Clear Sky Science · en
Benchmarking EGF signaling pathway inference using phosphoproteomics and kinase-substrate interactions
Why this matters beyond the lab
Our cells constantly listen and react to signals from their surroundings. One of the most important of these signals is epidermal growth factor (EGF), which helps control how cells grow, divide, and survive. When this communication system goes wrong, it can drive cancers and other diseases. This study asks a deceptively simple question with big implications: if we use today’s powerful measurement technologies and computer models, how much better can we really understand the EGF signaling pathway than what is shown in textbook diagrams?
From straight lines to real-life wiring diagrams
Traditional “signaling pathways” are often drawn as neat, almost linear chains of proteins passing a message from the cell surface to the nucleus. Those maps were built over decades using slow, targeted experiments that could only look at a few proteins at a time. The authors argue that this has left us with a narrow, streetlight-style view: we know a lot about the few proteins that were easy to study, and far less about the wider web of connections that likely exists in real cells.
Listening in on thousands of molecular switches at once
To move beyond that limited view, the team used phosphoproteomics, a mass-spectrometry technique that can track tens of thousands of tiny chemical tags called phosphate groups on proteins in a single experiment. These tags are placed by enzymes called kinases and act as fast on–off switches for signaling. The researchers first assembled the most comprehensive “who-can-phosphorylate-whom” maps to date by merging three types of information: curated findings from the literature, predictions from an artificial-intelligence model that reads protein sequences, and large experimental screens of kinase preferences. This greatly expanded the coverage of known or suspected kinase–substrate relationships from about 5% of measurable sites to as much as 70–80%.
Stress-testing EGF signaling across many experiments
Next, they focused on the EGF response, a classic pathway and major player in cancers of the lung, breast, skin, and brain. The group combined three state-of-the-art published datasets with three new, very deep time-resolved measurements in kidney-derived cells, plus two control studies where different stimuli were used. Together, these experiments measured over 40,000 phosphorylation sites per dataset and followed how they rose or fell in the minutes after EGF stimulation. While the overall patterns differed somewhat between cell types and technologies, key phosphorylation sites in the EGF pathway were repeatedly activated, confirming that the core signal is robust even across diverse experiments.
Reconstructing the hidden conversation between kinases
Having inferred which kinases became more or less active, the authors then tried to rebuild the underlying "conversation" among them: which kinases likely activate or influence which others. They converted their kinase–substrate maps into kinase–kinase networks and applied three computational strategies, ranging from simple ranking schemes to a more complex approach that forces the result to look like a directed tree rooted at the EGF receptor. They compared the resulting inferred pathways with several kinds of “ground truth”: a carefully curated EGF map from a database, large-scale experiments where single kinases were overexpressed, and pairs of kinase sites whose behavior was strongly correlated over time.
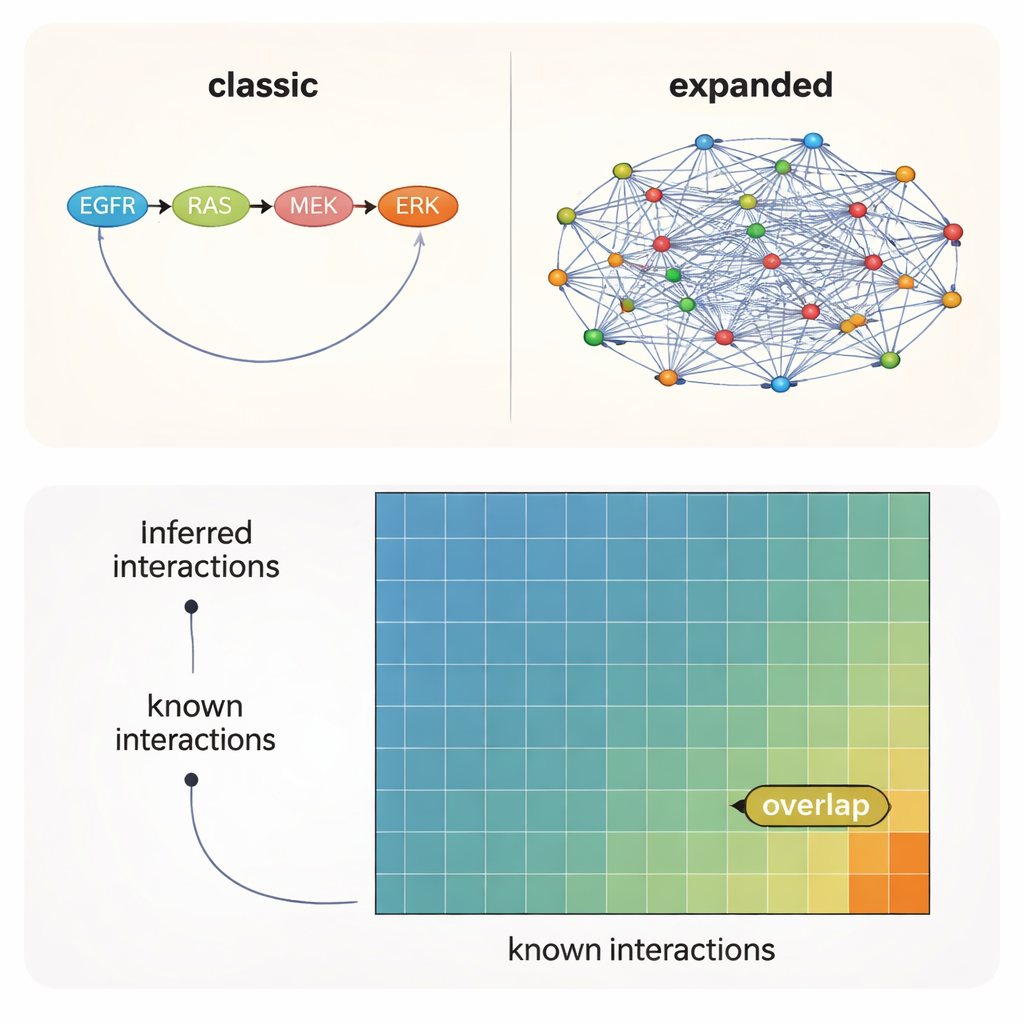
Most plausible links are still off the official map
Across all tests, one message was clear: the choice of prior knowledge—what starting network you feed into the algorithm—mattered more than the particular method or even the experimental context. Networks built from literature recovered the largest number of known interactions, while broader predictive resources pointed to many additional, plausible connections that have not yet been formally documented. Even under the best conditions, however, only about 10–12% of the interactions suggested by the data and models were present in any of the reference sets. In other words, roughly 90% of the supported kinase–kinase links sit in uncharted territory.
What this means for future treatments
For non-specialists, the takeaway is that the well-known EGF pathway diagram in textbooks is probably just the tip of the iceberg. Modern phosphoproteomics combined with advanced computational methods reveals a far richer and more tangled signaling network. Many of these newly suggested connections may represent overlooked ways that cancer cells reroute signals to escape drugs, or fresh opportunities for designing smarter drug combinations. The study does not prove every new link is real, but it provides a roadmap and open tools for researchers to test these ideas, ultimately aiming at more precise and effective therapies in diseases where EGF signaling plays a central role.
Citation: Garrido-Rodriguez, M., Potel, C., Burtscher, M.L. et al. Benchmarking EGF signaling pathway inference using phosphoproteomics and kinase-substrate interactions. Nat Commun 17, 2071 (2026). https://doi.org/10.1038/s41467-026-69332-0
Keywords: EGF signaling, phosphoproteomics, protein kinases, cell signaling networks, cancer biology