Clear Sky Science · en
Spatial perturb-seq: single-cell functional genomics within intact tissue architecture
Seeing Genes at Work in Living Brain Circuits
Many brain diseases, from Alzheimer’s to Parkinson’s, are linked to risky genes, but we still know little about how those genes affect individual cells inside the living brain. This study introduces a powerful way to watch what happens when specific genes are switched off in single cells, while those cells remain in their natural neighborhoods inside brain tissue. The approach, called Spatial Perturb-Seq, helps connect genetic risk to disrupted cell behavior and cell-to-cell communication in the intact brain.
A New Way to Test Many Genes at Once
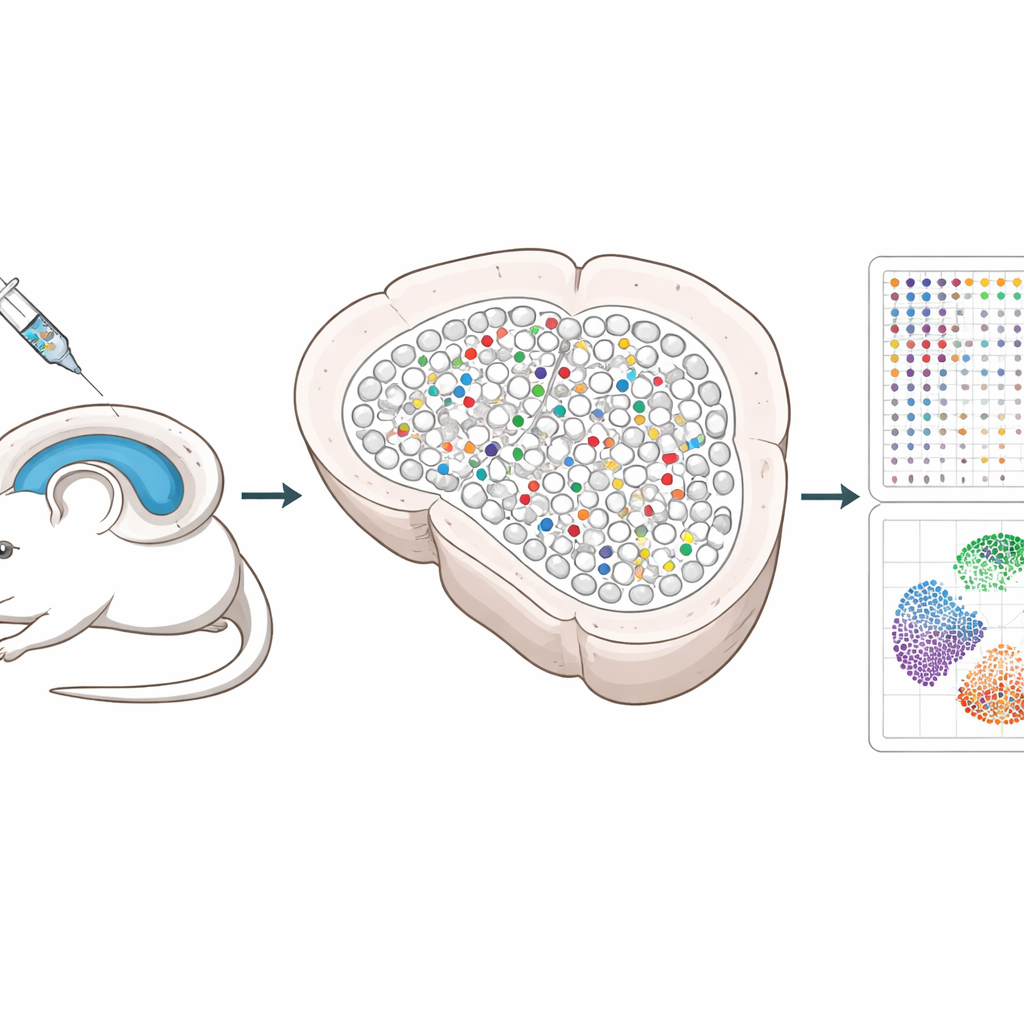
The researchers built a system that uses CRISPR gene-editing tools delivered by harmless viruses to the hippocampus, a brain region important for memory. Each virus carries three guides that disable a chosen gene, along with a unique DNA “barcode” and a fluorescent marker so edited cells can be found later. By mixing these viruses together and injecting them at low dose, only a small, scattered fraction of cells are edited, and most neighbors remain untouched. This sparse pattern lets scientists separate the changes happening inside edited cells from those happening in nearby healthy cells.
