Clear Sky Science · en
Seq-Scope-eXpanded: spatial omics beyond optical resolution
Seeing Cells in Sharper Detail
Our bodies are built from countless tiny cells, each bustling with activity that we usually can’t see, even with powerful microscopes. This paper introduces Seq-Scope-X, a technology that lets researchers zoom in on the inner workings of tissues with unprecedented clarity, mapping which genes and proteins are active in precise spots inside and between cells. That level of detail could change how we study development, the immune system, and diseases such as cancer or liver damage.
Why Map Molecules in Place?
Cells do not work in isolation: they live in neighborhoods, form layers, and specialize depending on where they are. Traditional gene sequencing methods break tissues apart, losing this spatial context. More recent “spatial omics” tools keep molecules in place, but there has been a trade-off. Methods based on imaging can see very fine detail but usually track only a limited set of genes. Sequencing-based methods can read almost every gene at once, but they blur fine features, often smearing signals over several micrometers—larger than many subcellular structures. The authors set out to close this gap: keep the rich, unbiased readout of sequencing, while reaching or even beating the sharpness of modern microscopes.
Stretching Tissue to Beat Optical Limits
The core idea behind Seq-Scope-X is deceptively simple: gently enlarge the tissue itself so that each original nanometer of structure becomes easier to resolve. The team first anchors RNA molecules—or special DNA tags attached to antibodies—into a soft hydrogel that is built around the tissue slice. They then digest away the original tissue and swell the gel in a salty solution, physically stretching everything by about threefold while keeping relative positions nearly intact. This expanded gel is laid on top of an ultra-dense sequencing chip coated with capture probes. By carefully heating the assembly, the anchored molecules are released from the gel and reattached to the chip, which can then be sequenced to reveal which genes or proteins were present at each tiny coordinate. 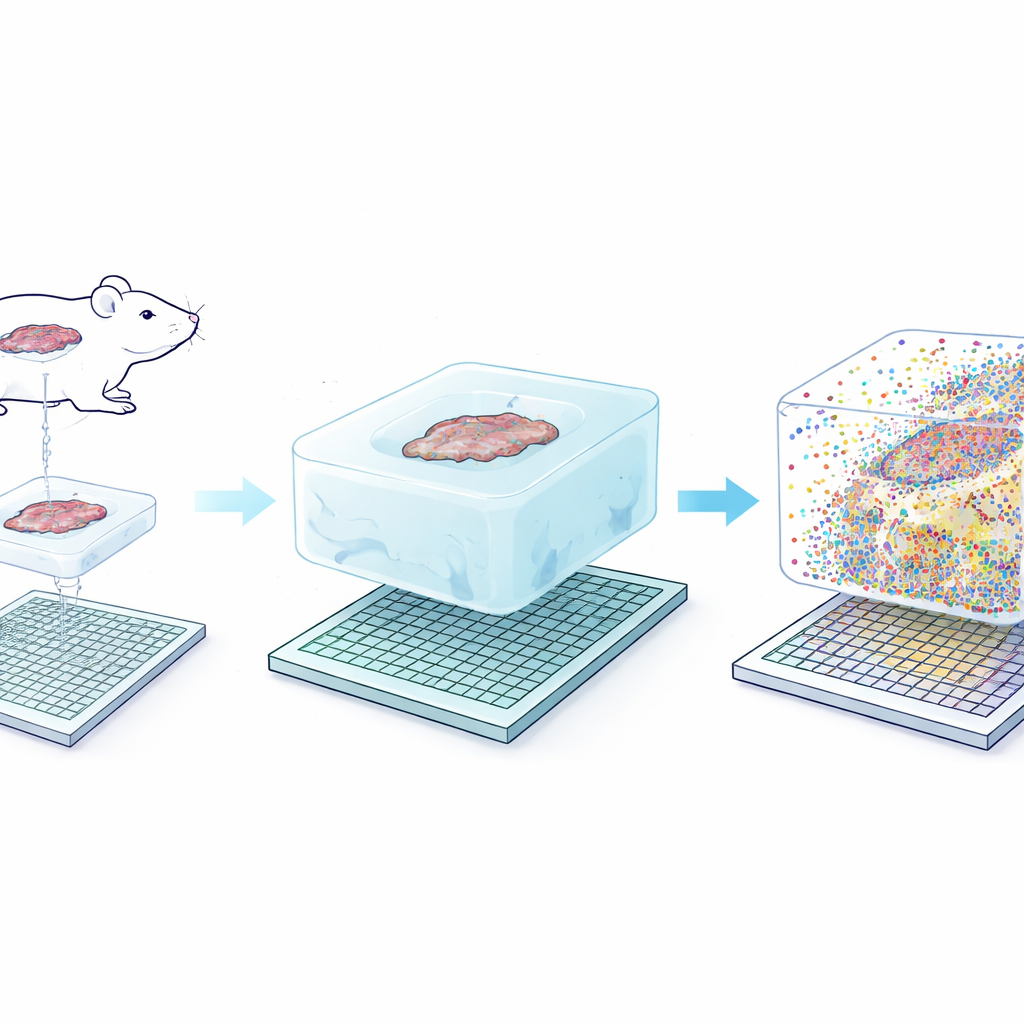
