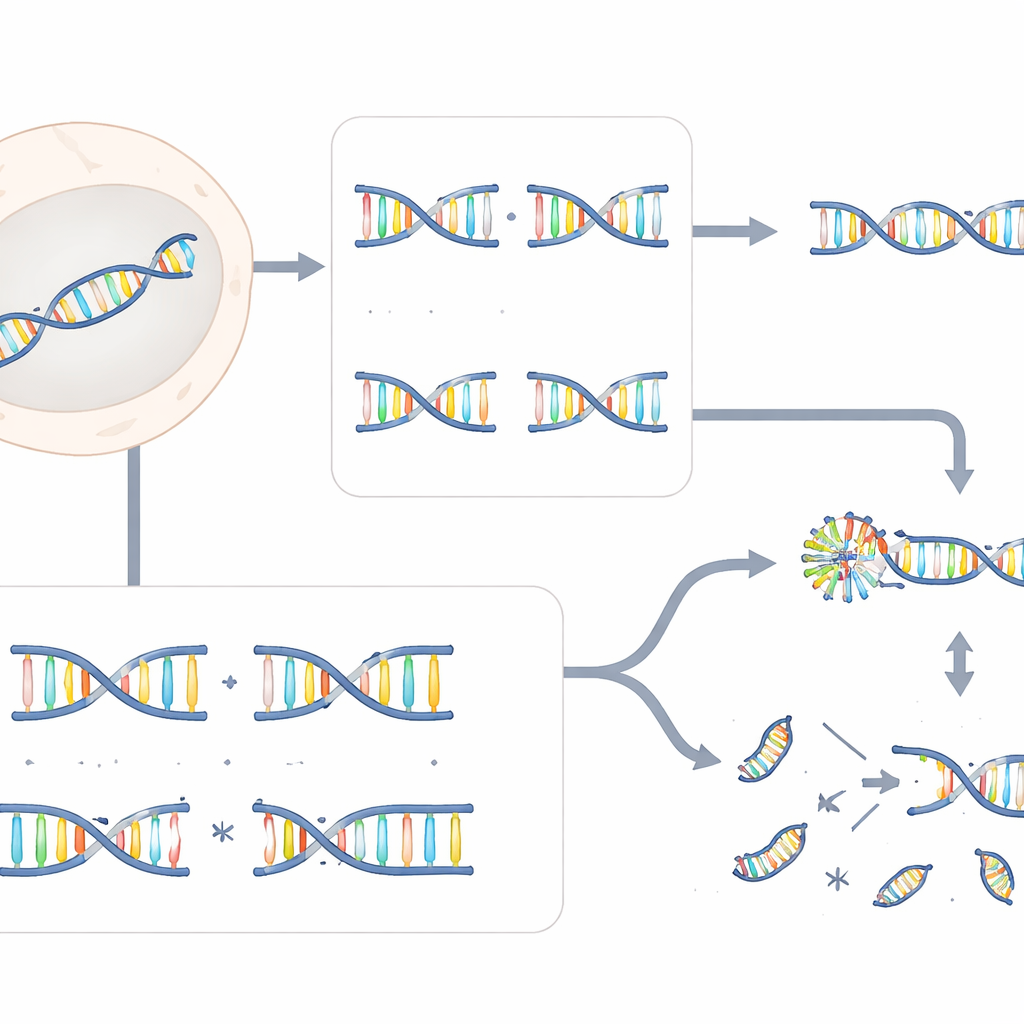
When many cuts pile up, cells begin to fail
The dangers of losing ERCC6L2 became even clearer when the team induced multiple staggered breaks across the genome at once. In these experiments, cells without ERCC6L2 struggled to survive and frequently formed micronuclei—small, misplaced DNA-containing bodies that signal chromosomes have shattered or been mis-segregated. The same vulnerability appeared when the researchers turned to a natural source of staggered breaks: the enzyme TOP2, which temporarily cuts DNA to relieve twisting during normal cell activity. A chemotherapy drug, etoposide, traps TOP2 in its broken state, converting these temporary nicks into lasting staggered breaks. Cells missing ERCC6L2 were dramatically more sensitive to etoposide and showed excessive erosion of DNA ends, mirroring what was seen with Cas12a-induced breaks.
How ERCC6L2 controls the damage machinery
To understand how ERCC6L2 works at the molecular level, the team reconstituted the process in a test tube with purified proteins and DNA. They found that ERCC6L2 binds many DNA shapes but has a special talent: it can actively "melt" or unwind short overhanging ends on staggered breaks, and this requires its energy-consuming motor activity. In cells, ERCC6L2 counterbalances another repair machine, the MRN complex, which normally nibbles back DNA ends to create stretches of single-stranded DNA. This resection is helpful in some repair routes but becomes dangerous when overdone, leading to large deletions and broken chromosomes. In ERCC6L2-deficient cells, MRN-driven resection runs unchecked at staggered breaks. Blocking MRN or its regulator ATM reversed the excessive DNA chewing and reduced the frequency of large deletions, showing that ERCC6L2 normally keeps this pathway in check.
