Clear Sky Science · en
Local PI(4,5)P2 synthesis by septin-associated PIPKIγ isoforms controls centralspindlin association with the midbody during cytokinesis
How Cells Finish Splitting in Two
Cytokinesis—the final pinch that splits one cell into two—is easy to take for granted, but when it goes wrong it can help drive cancer and other diseases. This study uncovers how a specific set of molecules acts like construction foremen at the narrow bridge connecting two newborn cells, making sure the split finishes cleanly instead of snapping back or stalling.
The Final Bridge Between Daughter Cells
After a cell duplicates its genetic material and separates its chromosomes, it must physically divide. A contractile ring of protein filaments tightens around the cell’s middle, creating a deep groove called the cleavage furrow. As this furrow pulls inward, the two daughter cells remain linked by a thin intercellular bridge that contains a dense core, the midbody. The midbody is a tiny but complex structure made of bundled microtubules, scaffold proteins, and signaling lipids, and it acts as both a tether and a control center that coordinates the last steps of division and the final cut between cells. 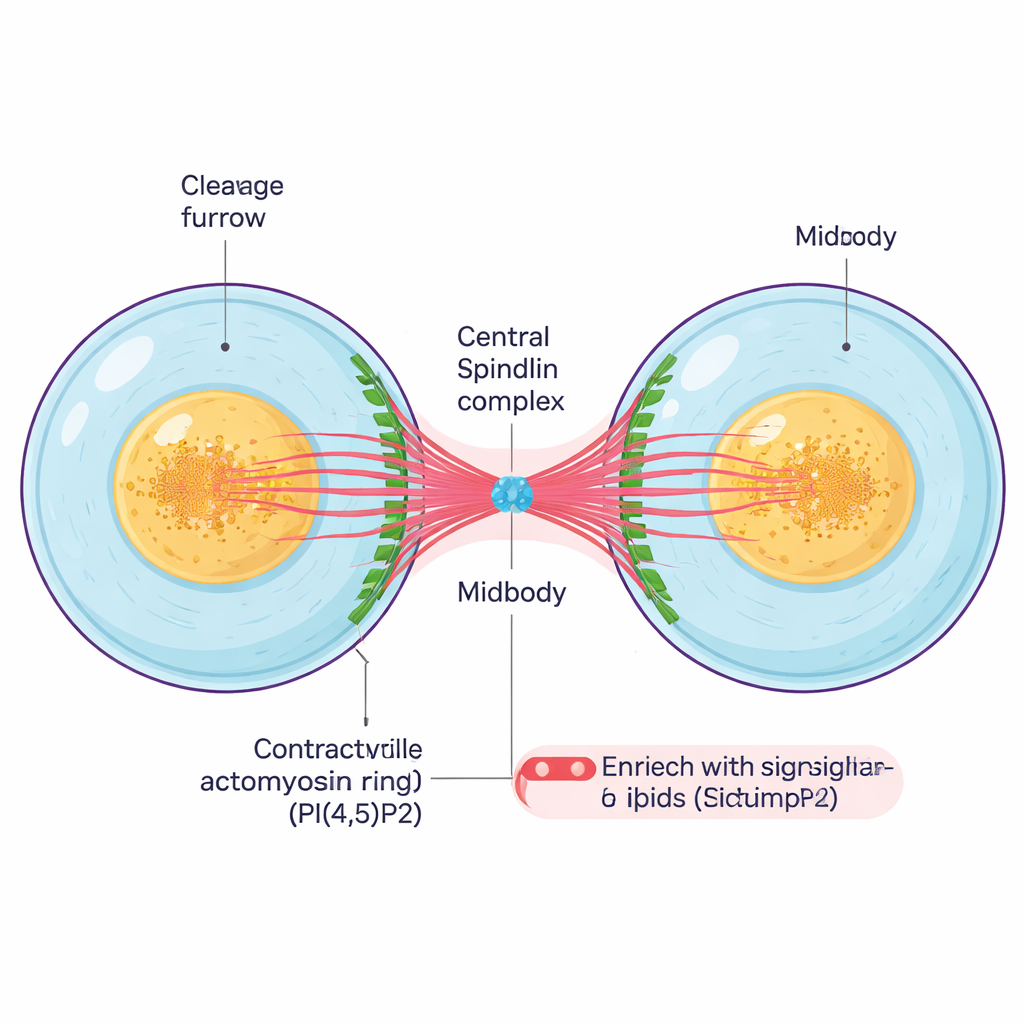
A Lipid Signal That Must Be in the Right Place
A key player in this process is a signaling fat molecule in the cell membrane called PI(4,5)P2. This lipid helps anchor many proteins that build and constrict the contractile ring and stabilize the bridge. The puzzle has been how cells control where and when PI(4,5)P2 appears, given that it can diffuse in the membrane. The authors focused on enzymes called type I PIP kinases, which make PI(4,5)P2, and in particular on one gene, PIPKIγ, that exists in several subtly different splice forms. By selectively reducing each kinase in human cells and observing the effects with advanced microscopy, they found that PIPKIγ has a special role late in cytokinesis, when the bridge and midbody are forming, even though it does not strongly change overall PI(4,5)P2 levels at the cell surface.
