Clear Sky Science · en
Interpretable and generative deep learning models explicate phase separating intrinsically disordered motifs
Why tiny protein segments matter
Inside each of our cells, vital molecules often gather into droplet-like blobs called biomolecular condensates. These droplets help organize chemistry without the walls of a membrane, shaping how genes are turned on, how signals are passed, and how cells respond to stress. Many such droplets are formed by floppy stretches of proteins known as intrinsically disordered regions. Yet biologists still struggle to pinpoint the short pieces of sequence that actually make these droplets form. This study introduces a deep learning framework, PhaSeMotif, that can both find these key segments and design new ones, giving researchers a powerful new way to probe and rewire cellular droplets.
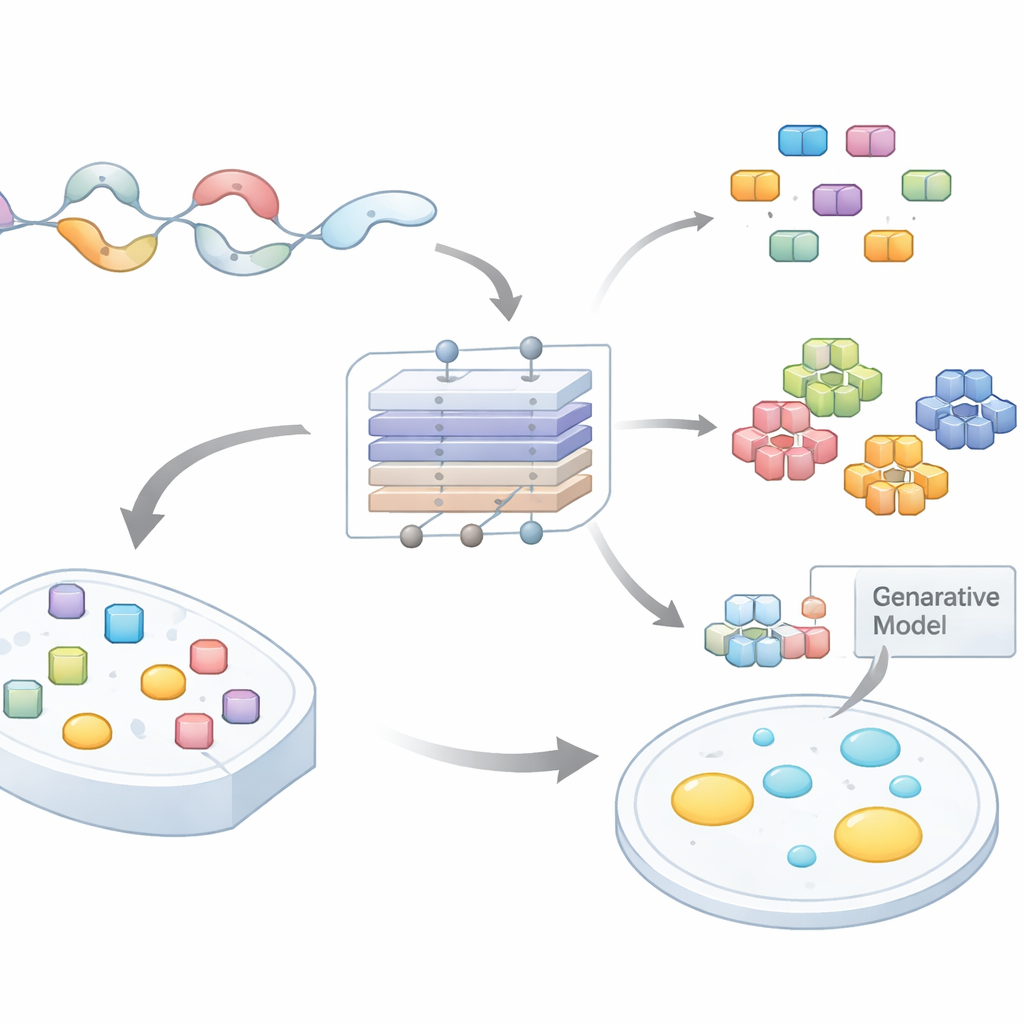
From messy protein tails to testable ideas
Many proteins contain long, flexible tails that do not fold into fixed shapes. These disordered regions are enriched in certain amino acids and often harbor repeated patterns or short motifs. A growing body of work shows that such motifs drive condensation by enabling many weak interactions at once. However, scanning entire proteomes to find which short stretches matter, and why, has been a major bottleneck. Existing computational tools usually rate whole proteins or large regions, offering little guidance on where to mutate or what to test in the lab. The authors set out to build a model that predicts not only whether a disordered region can form droplets, but also which exact subsequences are doing the heavy lifting.
A deep learning map of droplet-driving motifs
The team compiled large datasets of disordered regions across several species and labeled them according to whether their host proteins were likely to undergo phase separation. They then trained an attention-based neural network, PhaSeMotif, that takes an amino acid sequence of any length and outputs a droplet-forming score. Crucially, the network uses a combination of convolutional layers and attention mechanisms to evaluate how much each short window of the sequence contributes to that score. By tracing back through the model (using techniques akin to guided backpropagation), the authors extracted high-importance patches—short motifs often less than 20 residues long—that the model deemed essential for droplet formation.
Putting predictions to the test in living cells
To see whether these motifs really mattered, the researchers turned to a light-activated system in human cells. They fused predicted droplet-forming disordered regions to a light-sensitive oligomerization module and a fluorescent tag. Under blue light, these constructs rapidly condensed into bright puncta, reporting phase separation in real time. The team then surgically removed individual motifs by replacing them with neutral, flexible linkers of the same length. In 82% of the 17 altered sequences tested, droplet formation was dramatically weakened or vanished altogether, while control mutations outside PhaSeMotif segments often had little effect. Importantly, many of these key motifs overlapped with sites where disease-linked mutations are known to disrupt condensation, underscoring their biological relevance.
