Clear Sky Science · en
Molecular mechanism of action of small molecule SMT-738 on bacterial lipoprotein transporter LolCDE
Why this new antibiotic matters
As more hospital infections shrug off our best antibiotics, doctors urgently need drugs that can kill dangerous bacteria without wrecking the helpful microbes we carry. This study unpacks how a promising experimental compound, called SMT-738, shuts down a crucial transport machine in the outer shell of certain hard-to-treat bacteria. By revealing exactly where and how this drug jams the machinery, the work points the way to smarter antibiotics that hit superbugs while sparing much of the gut microbiome.
The outer armor of bad bacteria
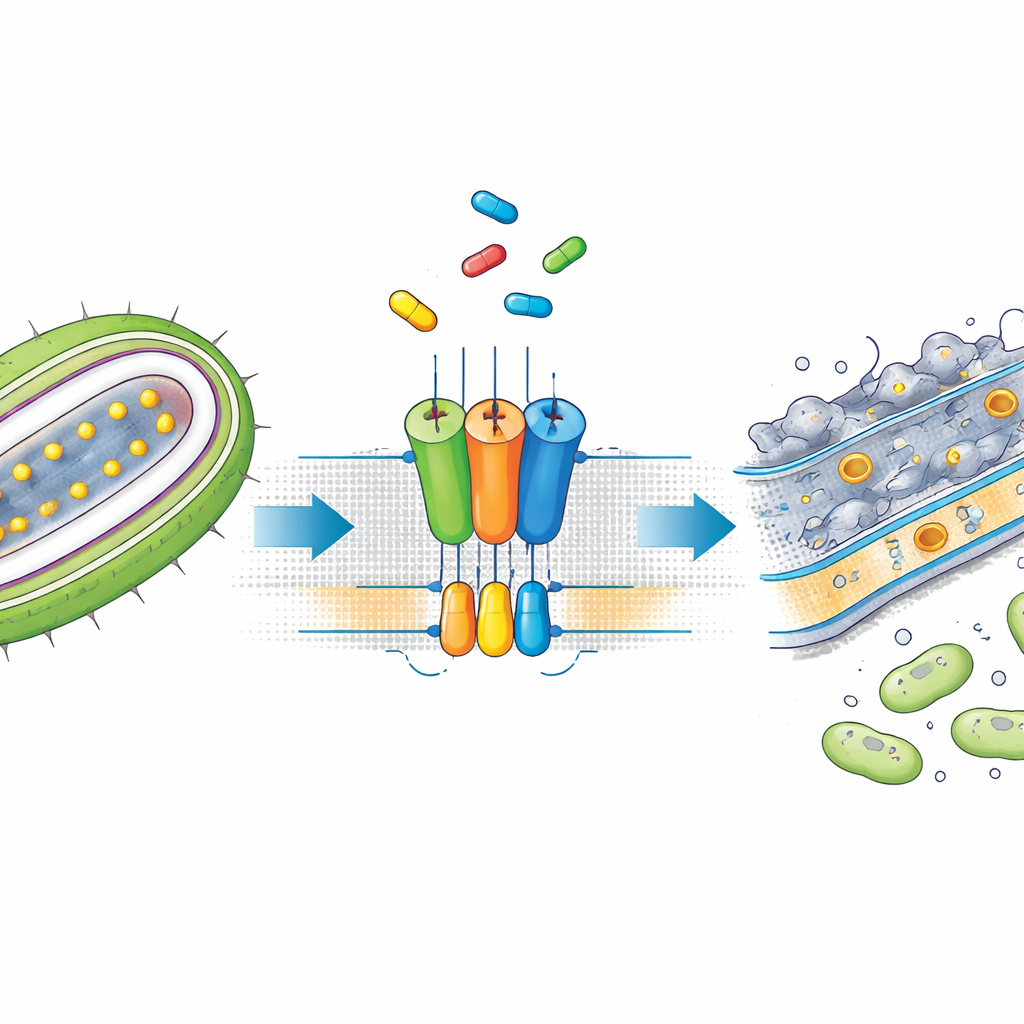
Many of the most worrisome hospital pathogens belong to a group known as Gram-negative bacteria. They are tough to kill because they wrap themselves in a double-layered outer armor. This outer layer is built and maintained by several molecular machines that move key building blocks from the inner membrane, where they are made, to the surface. One important set of cargo is lipoproteins, which help keep the outer shell intact and support processes like nutrient uptake and antibiotic resistance. A transport system called the Lol pathway, and in particular a pump named LolCDE, extracts lipoproteins from the inner membrane and hands them off to chaperones that deliver them to the outer surface. Because these steps are essential, they are attractive targets for new antibiotics.
