Clear Sky Science · en
A role for condensin-mediator interaction in mitotic chromosome organization
How dividing cells keep their genetic house in order
Every time a cell divides, it must carefully pack and separate meters of DNA so that each daughter cell gets a complete set of chromosomes. When this process fails, the result can be cell death, developmental disorders, or cancer. This study uncovers how two major molecular machines—one that controls gene activity and another that compacts DNA—work together to fold chromosomes during cell division, revealing a hidden link between which genes switch on in mitosis and whether chromosomes are shared faithfully.
Key players: the packer and the coordinator
Inside the nucleus, a protein complex called condensin acts as a molecular packer, looping and tightening DNA into the thick chromosomes seen during mitosis. Another large complex, called mediator, helps control which genes are turned on or off by working with RNA polymerase, the enzyme that copies DNA into RNA. The authors studied these complexes in fission yeast, a well-established model organism whose chromosome biology is strikingly similar to that of human cells. They discovered that one subunit of condensin, Cnd1, physically binds to a mediator subunit called Pmc4. This interaction happens at highly active genes and at a special set of genes that turn on only during mitosis. These “mitotic genes” sit at the edges of large chromosomal neighborhoods, or domains, and seem to act as boundary markers.
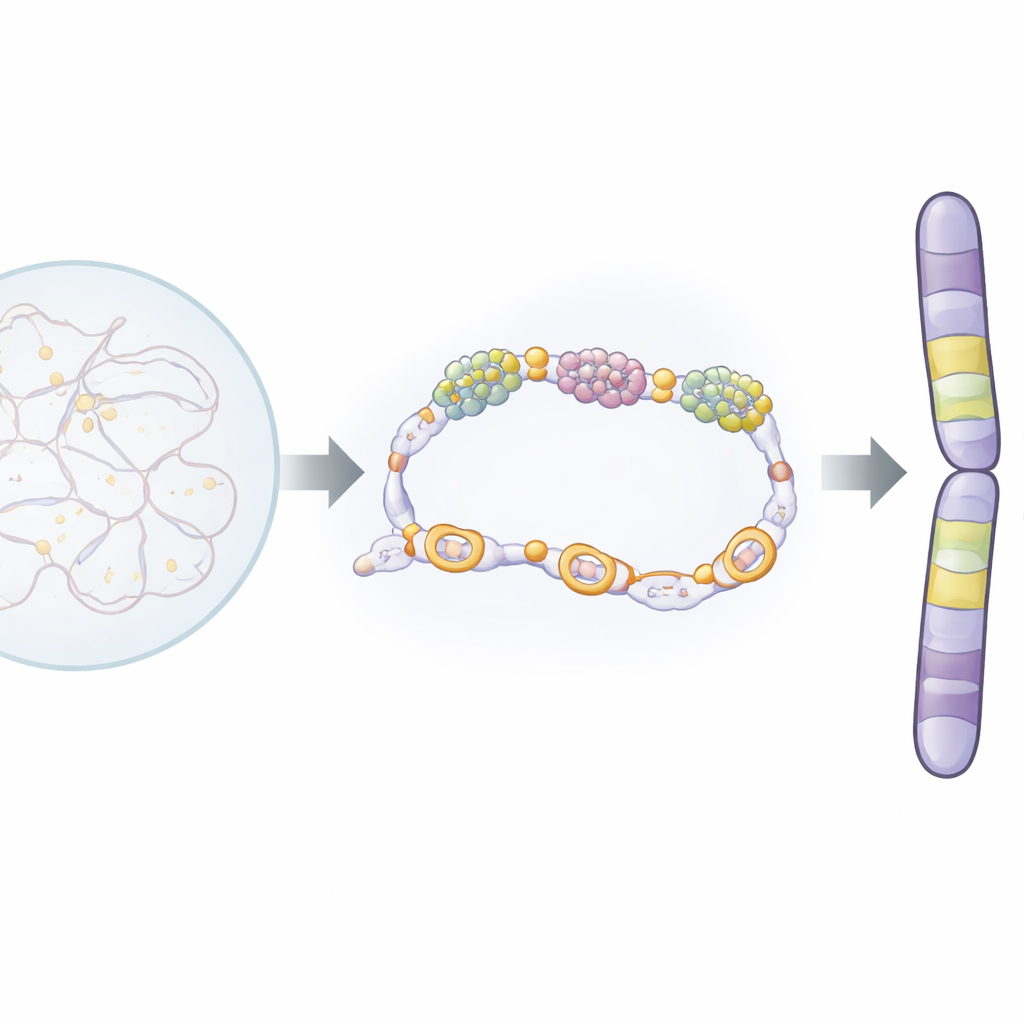
A single molecular handshake with big consequences
To test how important this handshake is, the researchers engineered a very precise mutation in Cnd1, changing just one amino acid (K658E) so that Cnd1 can no longer bind Pmc4 while still forming a normal condensin complex. Yeast cells carrying this mutation were viable, but their chromosomes missegregated more often, leaving lagging pieces of DNA during