Keeping the Brain’s Map Intact
Traditional single-cell methods require tissues to be broken apart, which destroys the precise positions of cells and can even lose fragile neurons. Spatial Perturb-Seq instead reads gene activity directly from thin brain slices, so every cell’s location is preserved. Using a technology called Stereo-seq, the team captured full gene-expression profiles for more than 200,000 cells, while also reading the CRISPR barcodes and recording each cell’s coordinates in the tissue. They then used advanced image-based algorithms to outline every cell and computational tools to identify cell types and local neighborhoods, focusing especially on hippocampal neurons and their close contacts.
How Edited Cells Influence Their Neighbors
With the tissue map in hand, the scientists compared edited neurons to unedited neurons and, separately, to the unedited neighbors surrounding each edited cell. This allowed them to distinguish “cell-autonomous” effects (inside the edited cell) from “non-cell-autonomous” effects (in the surrounding microenvironment). Knocking out 18 different genes, many tied to neurodegenerative disease risk, produced distinct patterns of gene changes inside the edited neurons. Some edits, such as those targeting the gene Cfap410, also triggered strong shifts in gene activity in nearby cells, reflecting altered local signaling and support.
Clues to Neurodegenerative Disease Pathways
Several genes offered especially intriguing insights. Disabling Lrrk2, a major Parkinson’s disease risk gene, caused large changes in the edited neurons, including reduced levels of Bc1, an RNA molecule important for tuning protein production at synapses. At the same time, neighboring cells showed altered expression of genes involved in synapse structure, protein trafficking, and calcium handling, hinting that Lrrk2-linked problems ripple through local circuits. Knocking out another gene, Srf, disturbed a network of genes tied to neuronal growth and plasticity, and weakened specific ligand–receptor signaling pathways between neurons, indicating disrupted cell-to-cell communication. By systematically scoring these signaling pairs, the team could pinpoint which communication routes were most affected for each gene.
Flexible Tools for Mapping Genetic Effects
To show that Spatial Perturb-Seq is not locked to a single platform, the researchers also adapted it to an imaging-based system called Xenium. There, pre-designed probe panels and custom barcode probes read out selected genes and barcodes directly in the tissue. While this targeted approach covers fewer genes than whole-transcriptome Stereo-seq, patterns of gene changes for key edits such as Lrrk2 and Srf agreed well between the two methods and with a large external brain atlas. This cross-checking supports the robustness of the observed gene and neighborhood effects.
What This Means for Brain Health Research
In everyday terms, Spatial Perturb-Seq lets scientists flick off many different genes in individual brain cells, then watch how those cells and their immediate neighbors react, all while the tissue’s wiring diagram stays intact. This makes it possible to trace how genetic risk factors disturb not just single cells but also the conversations between them, which are crucial for healthy brain function. Over time, as sequencing becomes cheaper and larger experiments are run, this approach could help reveal which genes and local circuits are most important in early stages of disorders like Alzheimer’s, Parkinson’s, and ALS, guiding more precise strategies for treatment.
Citation: Shen, K., Seow, W.Y., Keng, C.T. et al. Spatial perturb-seq: single-cell functional genomics within intact tissue architecture. Nat Commun 17, 3018 (2026). https://doi.org/10.1038/s41467-026-69677-6
Keywords: spatial genomics, CRISPR screens, single-cell sequencing, neurodegeneration, cell-cell communication