Clear Sky Science · en
Non-hydrolyzable acetyllysine analogs to study protein acetylation in vitro and in cells
Why tiny chemical tags on proteins matter
Inside every cell, proteins are constantly tweaked with small chemical tags that act like on/off switches or dimmers for their activity. One of the most important of these tags is called acetylation, which can change how a protein behaves without changing its underlying genetic code. Understanding exactly what acetylation does at one specific spot on one specific protein has been surprisingly hard, because cells can remove these tags just as quickly as they add them. This study introduces a clever chemical "stand‑in" for acetylation that cannot be removed, giving scientists a way to freeze these switches in place and watch what happens.
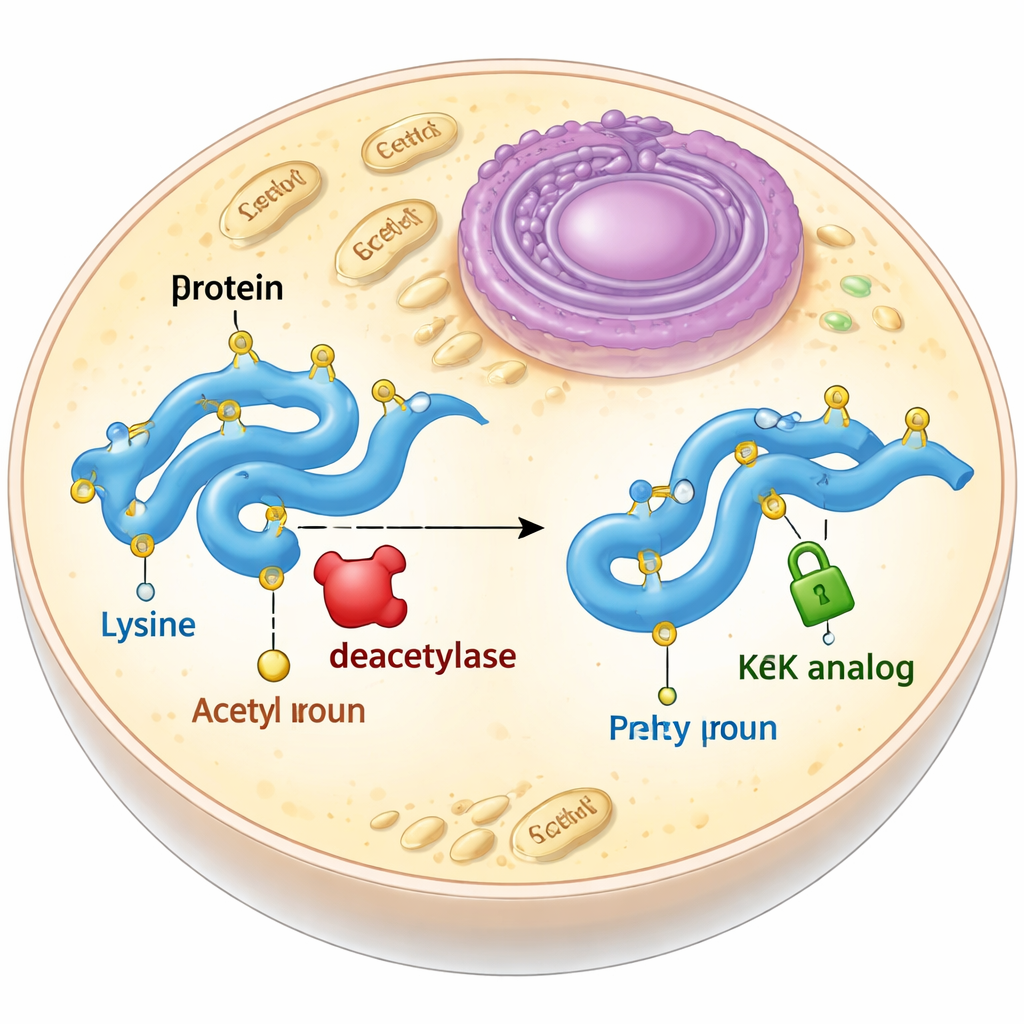
A stable stand‑in for a fleeting tag
Acetylation usually happens on a building block of proteins called lysine. When a lysine is acetylated, it loses its positive charge and its side chain becomes a bit longer. Biologists often try to mimic this by mutating lysine to another amino acid, glutamine, which is uncharged but shorter, or by adding true acetyllysine using an expanded genetic code. Both approaches have drawbacks: glutamine does not fully reproduce the shape changes caused by acetylation, and true acetyllysine is quickly stripped off by enzymes called deacetylases inside cells. The authors set out to design and test acetyllysine look‑alikes that keep the right size and shape but cannot be removed.
Testing new chemical mimics on a model protein
The team focused first on ubiquitin, a small, well‑studied protein that itself gets decorated with many different tags. Using advanced genetic engineering, they placed different lysine variants at one position in ubiquitin: natural acetyllysine, two non‑removable analogs called TFAcK and ketolysine (KeK), and common stand‑ins such as glutamine, alanine, and arginine. They then examined how these changes affected ubiquitin’s shape and function. High‑resolution NMR measurements showed that acetyllysine, TFAcK, and KeK all produced almost identical structural changes, especially around a central helix, while glutamine and other simple mutations did not. Functional tests using an enzyme called HDM2 revealed that ubiquitin carrying acetyllysine, TFAcK, or KeK behaved similarly and differently from the glutamine version, underscoring that the length and bulk of the side chain—not just its charge—are crucial.