What this means for patients and genome editing
Putting these pieces together, the authors propose that ERCC6L2 acts as a specialist guardian for staggered DNA breaks. By melting the overhangs and promoting quick, tidy rejoining, it prevents long tracts of DNA loss and mis-joining between distant chromosomes. In people born with ERCC6L2 mutations, everyday sources of staggered breaks—particularly those created by TOP2—likely accumulate unrepaired or misrepaired, driving bone marrow failure, leukemia, and possibly certain neurological problems. For gene therapy, the work sends a clear message: genome editing tools that deliberately create overhanging cuts, such as Cas12a, TALENs, or some nickase designs, could be especially risky in these patients. Choosing editing strategies that avoid staggered breaks may be essential to ensure that fixing one gene does not accidentally destabilize the rest of the genome. 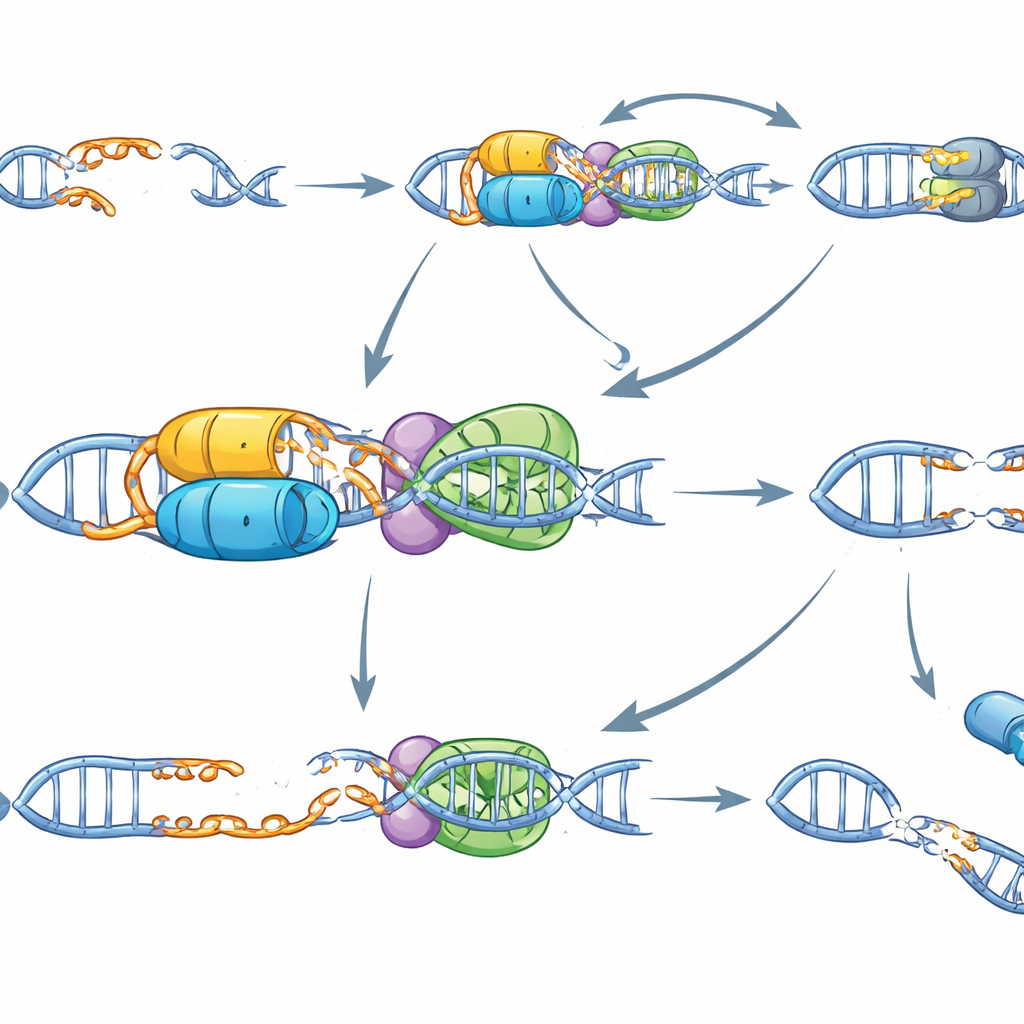
Citation: Aird, E.J., Serrano-Benitez, A., Siegner, S.M. et al. ERCC6L2 ensures repair fidelity for staggered-end DNA double-strand breaks. Nat Commun 17, 2743 (2026). https://doi.org/10.1038/s41467-026-69843-w
Keywords: DNA repair, genome editing, double-strand breaks, ERCC6L2, chromosomal instability