Clear Sky Science · en
Structural and molecular basis for allosteric regulation and catalytic coupling of human phosphoribosylformylglycinamidine synthase
Why This Enzyme Matters for Health
Every cell in your body depends on purines, small molecules that power reactions, store energy, and carry genetic information. Cancer cells, which divide rapidly, are especially hungry for purines and often rewire how they are made. This study focuses on a human enzyme called FGAMS, a key player in building purines from scratch. By revealing FGAMS’s three-dimensional structure and how its parts move and communicate, the authors uncover how the enzyme is switched on and off and how it might be targeted to slow tumor growth.
Building Blocks of Life Under Control
Cells make purines through a demanding assembly line known as the de novo pathway, which uses energy and several enzymes to stitch together a complex ring. FGAMS performs the fourth step in this sequence and sits at the heart of a larger enzyme cluster called the purinosome, which helps channel ingredients efficiently. Because fast-growing cancer cells often boost this pathway, enzymes like FGAMS are attractive drug targets: blocking them can starve tumors of the nucleotides they need while healthy cells fall back on recycling routes.
A Shape-Shifting Molecular Machine
FGAMS is built from three connected parts, or domains, that each carry out a different job and must cooperate in a precise order. One domain cuts ammonia from the amino acid glutamine, another uses that ammonia to modify a small sugar-based molecule, and a third domain acts as a regulatory hub. Using high-resolution cryo-electron microscopy, the researchers captured FGAMS in three distinct working states: with an energy-carrying molecule bound, with a short-lived reaction intermediate trapped, and with multiple substrates and intermediates present at once. Comparing these states revealed that flexible loops and an N-terminal “arm” move dramatically, reconfiguring the enzyme as chemistry progresses.
Hidden Tunnel and Molecular Gates
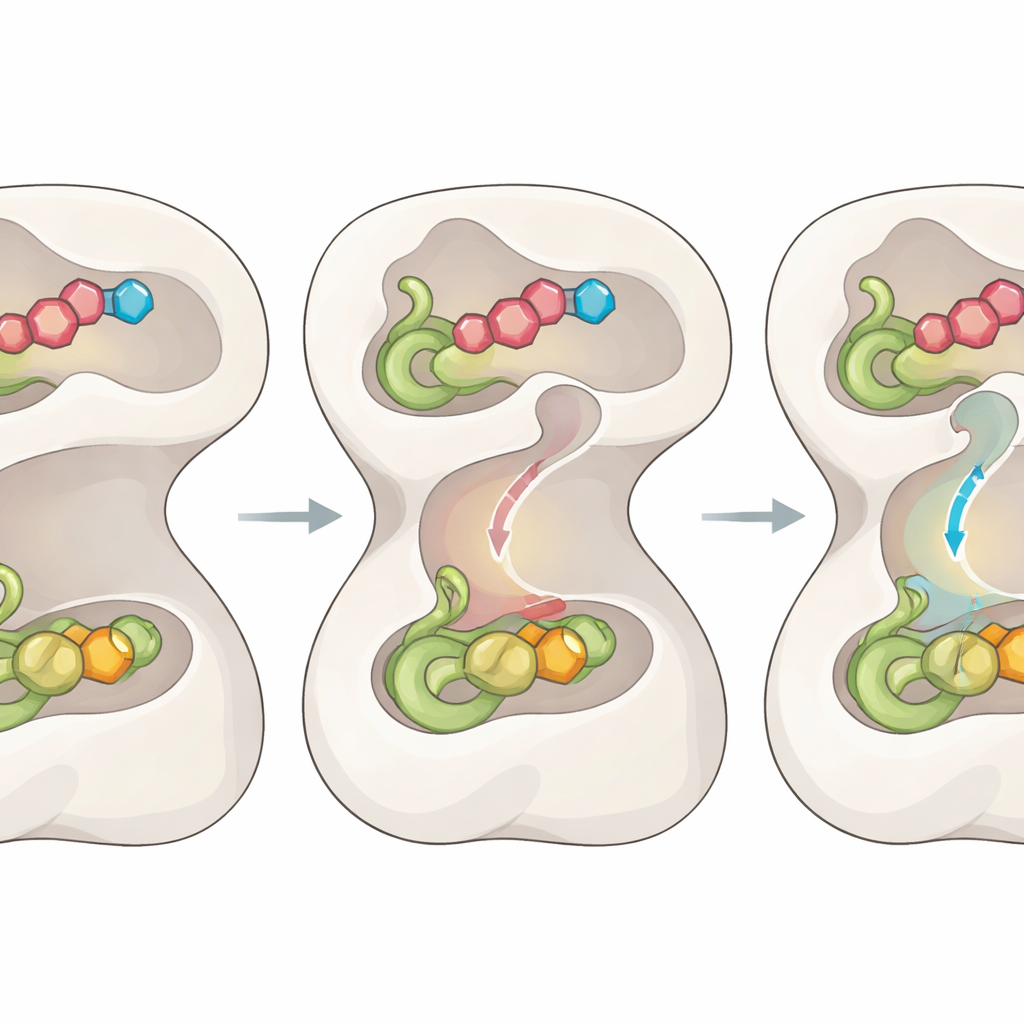
A key puzzle has been how the ammonia released in one domain travels safely to the distant site where it is used, without diffusing away and causing damage. The new structures expose a previously unseen internal tunnel that connects the two sites, along with a series of “gates” formed by specific amino acids. When the proper substrates bind and the intermediate forms, a central loop snaps into an ordered position, tugging on the N-terminal domain and rearranging nearby loops in the catalytic domain. This choreography opens two entry gates so glutamine can be processed and briefly reshapes a region that stabilizes the high‑energy intermediate needed for bond breaking.
Coupling Chemistry Through Motion
As the reaction proceeds, the enzyme does not simply wait; it uses each chemical step as a signal. Once a covalent thioester intermediate forms in the ammonia-generating domain, the earlier motions are partially reversed. This relaxes the central loop and repositions the gates to close off escape routes, while two additional gates at either end of the tunnel swing open. Subtle shifts in a handful of “bottleneck” residues widen the inner passage, allowing an ammonia molecule to travel through the tunnel directly into the second active site, where it reacts with the trapped intermediate to produce the final product. Mutating the gate residues causes ammonia to leak or the two halves of the reaction to uncouple, confirming that this gating-and-channel system is essential for efficient catalysis.
From Structural Insight to Therapeutic Avenues
By following FGAMS through its catalytic cycle, the study lays out a detailed picture of how domain motions, flexible loops, internal gates, and a transient tunnel are all wired together to coordinate chemistry. For a lay audience, the bottom line is that the enzyme acts like a tiny programmable factory: it only opens its doors and internal conveyor belt when all the right parts are in place, ensuring speed and safety. Because FGAMS supports purine production in cancer cells and is linked to poor survival in liver tumors, the structural blueprints and newly identified control points offer multiple footholds for designing drugs that jam the tunnel, freeze the gates, or trap key intermediates, potentially yielding a new class of targeted anticancer therapies.
Citation: Sharma, N., Zhou, W. & French, J.B. Structural and molecular basis for allosteric regulation and catalytic coupling of human phosphoribosylformylglycinamidine synthase. Nat Commun 17, 2732 (2026). https://doi.org/10.1038/s41467-026-69423-y
Keywords: purine biosynthesis, allosteric regulation, ammonia channeling, cancer metabolism, cryo-EM structure