Clear Sky Science · en
XL-MSDigger: a deep learning-based, versatile solution for cross-linking mass spectrometry
Seeing How Proteins Hold Together
Every process in our bodies depends on proteins not only folding into the right shapes but also finding the right partners. Yet watching these molecular relationships in action is notoriously hard. This study introduces XL-MSDigger, a software platform that uses modern artificial intelligence to pull much clearer signals out of a noisy experimental technique called cross-linking mass spectrometry, helping scientists map how proteins are arranged and who they interact with inside cells.
Untangling a Crowded Molecular World
To learn how proteins are built and how they connect, researchers often use cross-linking mass spectrometry. In this approach, small chemical “bridges” link nearby parts of proteins together. The linked pieces are then broken into fragments and weighed in a mass spectrometer. In principle, the pattern of fragments reveals which protein pieces were close in space, like finding which pages of a book were clipped together. In practice, however, the resulting data are extremely complex. Existing computer tools mostly look only at the basic mass information and struggle with the enormous number of possible combinations, leading to missed connections and spurious matches.
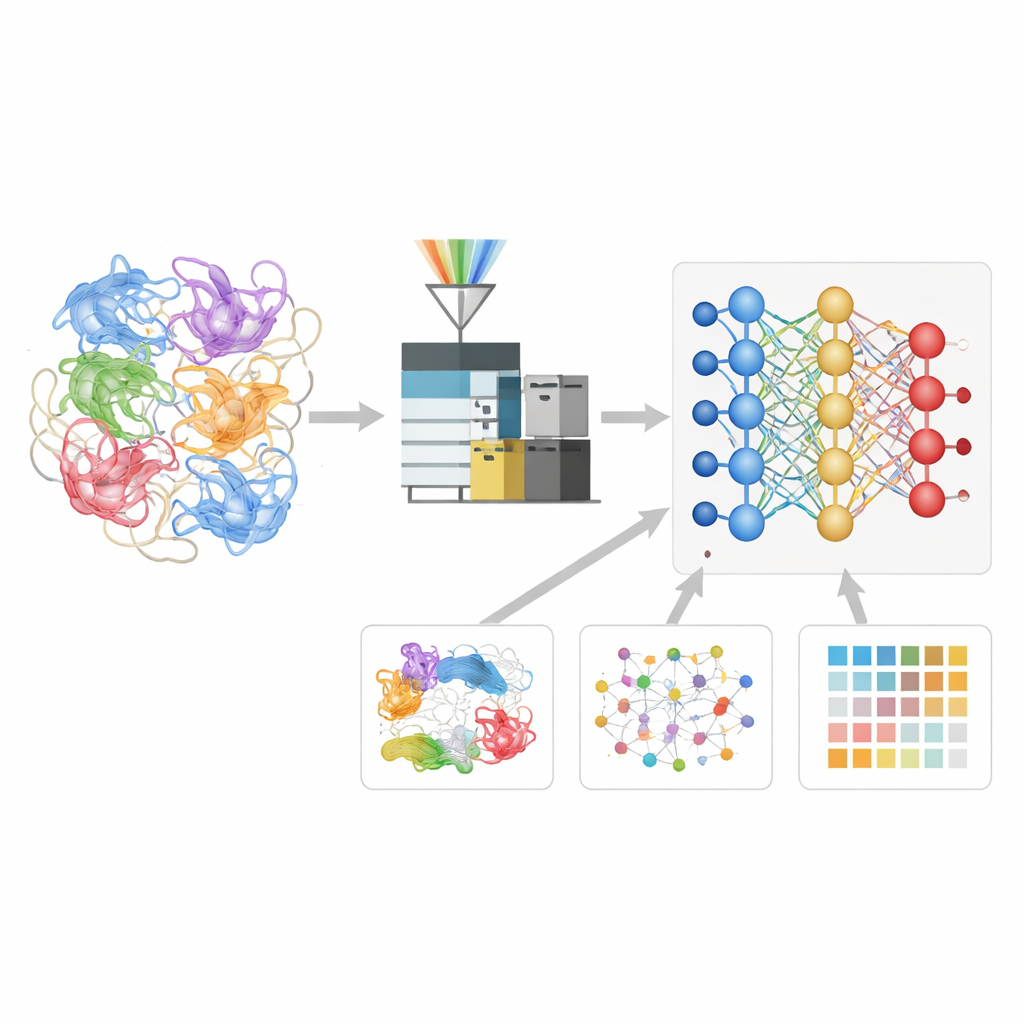
Teaching a Neural Network the Language of Protein Fragments
The authors built a deep learning model called Deep4D-XL to better interpret these cross-linking experiments. They first created a large reference set by cross-linking proteins from human cells, breaking them into peptides, and recording not just their masses but also how long they took to travel through the instrument and how they moved through an ion-mobility chamber. Each cross-linked pair was encoded for the model, which uses a twin “Siamese” design to read both peptide partners and a cross-attention step to combine their information. From this, the network learns to predict three key properties of any new cross-linked peptide: when it should appear in the experiment, how it should move, and what its fragmentation pattern should look like.
Turning Predictions into Cleaner Signals
XL-MSDigger wraps this prediction engine in analysis workflows for two major data-collection styles. In the traditional, targeted style, the instrument selectively records fragments from ions it chooses on the fly. XL-MSDigger takes the initial matches from established search software and re-evaluates them using the model’s predicted behavior for each candidate. A second neural network compares prediction and experiment along several dimensions and assigns improved scores. This rescoring step nearly doubles the number of confidently detected links between different proteins in yeast and human samples while keeping error rates low, revealing many more protein–protein interactions than before.
