Clear Sky Science · en
A missense variant in ASCL5 leads to lobodontia
When Teeth Look Like a Predator’s
Lobodontia is an extremely rare condition in which human teeth take on a striking, almost carnivore-like appearance, with extra sharp bumps and unusual roots. Until now, scientists thought a calcium channel gene called CACNA1S was to blame, but the evidence was thin. This study revisits that story using modern genomic tools and animal experiments, and shows that a different gene, ASCL5, is the real driver—revealing how a tiny change in our DNA can reshape the architecture of our teeth and jaws.
Strange Teeth in Otherwise Healthy Families
The researchers examined 17 people from six Thai and Croatian families who shared the same unusual dental pattern. Their canines were elongated and fang-like, premolars had sharp, pointed ridges, and molars carried multiple extra cusps, reminiscent of carnivorous animals. X‑rays showed further quirks: folds of enamel diving into the tooth, enlarged pulp chambers, and lower molars with a single thick, pyramid-shaped root instead of the usual branching roots. Despite these dramatic changes in their mouths, all affected individuals were otherwise healthy and had normal development and intelligence. The condition appeared in every generation and in both sexes, pointing to a single dominant genetic change.
From a Suspect Gene to the Real Culprit
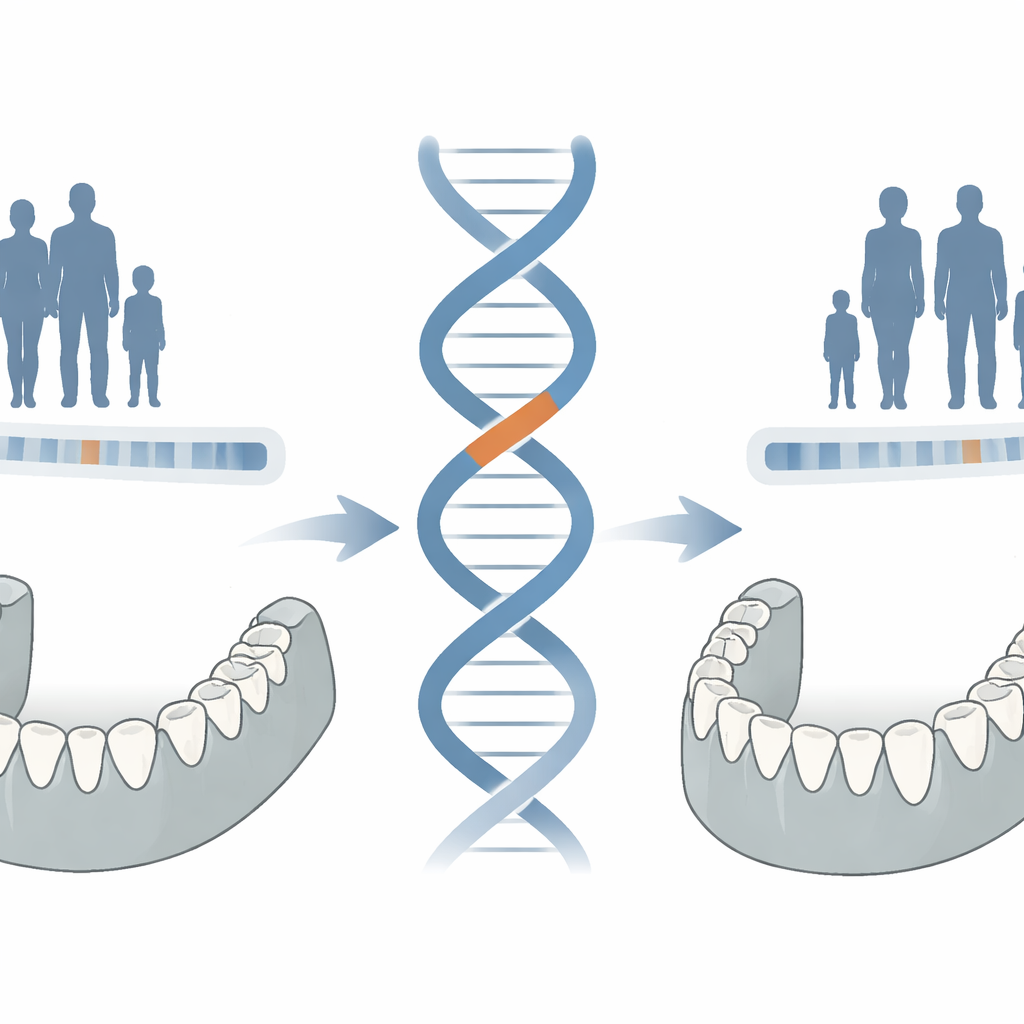
Earlier reports had linked lobodontia to a variant in CACNA1S, a gene better known for its role in muscle function. In this new study, the team found that all Thai patients carried that CACNA1S change, but Croatian patients with the same dental features did not. Even more telling, a healthy Thai person with perfectly normal teeth also carried the CACNA1S variant. That raised a red flag: perhaps this change was merely traveling together with the true cause on the same