Septin Scaffolds Recruit a Specialized Enzyme
The team discovered that two splice versions of PIPKIγ, called i3 and i5, act like precision tools that are brought to the right spot by structural proteins known as septins. Septins form filaments and rings that help shape the bridge and organize other components. Biochemical pull-downs and cell imaging showed that PIPKIγ-i3 and PIPKIγ-i5 physically bind to septin complexes and decorate septin filaments, whereas other PIPKIγ variants do not. When the researchers mutated two specific amino acids in these splice inserts, the enzymes could no longer bind septins. In dividing cells, normal PIPKIγ-i3/i5 move from a diffuse membrane pattern to concentrate at the cleavage furrow and then outline the intercellular bridge and midbody in step with septins and another scaffold protein, anillin. Removing only the i3 and i5 variants scattered anillin and septins away from the midbody along the bridge, and many cells failed cytokinesis, ending up with multiple nuclei.
Local Lipid Production Locks in the Division Machinery
To test whether these effects depended on local PI(4,5)P2 production, the authors measured PI(4,5)P2 around the midbody and manipulated enzymes that add or remove this lipid. Depleting PIPKIγ-i3/i5 reduced PI(4,5)P2 at the midbody, while blocking a PI(4,5)P2-degrading enzyme had the opposite effect. Importantly, restoring a normal, septin-binding, catalytically active PIPKIγ splice form rescued the proper clustering of anillin and septins at the bridge, but kinase-dead or septin-binding–deficient mutants did not. Using live imaging and an expansion microscopy technique that physically inflates the sample to reveal fine details, the team showed that without PIPKIγ-i3/i5, septins no longer move onto bridge microtubules, the bridge becomes shorter and less bundled, and a crucial complex called centralspindlin fails to remain tightly associated with the midbody. 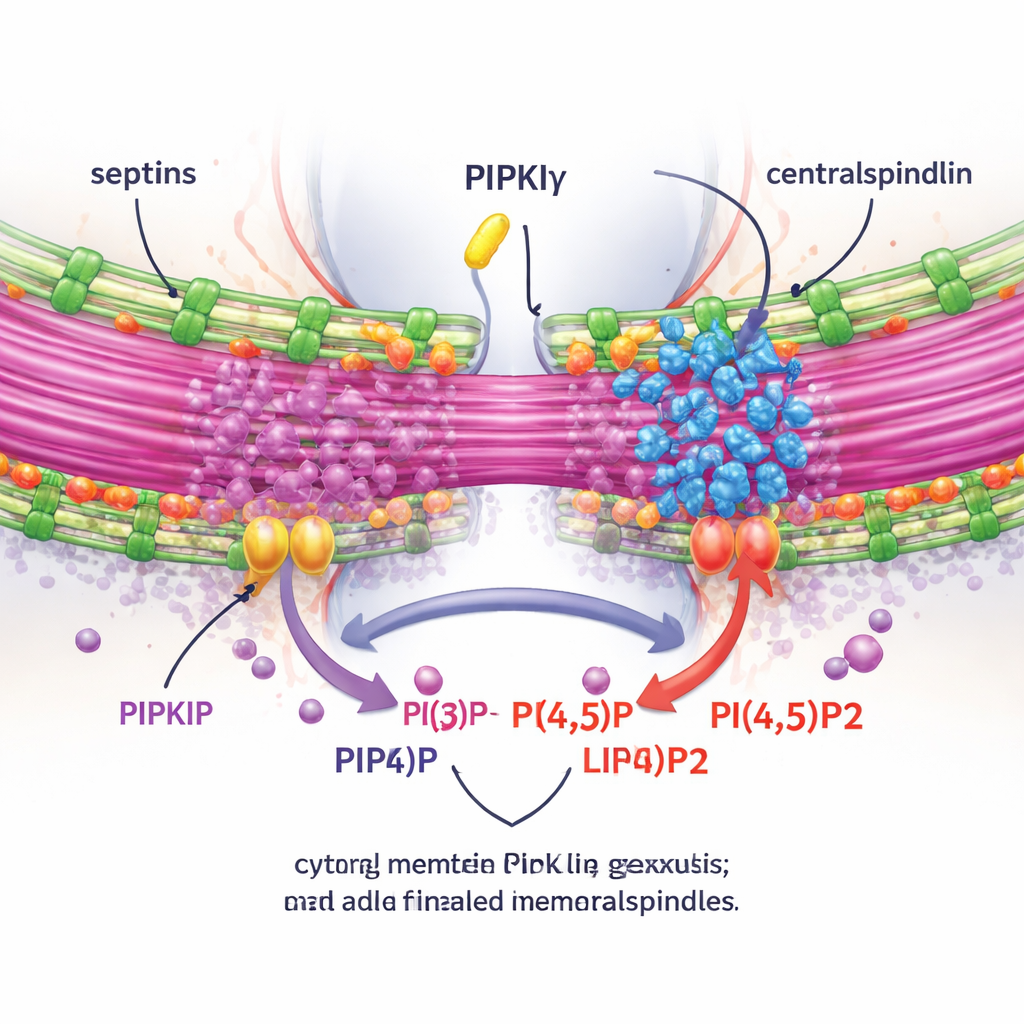