mitosis. Using 3D genome-mapping (Hi-C) and microscopic distance measurements between selected DNA spots, the team showed that the mutant cells had weaker condensin-driven contacts and less compact chromosomal domains. In other words, without a proper Cnd1–Pmc4 interaction, condensin could not efficiently fold chromosomes into robust, domain-based structures during cell division.
Gene activity as a boundary builder
The study then asked how mediator contributes to this folding. When Pmc4 was depleted, condensin binding at many genes dropped, and the strength of condensin-made domains decreased. Strikingly, the borders between neighboring domains became fuzzy, and DNA interactions began to spill across what used to be sharp boundaries. Detailed RNA measurements revealed that Pmc4 loss strongly reduced expression of a subset of mitotically activated genes controlled by the Ace2 transcription factor. These boundary genes normally show very high occupancy of mediator, condensin, and a basic transcription factor called TBP. The data support a model in which mediator and TBP first recruit condensin to promoters of highly active and mitotic genes; as transcription proceeds, condensin is pushed along gene bodies and helps knit nearby DNA into loops, with the Ace2 target genes setting the edges of each condensed domain.
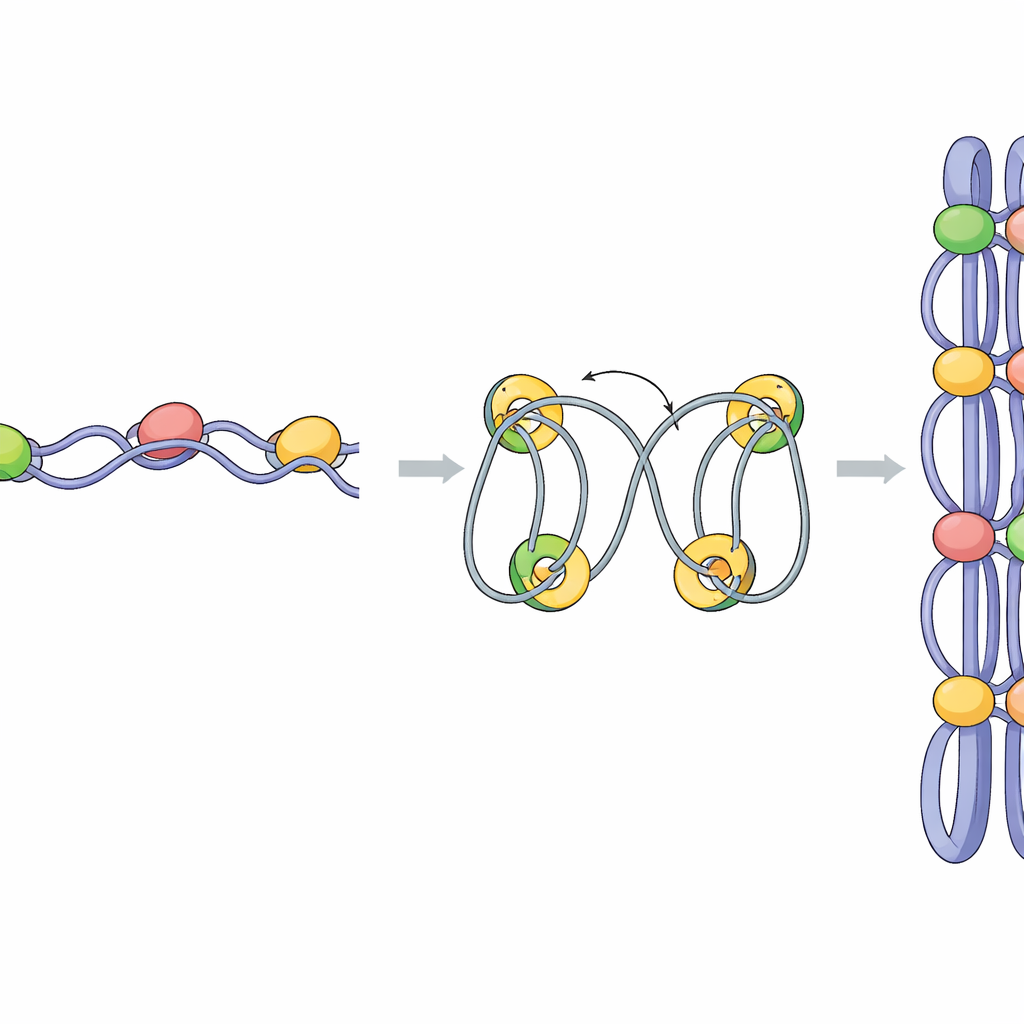
A possible role for liquid-like droplets
Mediator is known in human cells to form liquid-like droplets through a process called phase separation, which can concentrate transcription machinery in small nuclear hubs. The authors found that the fission yeast mediator behaves similarly: the Pmc4 subunit can form droplets in a test tube, and treatment of cells with 1,6-hexanediol, a chemical that disrupts such condensates, rapidly dispersed mediator foci in the nucleus. Mild hexanediol treatment weakened mediator and condensin binding at key genes and specifically blurred the boundaries between chromosomal domains, even though overall compaction of chromosomes still proceeded. This suggests that mediator-rich droplets at mitotic boundary genes may help shape where domains begin and end by focusing transcription and condensin loading at those sites.