A small molecule that stops the conveyor belt
SMT-738 emerged from earlier screens as a potent blocker of Gram-negative pathogens belonging to the Enterobacteriaceae family, including multidrug-resistant strains of Escherichia coli and Klebsiella pneumoniae. Yet its precise mode of action was unclear. The authors first showed that SMT-738 prevents a test lipoprotein from being released by LolCDE in engineered bacteria, confirming that the drug really stalls the transport step. Importantly, when bacteria carried certain mutations in the genes for the LolCDE complex, they became resistant to SMT-738, suggesting that the drug works by binding directly to this pump. These observations set the stage for a structural deep dive into how the compound interacts with the transporter.
Jamming the pump from the inside
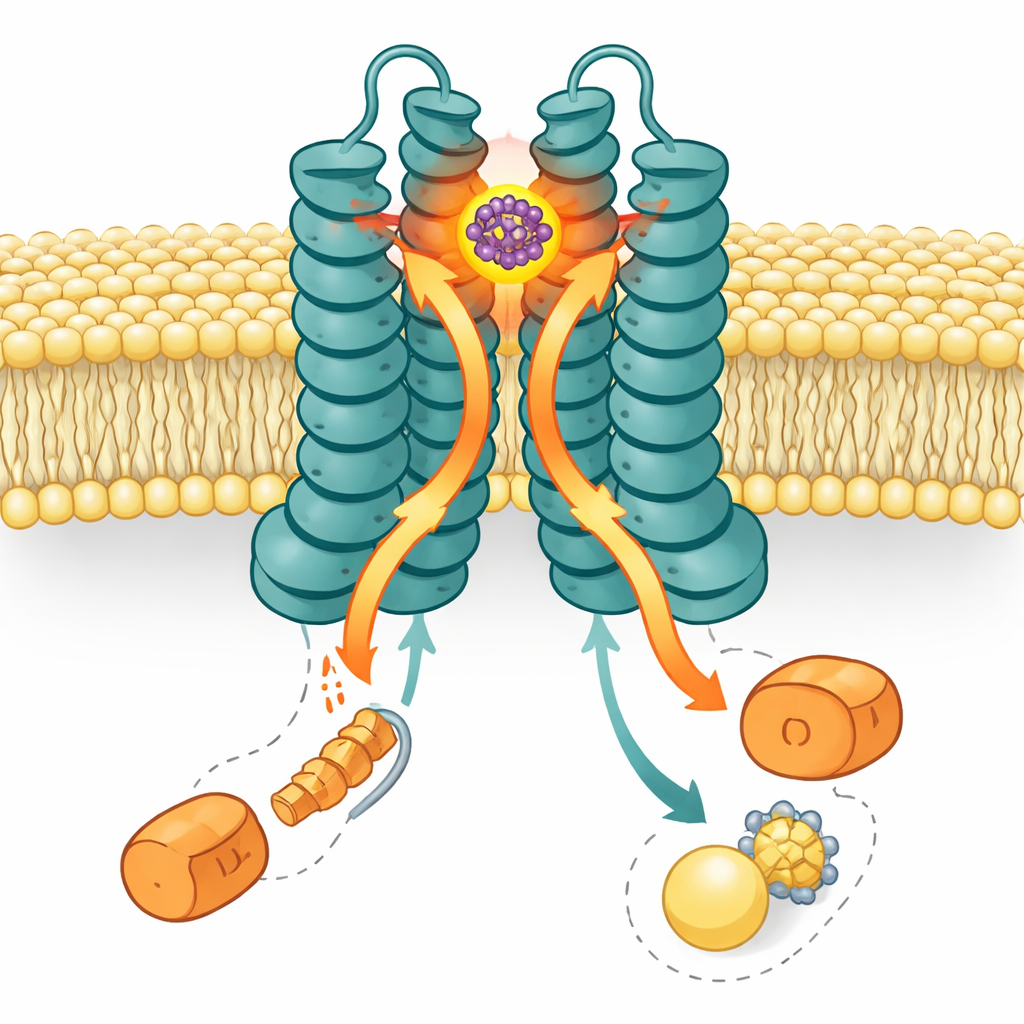
Using high-resolution cryo–electron microscopy, the researchers captured a three-dimensional image of LolCDE with SMT-738 locked in place. The structure shows the drug wedged in a pocket at the periplasmic end—the side facing the space between the inner and outer membranes—between two core subunits, LolC and LolE. This site overlaps the normal docking area for lipoprotein cargo. When SMT-738 occupies this pocket, surrounding amino acids in LolC and LolE shift position by up to about one nanometer. Those movements create steric clashes with where the lipoprotein and its fatty tails would normally sit, effectively blocking cargo from entering the pump. Mutating many of these pocket-lining residues either weakened drug binding or made bacteria resistant, neatly matching the structural picture.
Triggering a one-sided breakdown
The story does not end at the binding site. LolCDE is powered by two copies of a motor subunit called LolD, which sit on the cytoplasmic side and burn ATP to fuel transport. Strikingly, in the SMT-738–bound structure, only one LolD remains attached; the partner LolD has dissociated. Biochemical gels and ATPase assays confirmed that binding of SMT-738 causes loss of one LolD and sharply reduces the pump’s energy-turning activity. Computer simulations of the transporter in a membrane reinforced this view: as SMT-738 settled into its pocket, conformational changes rippled downward, nudging the coupling helices that connect LolC and LolE to LolD. These shifts pushed the LolD bound to LolC into an unfavorable clash, causing it to detach, while the LolD bound to LolE stayed put. The result is an asymmetric, “deadlocked” machine that can no longer move lipoproteins.
