Clear Sky Science · en
FLEXTAG: a small and self-renewable protein labeling system for anti-fading multi-color super-resolution imaging
Seeing the Invisible Inside Cells
Much of modern biology depends on taking pictures of the molecules that keep our cells alive. But even our best light microscopes struggle with a simple problem: the glowing tags used to highlight proteins quickly fade, and it is hard to label many different targets at once without disturbing how cells work. This paper introduces FLEXTAG, a new family of tiny, renewable tags that keep glowing far longer, support multiple colors, and work with the most powerful microscopes scientists use to see the inner architecture of cells.
Why Sharper Cellular Photos Are So Hard
Conventional fluorescence microscopes can reveal structures about a quarter of a micrometer across, which is still far bigger than most individual proteins. Super-resolution methods squeeze out more detail, reaching down to a few billionths of a meter, but only if the fluorescent labels behave perfectly. Today’s popular protein labels are either bulky antibodies, dim fluorescent proteins, or chemical tags whose dyes quickly burn out under the intense light needed for these techniques. Fixing cells with chemicals for detailed imaging can also lock proteins together in ways that block dyes from reaching their targets, and stray dye molecules may stick nonspecifically, clouding the picture with background glow.
A New Kind of Refillable Protein Tag
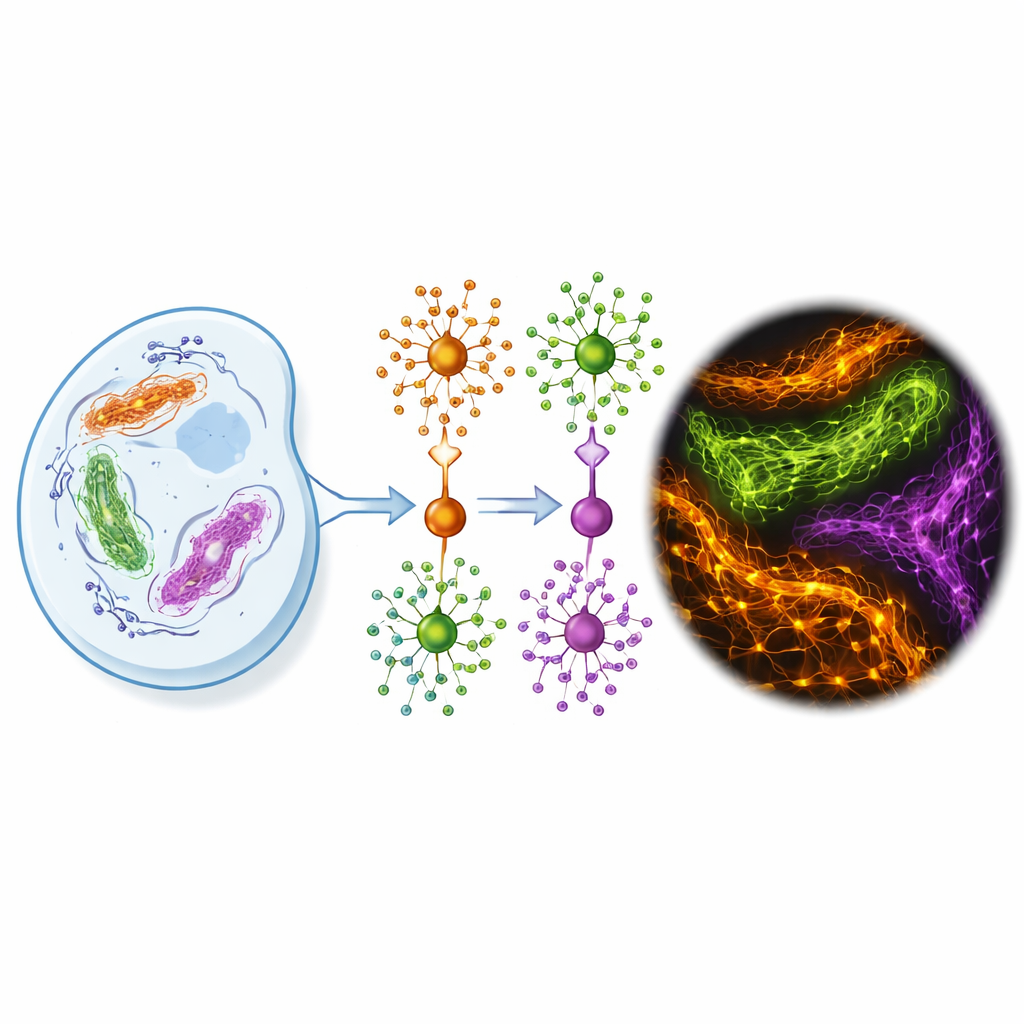
The authors designed FLEXTAG (short for Fluorescent Labeling for Exchangeable, X-resilient Tagging in Advanced Generic Nanoscopy) to tackle these limitations head-on. FLEXTAG is not a single tag but a coordinated trio—FLEXTAG1, FLEXTAG2, and FLEXTAG3—each a small engineered protein (12–18 kilodaltons, roughly half the size of classic tags like GFP and far smaller than HaloTag). Each FLEXTAG binds a matching small-molecule ligand that carries a bright organic dye. Crucially, this binding is reversible: dye–ligand molecules continuously come and go. When one fluorophore is damaged by light, a fresh one from the surrounding solution takes its place, so the signal effectively “self-renews” instead of steadily fading.
