Clear Sky Science · en
Discovery of antimicrobial peptides targeting Acinetobacter baumannii via a pre-trained and fine-tuned few-shot learning-based pipeline
Why this matters for everyday health
Drug‑resistant infections are rising so quickly that, by mid‑century, they could kill more people each year than cancer. One of the most worrisome culprits is Acinetobacter baumannii, a tough hospital germ that shrugs off many antibiotics and often infects people on ventilators. This study shows how scientists combined artificial intelligence with lab experiments to rapidly discover new, tiny drug candidates that can kill this bacterium while causing far less harm to the body than our current last‑resort medicine.
A hospital superbug that is hard to kill
Acinetobacter baumannii is a Gram‑negative bacterium that thrives in intensive‑care units, particularly in patients relying on ventilators. Its rugged outer shell and fast‑evolving genome make it resistant to many standard antibiotics. Today, one of the few remaining options is a drug called polymyxin B, but it can severely damage the kidneys and the bacterium can still develop resistance. At the same time, a promising class of small protein fragments called antimicrobial peptides has been explored against other microbes, yet very few have been found that specifically work well on A. baumannii. Traditional trial‑and‑error screening is simply too slow and costly to sift through the astronomically large number of possible short peptide sequences.
Using smart algorithms to search a huge space
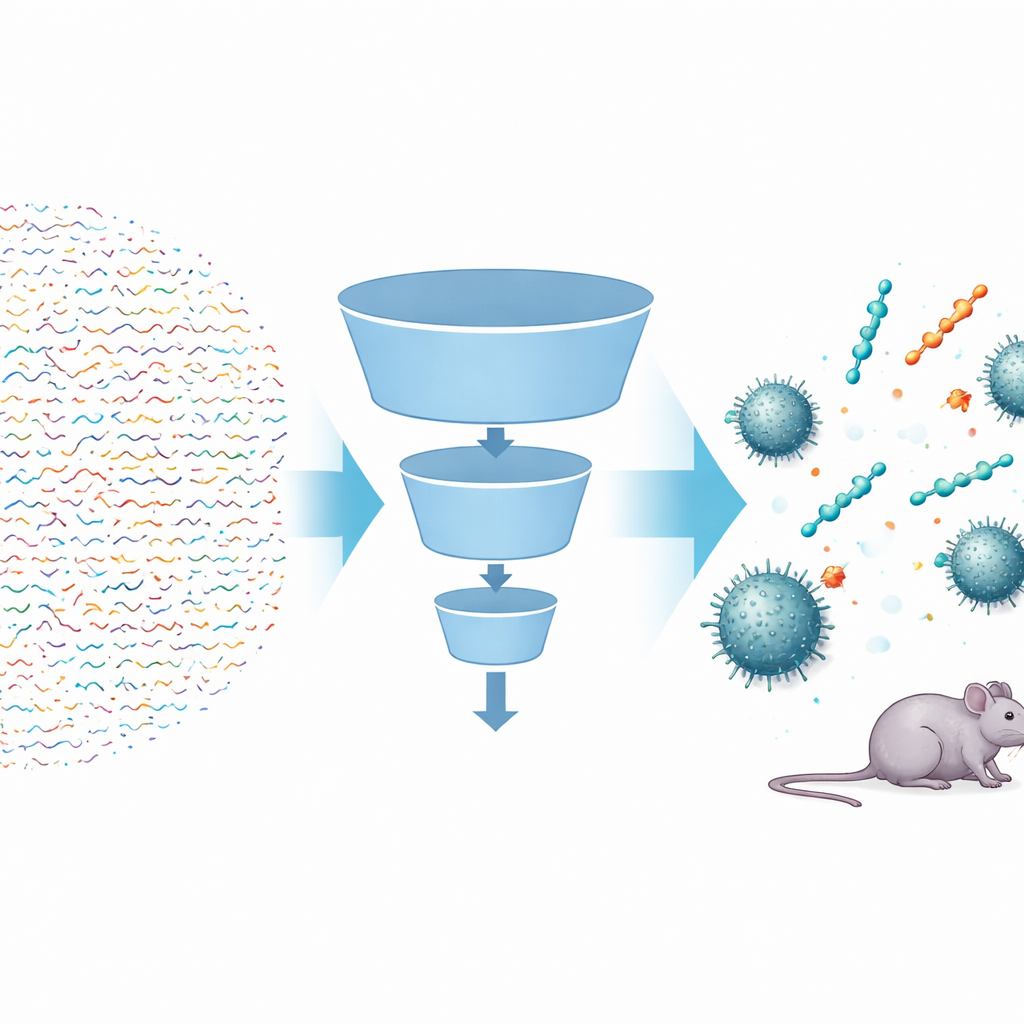
The researchers built a multi‑step artificial intelligence pipeline, called FSLSMEP, to scan entire libraries of short peptides—those with six, seven, or eight building blocks—amounting to tens of billions of candidates. The challenge was that they had only 148 known peptides that work against A. baumannii, far too few for standard machine‑learning methods. To get around this, they started with a powerful pre‑trained model that had already “read” hundreds of millions of natural protein sequences and learned general rules about how such molecules behave. They then fine‑tuned this model in two stages: first on a larger set of peptides active against a related bacterium, Pseudomonas aeruginosa, and finally on the scarce A. baumannii data. Along the way, three linked modules—one to sort likely from unlikely peptides, one to rank them, and one to estimate how strong each might be—acted like successive filters in a funnel.
