Clear Sky Science · en
Assessing conformation validity and rationality of deep learning-generated 3D molecules
Why AI-Designed Molecules Need a Reality Check
Artificial intelligence is rapidly learning to design small, three-dimensional molecules that can nestle into the nooks and crannies of disease-related proteins. These AI-designed structures could one day accelerate drug discovery. But there is a catch: many of the computer-generated molecules look fine on screen yet break the basic rules of chemistry. They may twist into impossible shapes or pack atoms so tightly that they would never exist in real life. This study introduces a fast, physics-aware quality control system to tell which AI molecules are likely to be real—and which ones belong in the digital trash bin.
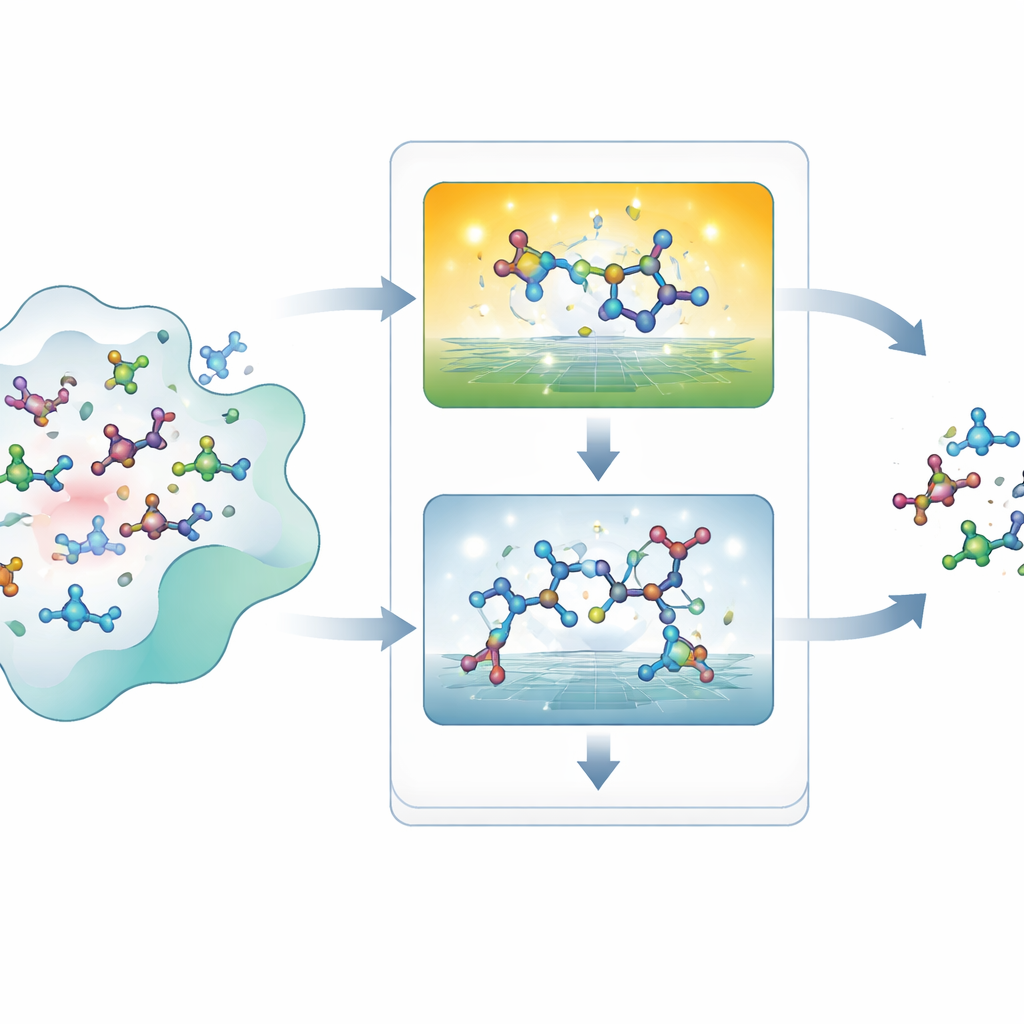
When Pretty Pictures Hide Impossible Shapes
Modern AI systems can propose thousands of 3D molecules for a given protein pocket, but checking whether each proposal is physically reasonable is surprisingly hard. Traditional “geometry checks” look at bond lengths, bond angles, and how closely atoms approach each other, or compare shapes to known reference structures. These rules can miss many subtle problems and can give misleading answers when a new molecule is unlike anything in the reference set. More rigorous energy calculations from quantum mechanics are far more trustworthy but painfully slow, making them impractical for screening millions of candidates. As a result, developers of generative models have lacked a clear, scalable way to measure whether their creations obey basic chemical physics.