Building Three Compact and Reliable Tags
To create FLEXTAG1–3, the team repurposed three well-known protein scaffolds from cell biology and drug design, then used structural modeling and imaging-based assays to reshape their behavior. FLEXTAG1 is derived from a bromodomain protein that recognizes a modified small molecule; the authors introduced mutations to break up its tendency to pair up and form clumps, while preserving strong binding to its ligand. FLEXTAG2 comes from bacterial dihydrofolate reductase. By adding a strategic disulfide bond and tuning flexible linkers, they stabilized the protein and dramatically increased how many tags carried dye at any moment, all while keeping binding reversible. FLEXTAG3 is based on a human FKBP protein used in chemical biology; here, the team balanced binding strength and speed so that dyes come off quickly enough to be replaced, yet still bind strongly enough to give bright images and avoid excessive aggregation.
Protecting Tags During Fixation and Cutting Background Glow
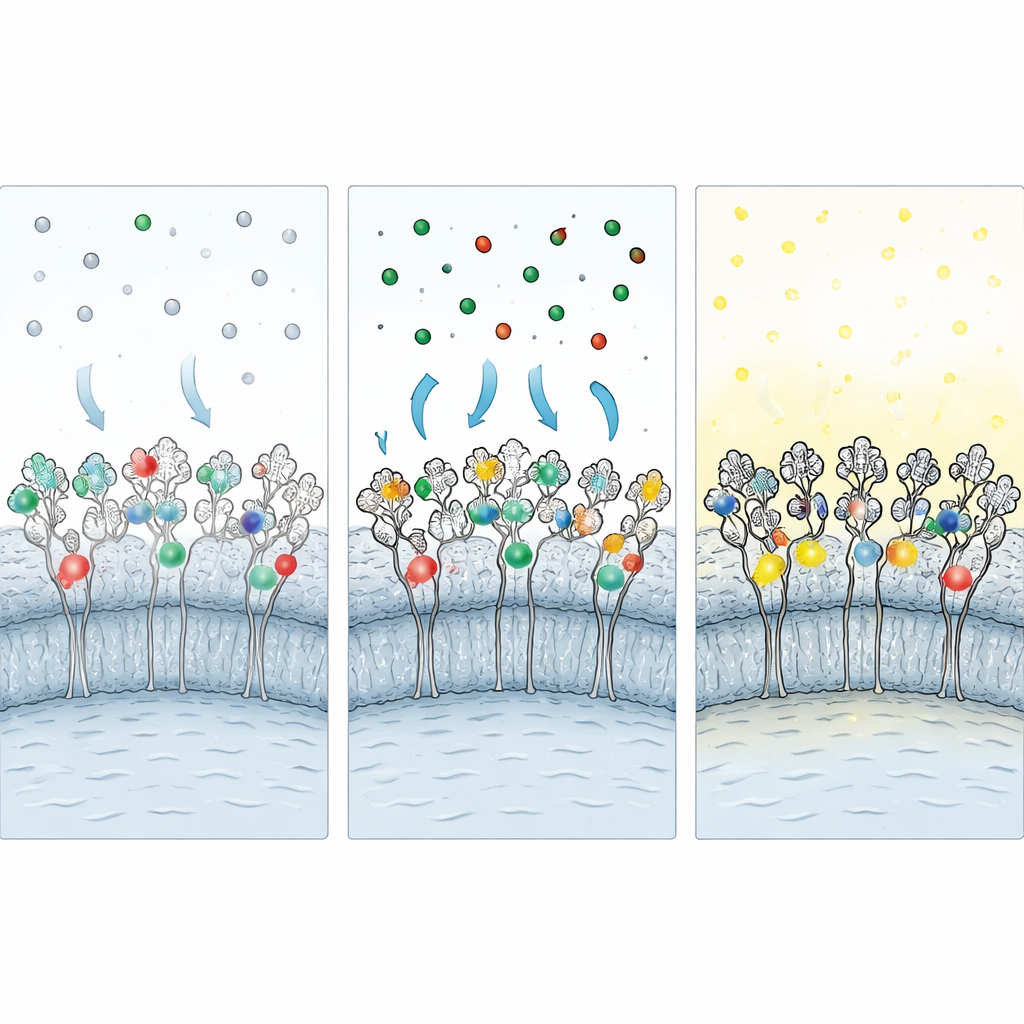
Because many important experiments require imaging fixed cells, the researchers developed a “protective fixation” strategy. Before adding aldehyde fixatives, they flood living cells with unlabeled versions of each ligand, plugging the tag’s binding pocket. During fixation, these protective ligands shield vulnerable amino acids from chemical crosslinking. After fixation, the protective ligands are washed out and replaced with fluorescent ones, restoring access to the tag. Additional steps—chemical reduction to neutralize leftover reactive groups and a blocking cocktail containing albumin, detergent, and a chaotropic salt—further reduce sticky, nonspecific binding of free dyes. Together, these measures preserve around 60–70% of the labeling seen in live cells while sharply improving signal-to-noise.
Sharper, Longer-Lasting Movies Across Many Methods