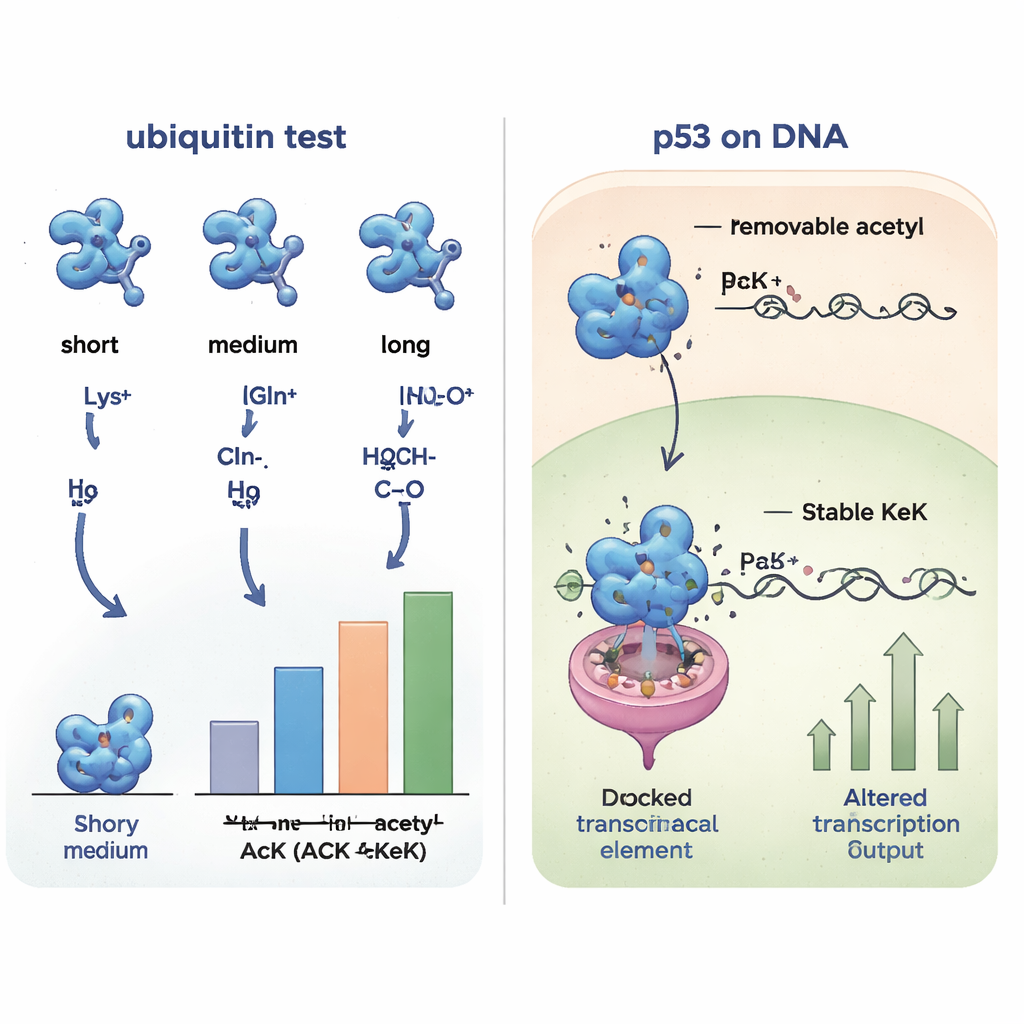
Probing protein partnerships across the cell
Because small structural changes can alter which partners a protein interacts with, the researchers next compared how the various ubiquitin versions bound to other proteins from human cell extracts. By fishing out binding partners and identifying them with mass spectrometry, they found that ubiquitin containing acetyllysine, TFAcK, or KeK shared very similar interaction patterns, while forms containing glutamine or alanine clustered together and looked different. Positively charged variants (lysine or arginine) formed yet another group. This systematic comparison showed that the non‑removable analogs TFAcK and especially KeK mimic both the structure and the binding behavior of true acetylation far better than the traditional glutamine substitution.
Locking in acetylation on the tumor suppressor p53
To see whether these tools work inside living cells, the authors turned to p53, a key tumor suppressor often called the "guardian of the genome." Specific lysines in p53’s DNA‑binding region can be acetylated, shifting which genes it turns on, such as those that stop the cell cycle or trigger cell death. The team engineered human cells so that p53 would incorporate acetyllysine, TFAcK, or KeK at two important lysine positions (120 and 164) only when the corresponding non‑standard amino acid was supplied. All three could be built into p53, but detailed measurement showed that, in cells, the acetyl and TFAcK groups were largely removed by deacetylases, whereas KeK stayed intact. When they tested p53’s ability to activate a reporter gene and the natural p21 gene, variants carrying KeK at these sites were much less active, matching what is expected when acetylation at those positions is locked in, while acetyllysine and TFAcK behaved more like normal p53 because they had been deacetylated.
What this means for studying cell decisions
Overall, the study shows that KeK is a faithful and non‑removable stand‑in for acetyllysine: it closely reproduces the structural changes, binding preferences, and functional consequences of acetylation, but it cannot be erased by cellular enzymes. For non‑specialists, this means researchers now have a way to "freeze" a normally reversible chemical switch at a single site on a protein and then watch how that affects processes such as gene regulation and cell fate decisions. This should make it much easier to disentangle the specific roles of acetylation from those of other overlapping modifications, and could ultimately clarify how mis‑regulated protein acetylation contributes to diseases like cancer.
Citation: Kienle, S.M., Sigg, M., Schneider, T. et al. Non-hydrolyzable acetyllysine analogs to study protein acetylation in vitro and in cells. Nat Commun 17, 1985 (2026). https://doi.org/10.1038/s41467-026-69782-6
Keywords: protein acetylation, posttranslational modification, ubiquitin, p53, genetic code expansion