From Blurry Cells to Subcellular Maps
When the researchers applied Seq-Scope-X to mouse liver, they found that the method sharpened spatial resolution from roughly 0.6 micrometers to about 0.2 micrometers and increased the number of measurable positions per square millimeter by nearly tenfold. In practical terms, what used to look like fuzzy blobs of signal now resolved into crisp outlines of individual cells. Even more striking, clusters of RNA that had not yet been fully processed (unspliced) lined up tightly with the centers of nuclei, while mature RNA (spliced) traced the surrounding cytoplasm. This allowed the authors to draw cell boundaries using RNA alone and to separate nuclear and cytoplasmic gene readouts for nearly every hepatocyte in the tissue, something previous sequencing-based methods could only hint at.
Cells with Split Personalities
With this new clarity, the team discovered an unexpected form of molecular “double life” inside liver cells. Along the well-known gradient from blood vessels at the portal side of the liver lobule to those at the central side, hepatocytes specialize in different metabolic tasks. Seq-Scope-X showed that, in many cells, the pattern of genes in the nucleus did not match the pattern in the surrounding cytoplasm. Roughly one-third of hepatocytes appeared to have a nuclear profile aligned with one zone and a cytoplasmic profile aligned with a neighboring zone. Independent imaging-based methods, including MERFISH and single-molecule RNA fluorescence assays, confirmed that individual transcripts can be concentrated either in the nucleus or the cytoplasm. Together, these findings suggest that hepatocytes can dynamically shift their metabolic roles over time, with the nucleus preparing for a future state while the cytoplasm reflects the current one. 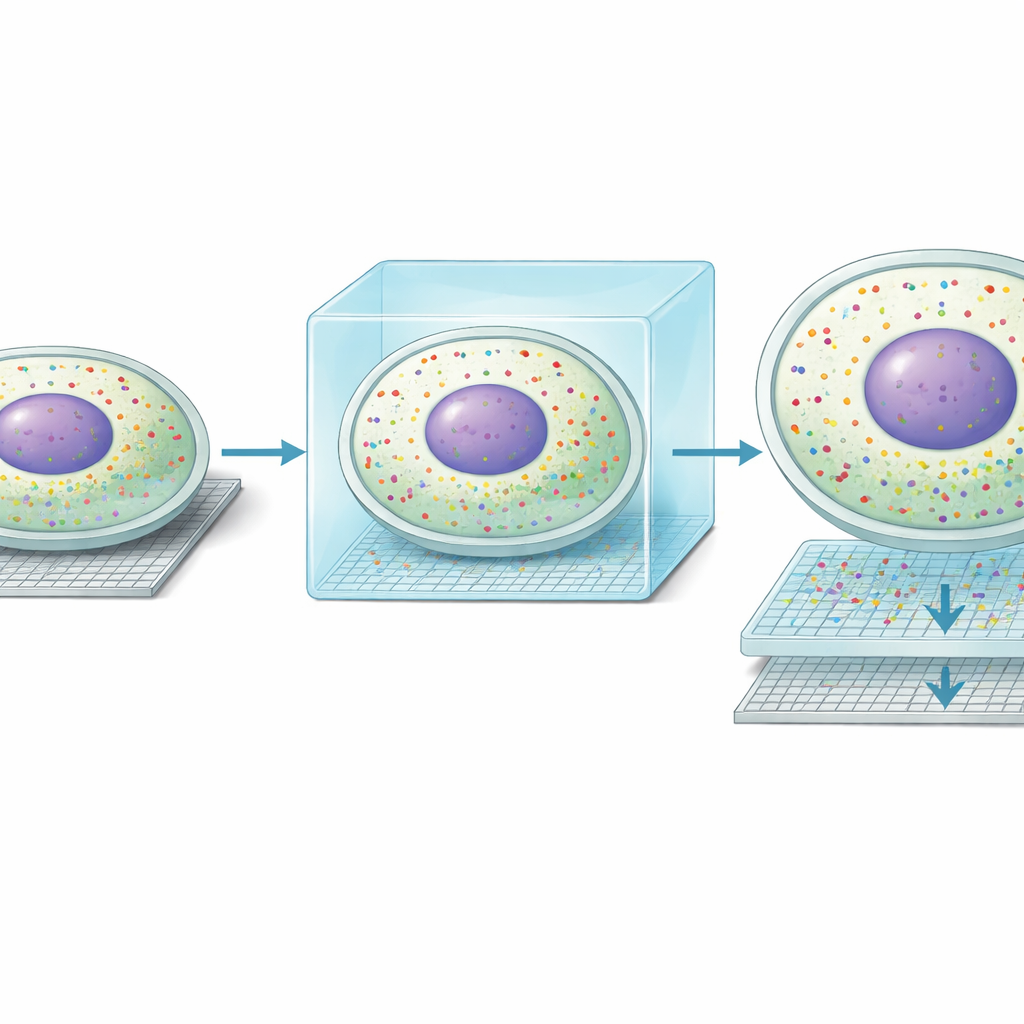