A Two-Step Health Check for 3D Molecules
The authors propose a two-stage framework that combines the speed of machine learning with the accuracy of advanced quantum chemistry. The first stage, called the “validity test,” targets grossly unrealistic structures before any clean-up is done. It uses a machine-learning force field to estimate the energy of each atom in a molecule based on its local surroundings. Atoms that sit in extremely high-energy environments—such as severe clashes, twisted rings, or misplaced hydrogens—raise red flags. This module, named HEAD (high-energy atom detector), labels conformations as valid or invalid and can also flag problematic contacts between a molecule and its protein pocket.
From Rough Sketches to Chemically Sensible Poses
Even if a molecule passes this first filter, it might still strain its internal “hinges”—the rotatable bonds—into awkward angles. After a quick clean-up with a classical force field, the second stage, called the “rationality test,” examines these finer details. Here the TED (torsional energy descriptor) tool breaks a molecule into fragments around each rotatable bond and uses a deep learning model trained on millions of quantum-level calculations to predict how costly each twist is in terms of energy. If any bond sits in a state more than about 2 kilocalories per mole above its preferred range, the conformation is labeled irrational. TED focuses on these local torsional strains, which medicinal chemists care about because they often correlate with unstable or hard-to-make molecules.
Putting AI Molecule Generators Under the Microscope
To demonstrate the power of their approach, the researchers used HEAD and TED to scrutinize five state-of-the-art AI models that generate 3D molecules for 102 different protein targets. They first filtered out molecules that were unlikely to be useful drugs based on standard “drug-likeness” and synthetic accessibility scores. The remaining candidates were then passed through HEAD to check both ligand shapes and their fit inside protein pockets, and through TED to probe torsional strain after refinement. No single AI model excelled at everything: some produced molecules that interacted well with protein pockets but often had strained internal geometries, while others yielded more torsion-friendly structures but more frequent clashes. This side-by-side evaluation revealed distinct strengths and weaknesses that would not be obvious from simple docking scores or geometry checks alone.
A Practical Screening Pipeline for Future Drug Design
By chaining together drug-likeness filters, HEAD validity checks, and TED rationality checks, the authors built a full screening pipeline that can process thousands of AI-generated molecules in minutes on modern hardware. In this pipeline, only about one in five molecules from the best-performing models survived all stages, underscoring how much “fantasy chemistry” current generators still produce. Yet the framework is flexible: HEAD can plug into newer machine-learning force fields that support more elements, and TED can be improved with richer data and environmental information. For non-experts, the takeaway is straightforward: this work supplies a fast, physics-based safety net that helps separate chemically plausible AI-designed molecules from the many that would fall apart outside a computer, bringing AI-driven drug design a step closer to trustworthy reality.
Citation: Fan, F., Xi, B., Meng, X. et al. Assessing conformation validity and rationality of deep learning-generated 3D molecules. Nat Commun 17, 2481 (2026). https://doi.org/10.1038/s41467-026-69303-5
Keywords: AI-driven drug design, 3D molecular conformation, machine learning force fields, torsion energy, structure-based drug discovery