Clear Sky Science · en
Utilizing the predictable binding kinetics of DNA-PAINT to denoise super-resolution images
Sharper Views into the Tiny World
Modern microscopes can now “see” individual molecules inside cells, but these images are often speckled with misleading background dots. This study introduces a way to clean up such images by taking advantage of how pieces of DNA naturally stick together and let go over time. The result is crisper pictures of the molecular machinery inside cells, which matters for everything from basic biology to drug discovery.
How a Blinking Trick Reveals Hidden Detail
One powerful imaging method, called DNA-PAINT, turns the random sticking of short DNA strands into a super‑resolution microscope. One DNA strand is attached to the protein a scientist wants to see; a matching strand carrying a fluorescent dye floats in the solution. When the floating strand briefly binds to its partner, it produces a tiny flash of light. By recording many such blinks, and pinpointing their locations, researchers can reconstruct the positions of proteins with nanometer precision—far sharper than conventional light microscopes.
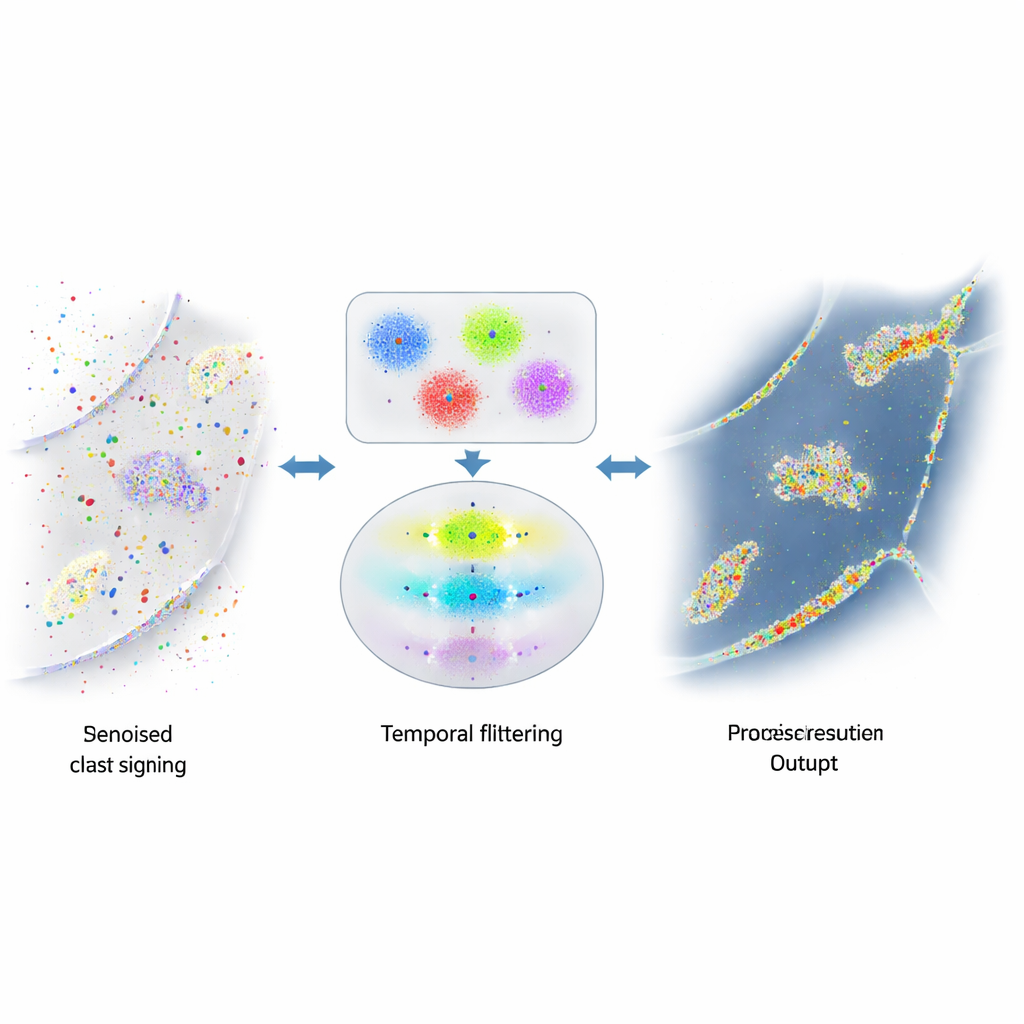
When Helpful Signals Are Buried in Noise
DNA-PAINT’s strength—lots of freely moving fluorescent strands—also creates a problem. These strands occasionally stick where they should not, for example to random cellular surfaces or to the abundant DNA inside the nucleus. Those accidental flashes are indistinguishable in appearance from the real ones and clutter the final image with false dots and clusters. Earlier attempts to reduce this noise focused on changing the chemistry to make unwanted sticking rarer, or on rough rules to spot obvious outliers. Yet a substantial amount of misleading signal remained, limiting how reliably scientists could measure protein numbers or spatial relationships.
Using Time to Tell Truth from Trickery
The authors realized that real DNA pairs in DNA-PAINT obey a simple timing rule: the pauses between genuine binding events follow a predictable, exponential pattern. In contrast, random sticking events do not. They built a multi‑step algorithm that first groups nearby blinks into small clusters based on where they occur, using a combination of density‑based and k‑means clustering to break complex shapes into localization‑sized units. For each cluster, they then construct a “time trace” of binding events, carefully merging rapid on‑off blinking from the same binding into single events. Finally, they apply a statistical test to ask whether the gaps between events in each cluster match the expected exponential behavior. Clusters that pass are treated as real; those that fail are discarded as noise.
Putting the Method to the Test in Real Cells
To test their approach, the team imaged the cell‑adhesion protein E‑cadherin in fruit fly egg chambers, where some cells produced the tagged protein and neighboring cells did not. This created side‑by‑side areas rich in genuine signal and areas that should contain only background. By tuning how many binding events a cluster must show before being tested, they found settings that removed over 90% of spurious clusters while keeping more than 98% of true ones at cell borders. In denoised images, fine features such as thin tubular structures and vesicles became much clearer. The same strategy worked on other systems, including mitochondria and microtubules, and even on proteins with more diffuse, amorphous distributions, where spatial patterns alone give no clue about what is real.
Clearer Molecular Maps for Future Studies
By listening to the “rhythm” of DNA binding rather than just looking at where blinks occur, this method reliably strips away misleading background from DNA‑PAINT data. For nonspecialists, the key outcome is simple: sharper, more trustworthy molecular maps inside cells. This makes it safer to count how many protein molecules are present, to judge how closely different proteins sit next to each other, and to build more accurate pictures of the inner life of cells. As DNA‑based imaging continues to grow, this kind of smart denoising will be essential for turning raw speckled images into solid biological insight.
Citation: Sirinakis, G., Allgeyer, E.S., Richens, J.H. et al. Utilizing the predictable binding kinetics of DNA-PAINT to denoise super-resolution images. Nat Commun 17, 2397 (2026). https://doi.org/10.1038/s41467-026-69304-4
Keywords: DNA-PAINT, super-resolution microscopy, image denoising, single-molecule imaging, protein mapping