Uncovering a vocabulary of motif types
With more than 17,000 motifs in hand, the authors next asked whether there were common "flavors" of droplet-driving segments. They analyzed amino acid composition and patterning, then clustered motifs into nine groups. Some clusters were rich in aromatic residues and glycine, consistent with sticky π–π and cation–π interactions. Others contained separated patches of positive and negative charges, favoring electrostatic attraction and selective partitioning into particular condensates. Additional clusters were dominated by proline and glycine, which support flexibility, or by long runs of glutamine that can form dense networks of hydrogen bonds. Different cell compartments and condensate types showed characteristic mixes of these motif classes, hinting that motif composition helps determine where and with what partners a protein will condense.
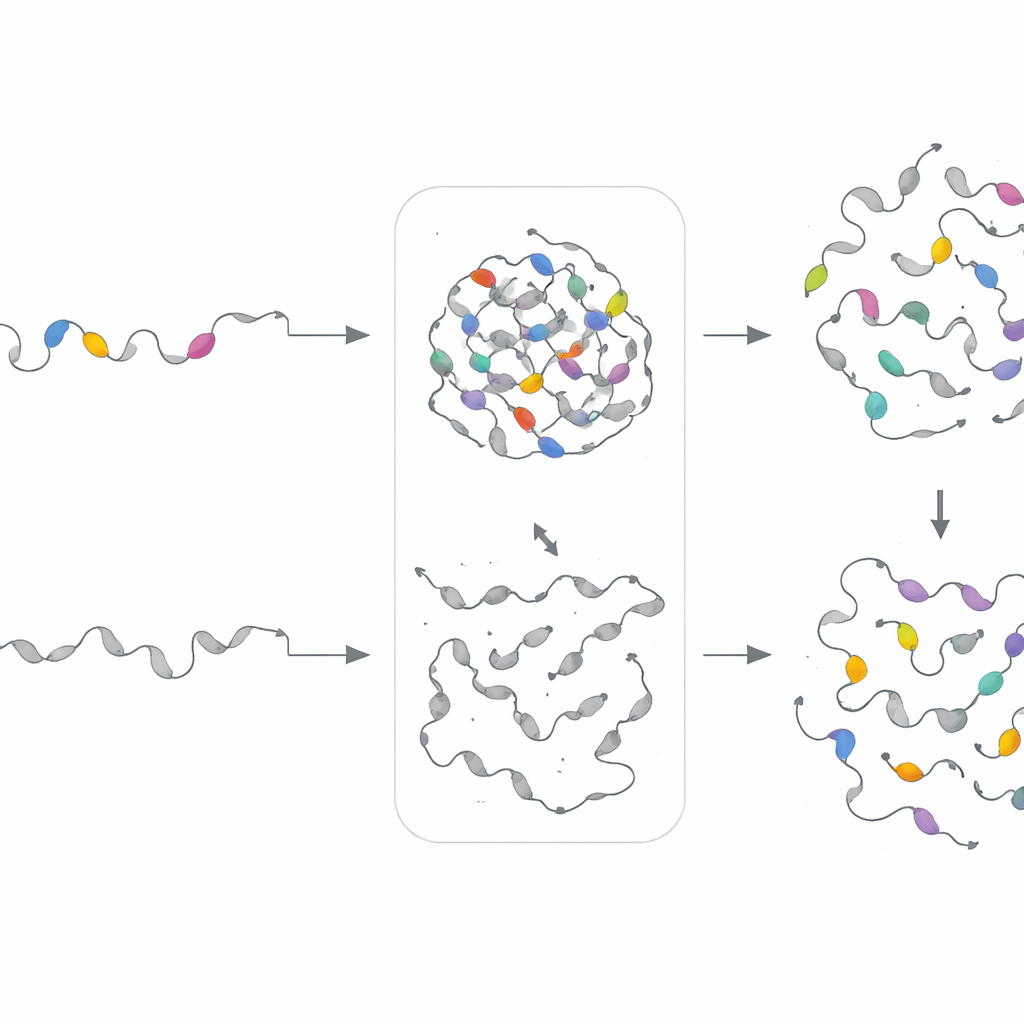
Designing new motifs to prove the rules
To test whether motif "recipes"—rather than exact sequences—govern droplet behavior, the team built separate generative models for each motif cluster. These variational autoencoders learned the statistical patterns for a given cluster and then produced new, artificial sequences sharing the same compositional fingerprints but different exact order. The researchers experimentally swapped these synthetic motifs into proteins where the original segments had been deleted. Remarkably, in 18 of 21 cases, the engineered motifs restored phase separation in cells, sometimes even tuning the speed or density of droplet formation. This shows that PhaSeMotif captures underlying design rules that can be reused to build or repair droplet-forming regions.
What this means for biology and disease
By linking interpretable deep learning with generative design and direct cellular tests, this work turns the vague notion of "disordered droplet-forming regions" into a concrete set of short, composable motifs. For non-experts, the takeaway is that scientists can now read and write the tiny protein segments that control how cellular droplets assemble, mix, and malfunction. This opens the door to faster discovery of disease-causing mutations in these segments, clearer mechanistic studies of how condensates organize cell physiology, and eventually the rational engineering of proteins that steer droplets for therapeutic or synthetic biology applications.
Citation: Yang, H., You, K., Ma, L. et al. Interpretable and generative deep learning models explicate phase separating intrinsically disordered motifs. Nat Commun 17, 2571 (2026). https://doi.org/10.1038/s41467-026-69252-z
Keywords: biomolecular condensates, intrinsically disordered proteins, phase separation, deep learning, protein motifs