From yeast lessons to human health
Finally, the team asked whether a similar partnership exists in human cells. Using an interaction assay, they found that MED4, the human counterpart of Pmc4, binds specifically to CAP-D3, a condensin II subunit closely related to yeast Cnd1. A human CAP-D3 mutation equivalent to the yeast K658E change disrupted this interaction and caused mitotic errors, such as misaligned chromosomes and micronuclei. Depleting MED4 also produced segregation defects. These parallels suggest that coupling between mediator and condensin is a conserved strategy by which cells coordinate gene activity with chromosome folding, ensuring that DNA is neatly packaged and evenly divided during cell division.
Why this matters for understanding disease
This work reveals that the way chromosomes are folded during mitosis is not just a matter of brute-force compaction. Instead, it depends on where and when particular genes are switched on, with mediator-driven transcription at boundary genes recruiting condensin to sculpt large-scale domains. By showing how a single interaction between two complexes can ripple out to affect whole-chromosome architecture and segregation fidelity, the study offers a framework for thinking about how subtle changes in gene regulation machinery or condensin function might lead to chromosomal instability, a hallmark of many cancers and genetic diseases.
Citation: Iwasaki, O., Tashiro, S., Chung, C.YL. et al. A role for condensin-mediator interaction in mitotic chromosome organization. Nat Commun 17, 2509 (2026). https://doi.org/10.1038/s41467-026-69270-x
Keywords: chromosome architecture, condensin, mediator complex, mitosis, 3D genome organization