stretch of chromosome, rather than causing the condition itself. By combining whole‑genome sequencing with fine‑scale genetic mapping in the Thai families, the researchers narrowed the search to a 15.4‑million‑base region on chromosome 1 that contained both CACNA1S and a little‑studied gene, ASCL5.
A Single Letter Swap in a Tooth-Specific Gene
Within this region, genome sequencing uncovered a striking find: every affected family member, Thai and Croatian alike, carried the exact same change in ASCL5—a single DNA letter switch that alters one amino acid in the encoded protein. None of the unaffected relatives carried this change, and it was missing from large population databases, underscoring its rarity. ASCL5 is a transcription factor, a protein that turns other genes on or off, and its close relative in mice, called AmeloD, is known to be active in developing tooth enamel. Computer modeling suggested that the new amino acid could weaken how ASCL5 grips DNA, potentially altering its control over key developmental genes.
Mouse Clues: When Jaw and Tooth Plans Go Awry
To test whether this DNA change truly disrupts development, the team used CRISPR gene editing to introduce the equivalent mutation into mice. Animals with one altered copy of the gene—mirroring the human situation—grew extra cusps on their molars and showed root abnormalities, closely echoing lobodontia. Mice with two altered copies fared far worse: they had shortened lower jaws, missing or severely malformed molars, and overgrown front teeth, indicating that ASCL5 is crucial for normal jaw and tooth formation. When the researchers examined gene activity in the developing lower jaws, they found that several genes already known to shape facial bones and teeth, including members of the DLX family and the signaling molecule Shh, were dialed down in mutant embryos.
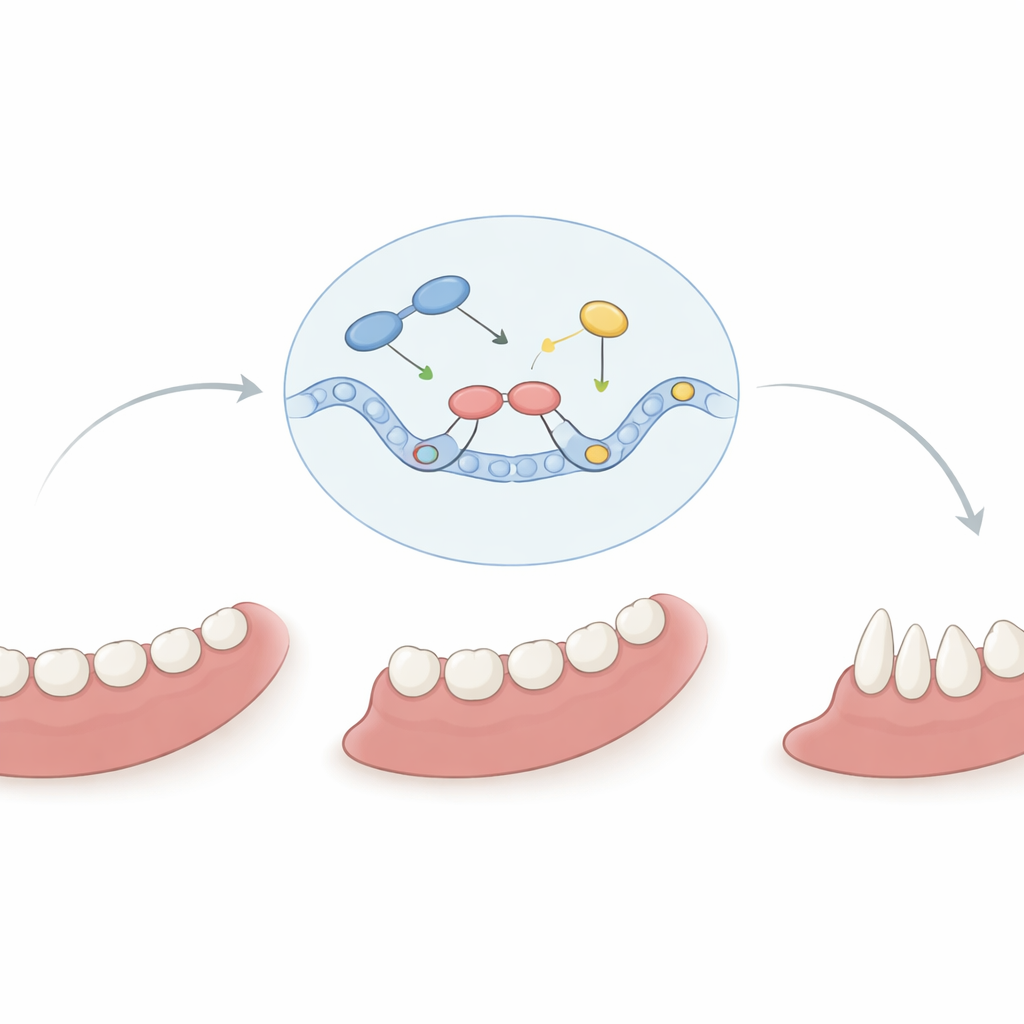