From computer predictions to test‑tube successes
After first discarding the least promising peptides using simple chemical rules, the team fed nearly four million candidates into their pipeline. The classifier weeded out most sequences that did not look antimicrobial; the ranking model favored those whose features suggested strong activity; and the regression model predicted the minimum drug concentration needed to stop bacterial growth. From sixty‑four million six‑unit peptides, the system proposed just ten top candidates. When synthesized and tested in the lab, nine of these showed real antibacterial activity, an impressive hit rate in drug discovery. The same trained models were then applied, without retraining, to much larger seven‑ and eight‑unit peptide spaces, from which the researchers chose additional top‑ranked sequences for testing. Again, most turned out to be active, including heptapeptides as potent as polymyxin B against some strains.
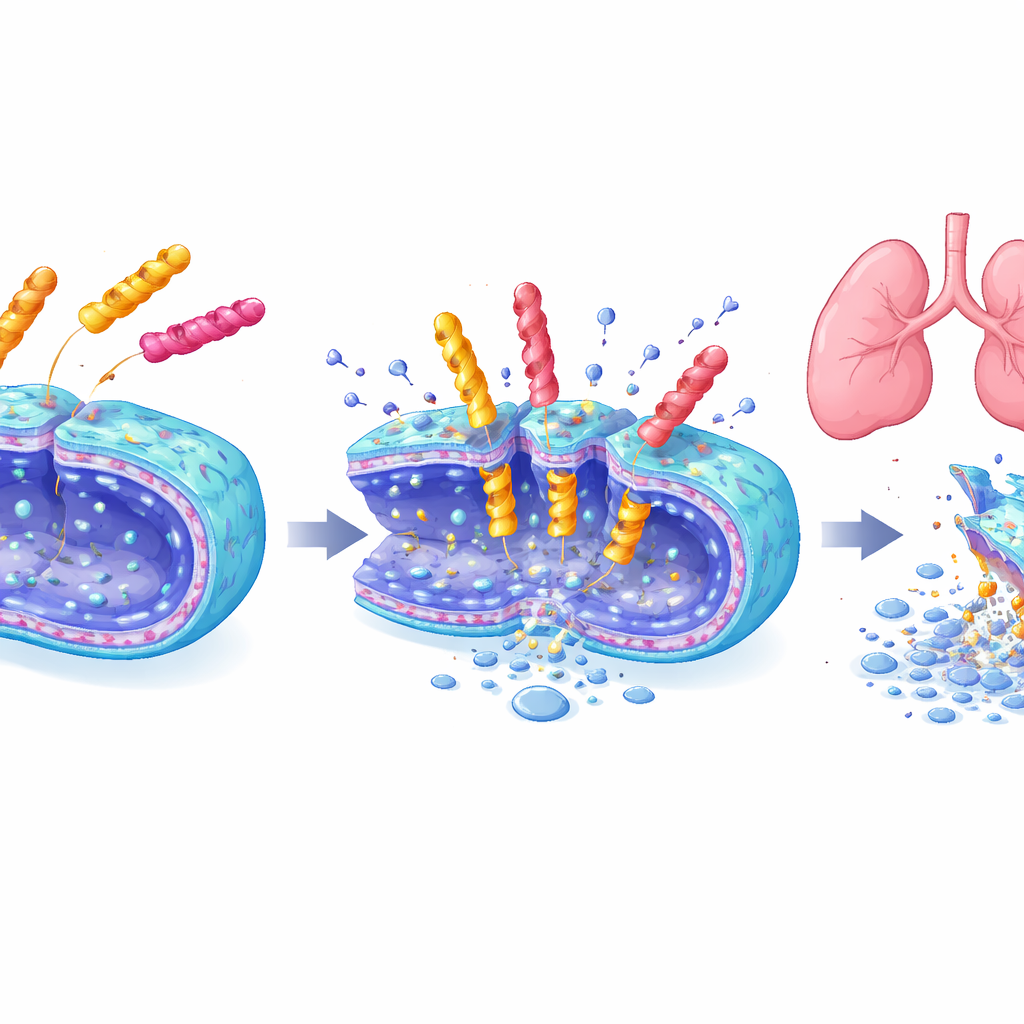
Potent killers with a gentler touch
Three lead peptides—one each from the six‑, seven‑, and eight‑unit libraries—were examined in detail. They rapidly wiped out A. baumannii populations in minutes to hours, damaged bacterial membranes as seen under electron microscopes, and also worked on several other dangerous microbes. At the same time, they caused little harm to cultured mammalian cells or to red blood cells, in contrast to some existing antimicrobial peptides. In long‑term exposure tests, bacteria did not readily develop resistance to these new molecules, while resistance to polymyxin B rose quickly. Most strikingly, in a mouse model of pneumonia, an inhaled dose of the heptapeptide EME7(7) cleared lung infections as effectively as polymyxin B but did not injure the kidneys, whereas polymyxin B caused clear kidney damage and increases in blood markers of renal stress.
What this means for future medicines
This work shows that a carefully designed AI pipeline can turn a tiny, incomplete dataset into a powerful engine for discovering new antimicrobial peptides. By combining broad pre‑training, stepwise fine‑tuning, and multiple layers of filtering, the researchers efficiently explored whole libraries of short peptides and found candidates that both fight a dangerous hospital superbug and appear safer for vital organs. The same strategy also succeeded in finding peptides active against the fungal pathogen Candida albicans, hinting that it could be reused to search for many other types of therapeutic peptides. For patients, this approach could eventually translate into new medicines that treat stubborn infections without the heavy side effects and rapid resistance that plague today’s last‑line antibiotics.
Citation: Huang, J., Zhang, W., Wang, A. et al. Discovery of antimicrobial peptides targeting Acinetobacter baumannii via a pre-trained and fine-tuned few-shot learning-based pipeline. Nat Commun 17, 2475 (2026). https://doi.org/10.1038/s41467-026-69306-2
Keywords: antimicrobial peptides, antibiotic resistance, Acinetobacter baumannii, machine learning drug discovery, few-shot learning