Beyond the Liver: Brain, Gut, and Immune Tissues
The authors then tested whether Seq-Scope-X works in other organs. In the brain, it cleanly separated different neuron types and supporting cells and revealed subcellular patterns of RNA around nuclei. In the colon, it resolved distinct goblet cell subtypes and colonocyte layers along the crypt-to-surface axis, again capturing nuclear signatures. Perhaps most dramatically, the team adapted the method to proteins by using DNA-barcoded antibodies in mouse spleen and human tonsil. Seq-Scope-X could simultaneously map over a hundred different cell-surface markers at single-cell resolution, distinguishing closely packed T and B cell subsets and myeloid cells. By switching to a more expandable gel chemistry, they pushed effective resolution toward the true nanoscale, while preserving tissue architecture well enough for detailed mapping.
What This Means Going Forward
Seq-Scope-X shows that physically enlarging tissues before sequencing can overcome long-standing resolution limits, delivering nearly microscope-level detail while still reading out thousands of genes or protein tags at once. For a lay observer, this means scientists can now build “molecular atlases” that not only say which cells are where, but also reveal what is happening inside different parts of each cell. Such maps could help explain how tissues respond to injury, how immune cells organize in lymphoid organs and tumors, or how subtle shifts inside cells foreshadow disease. Although further refinements are needed to scale up coverage and improve some staining panels, Seq-Scope-X opens a powerful new window on the microscopic landscapes that underlie health and illness.
Citation: Anacleto, A., Cheng, W., Feng, Q. et al. Seq-Scope-eXpanded: spatial omics beyond optical resolution. Nat Commun 17, 2564 (2026). https://doi.org/10.1038/s41467-026-69346-8
Keywords: spatial transcriptomics, tissue expansion, single-cell omics, spatial proteomics, liver zonation