Building a Checkpoint for a Clean Cut
Centralspindlin helps link the midbody’s microtubules to the surrounding membrane and regulates small molecular switches that control cell shape. The study finds that centralspindlin associates with both septins and PIPKIγ, and that losing either partner similarly weakens its presence at the midbody. The authors propose that septins recruit PIPKIγ-i3/i5 to the ingressed furrow, where these enzymes create a concentrated patch of PI(4,5)P2. This local lipid pool, in turn, stabilizes anillin, centralspindlin, and septins near the midbody, promotes the bundling and stabilization of bridge microtubules, and allows the intercellular bridge to mature until the final abscission cut occurs. If this system is disrupted, PI(4,5)P2 appears in the wrong places, scaffolding proteins disperse, microtubules are poorly bundled, and cytokinesis often fails.
Why This Matters Beyond Basic Cell Biology
By pinpointing how specific PIPKIγ splice variants and septins cooperate to generate a local lipid signal at the midbody, this work explains how cells give spatial precision to a highly mobile signaling molecule. That precision is crucial for the safe completion of cell division, helping prevent chromosome mis-segregation and abnormal cell numbers that can favor tumor development. The same midbody structures also influence cell fate and proliferation after division, so understanding how they are assembled opens new avenues for exploring how errors in these nanoscale scaffolds might contribute to cancer and possibly to manipulating stem cells.
Citation: Russo, G., Hümpfer, N., Jaensch, N. et al. Local PI(4,5)P2 synthesis by septin-associated PIPKIγ isoforms controls centralspindlin association with the midbody during cytokinesis. Nat Commun 17, 1482 (2026). https://doi.org/10.1038/s41467-026-69224-3
Keywords: cell division, cytokinesis, signaling lipids, septins, cancer biology