With the FLEXTAG framework in place, the authors put it through its paces across the major families of super-resolution microscopy. In patterned-illumination methods such as SIM and STED, FLEXTAGs allowed multi-color imaging of mitochondria, microtubules, the endoplasmic reticulum, and actin with far less fading than standard covalent tags; over dozens of imaging cycles, signal from conventional tags dropped by half or more, while FLEXTAG signals stayed nearly constant. In single-molecule methods such as PAINT and STORM, the fast on–off exchange of FLEXTAG ligands produced rich, stable streams of localizations that could be turned into three-dimensional, multi-color maps of subcellular structures, even in live cells over many minutes. FLEXTAG2 showed particularly favorable kinetics for PAINT, while FLEXTAG3 excelled in long-lived STORM movies. Because the tags are orthogonal to each other and compatible with a broad palette of dyes, scientists can label several proteins at once and then choose whichever imaging mode best suits their question without changing the underlying constructs.
What This Means for Looking Inside Cells
FLEXTAG offers a kind of universal, refillable connector between proteins and bright dyes. Its tiny size reduces the risk that the tag will distort where a protein goes or how it works, and its self-renewing dyes help sidestep the long-standing problem of photobleaching in high-powered microscopes. By pairing these tags with protective fixation and background-suppression chemistry, researchers can obtain cleaner, longer-lasting, and more colorful views of how proteins are organized and move inside cells. In practical terms, this means better maps of cellular architecture, more reliable tracking of molecular interactions over time, and a versatile toolkit that should benefit both basic cell biology and translational studies that rely on seeing disease-related changes at the nanoscale.
Citation: Zhang, H., Yao, Y., Wang, X. et al. FLEXTAG: a small and self-renewable protein labeling system for anti-fading multi-color super-resolution imaging. Nat Commun 17, 2156 (2026). https://doi.org/10.1038/s41467-026-69658-9
Keywords: super-resolution microscopy, fluorescent protein tagging, live-cell imaging, photobleaching resistance, cellular architecture