Making Sense of Floods of Unbiased Data
A newer way to run these instruments, called data-independent acquisition, records fragments for almost everything in a sample, improving coverage but generating overwhelming data. Until now, there has been no good way to estimate how many of the resulting cross-links were truly real. XL-MSDigger uses Deep4D-XL to build a carefully matched “decoy” library of fake cross-links, then analyzes both real and decoy entries together. By seeing how often decoys slip through, the software can estimate the false discovery rate and train another neural network to separate true from false matches. This rescoring boosts the number of trustworthy cross-linked signals by roughly fivefold and produces clear separation between real and decoy patterns.
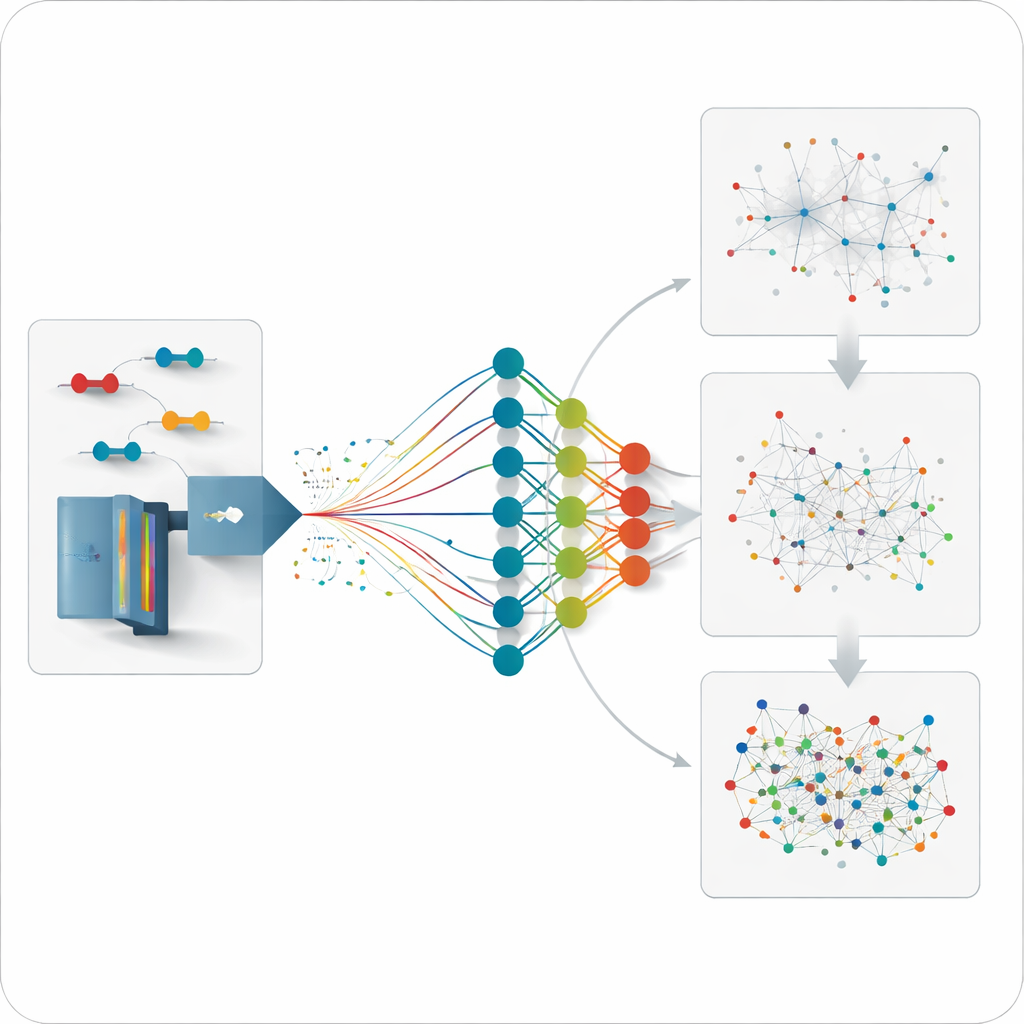
Predicting What Hasn’t Been Measured Yet
Because the model can forecast how any plausible cross-linked peptide should behave, the team can go a step further and analyze data for links that were never directly measured before. They generate moderate-sized predicted libraries focusing on selected proteins or interaction networks and then search the unbiased data against these libraries. This strategy uncovers additional links within single proteins and between partners of important chaperone proteins, with distances that agree well with known three-dimensional structures. It also recovers interactions missed by the traditional, more limited experimental libraries, especially for low-abundance connections.
Opening a Clearer Window on Protein Partnerships
For non-specialists, the key message is that XL-MSDigger acts like a highly trained pattern recognizer layered on top of an already powerful experimental method. By learning what genuine cross-linked signals should look like in several dimensions at once, it can sift through vast, messy datasets, discard likely impostors, and rescue real but previously hidden protein connections. While full, whole-proteome applications will still demand heavy computing power, this work shows that combining cross-linking experiments with deep learning can greatly sharpen our view of how proteins are arranged and who they meet inside the cell.
Citation: Chen, M., Hao, Y., Huang, X. et al. XL-MSDigger: a deep learning-based, versatile solution for cross-linking mass spectrometry. Nat Commun 17, 2554 (2026). https://doi.org/10.1038/s41467-026-69489-8
Keywords: protein interactions, cross-linking mass spectrometry, deep learning, proteomics, data-independent acquisition