Why some bacteria are hit and others are spared
SMT-738 has a desirable trait: it strongly attacks drug-resistant Enterobacteriaceae but leaves many other Gram-negative species in the gut largely untouched. To understand this selectivity, the team compared sequences of LolE from susceptible and resistant bacteria and focused on positions that contact SMT-738. Two residues in LolE, especially one corresponding to position D264 in E. coli, stood out. In sensitive pathogens, these positions matched the drug-contacting version in E. coli, whereas in many resistant gut commensals they were replaced by other amino acids. When the researchers engineered E. coli to carry such substitutions, the bacteria became highly resistant to SMT-738 while still expressing LolCDE. Simulations suggested that these changes alter local flexibility around the pocket, reducing the ability of SMT-738 to bind. Interestingly, another LolCDE inhibitor, lolamicin, relied on only partly overlapping contact points and showed a different resistance pattern, hinting at distinct ways of targeting the same pump.
New blueprints for future antibiotics
By combining structural imaging, mutational scans, biochemical tests, and computer simulations, this work shows that SMT-738 is not just a plug in a channel. Instead, it latches onto the periplasmic side of the LolCDE pump, blocks the lipoprotein entry site, and then triggers long-range shifts that knock off one of the two motor units, freezing the transporter in a non-working state. This “allosteric deadlock” mechanism represents a new way to disable a bacterial machine and helps explain why SMT-738 can zero in on dangerous pathogens while sparing much of the microbiome. The detailed map of the binding pocket and the key selectivity residue in LolE now offer a rational blueprint for designing next-generation antibiotics that exploit the same vulnerability with even greater potency and precision.
Citation: Li, H., Zhu, X., Zhang, D. et al. Molecular mechanism of action of small molecule SMT-738 on bacterial lipoprotein transporter LolCDE. Nat Commun 17, 2540 (2026). https://doi.org/10.1038/s41467-026-69411-2
Keywords: antibiotic resistance, Gram-negative bacteria, lipoprotein transport, LolCDE inhibitor, cryo-EM structure