How One Faulty Switch Misguides Tooth Building
Because ASCL5 works by controlling other genes, the scientists asked whether the lobodontia‑linked version could still perform its normal tasks. In cell-based experiments, the healthy ASCL5 protein boosted activity of the DLX2 gene, a key player in craniofacial development, while the mutant version did this much less effectively. At the same time, both the normal and mutant proteins could still repress a cell‑adhesion gene called E‑cadherin, suggesting that the mutation selectively disrupts some targets but not others. In young mutant mouse molars, additional genes tied to hard tissue formation were abnormally activated, hinting that the variant may also misdirect mineralization of teeth. Together, these results paint a picture of ASCL5 as a finely tuned master switch: when one critical amino acid is altered, the timing and balance of signals that sculpt teeth and jaws shift, producing carnivore‑like crowns, odd roots, and in severe cases, missing teeth.
What This Means for Rare Tooth Disorders
By firmly linking lobodontia to a specific ASCL5 mutation and reproducing its effects in mice, this work overturns the earlier focus on CACNA1S and firmly establishes ASCL5 as a key regulator of how mammalian teeth and jaws take shape. For families with lobodontia, it provides a clear genetic explanation and a basis for future diagnosis. More broadly, it shows how a subtle change in a developmental “control knob” can reorganize the form of our teeth without affecting the rest of the body, offering new insight into both rare dental conditions and the evolutionary flexibility of our smiles.
Citation: Theerapanon, T., Intarak, N., Rattanapornsompong, K. et al. A missense variant in ASCL5 leads to lobodontia. Nat Commun 17, 2643 (2026). https://doi.org/10.1038/s41467-026-69323-1
Keywords: lobodontia, ASCL5 gene, tooth development, craniofacial genetics, dental anomalies