Clear Sky Science · en
Single-cell structural biology with intracellular electron crystallography
Seeing the Shape of Life, One Cell at a Time
Proteins are the tiny machines that keep every cell alive, but to truly understand how they work, scientists need to see their three-dimensional shapes in fine detail. Traditionally, that has meant purifying huge amounts of protein and growing large, fragile crystals outside the cell—often a slow, failure‑prone process. This study presents a new way to read out protein structure directly from a single crystal inside a single cell, using electrons instead of X‑rays. It points toward a future where high‑resolution structural biology can be done in ordinary labs, and even at the level of individual cells.
Crystals Hidden Inside Living Cells
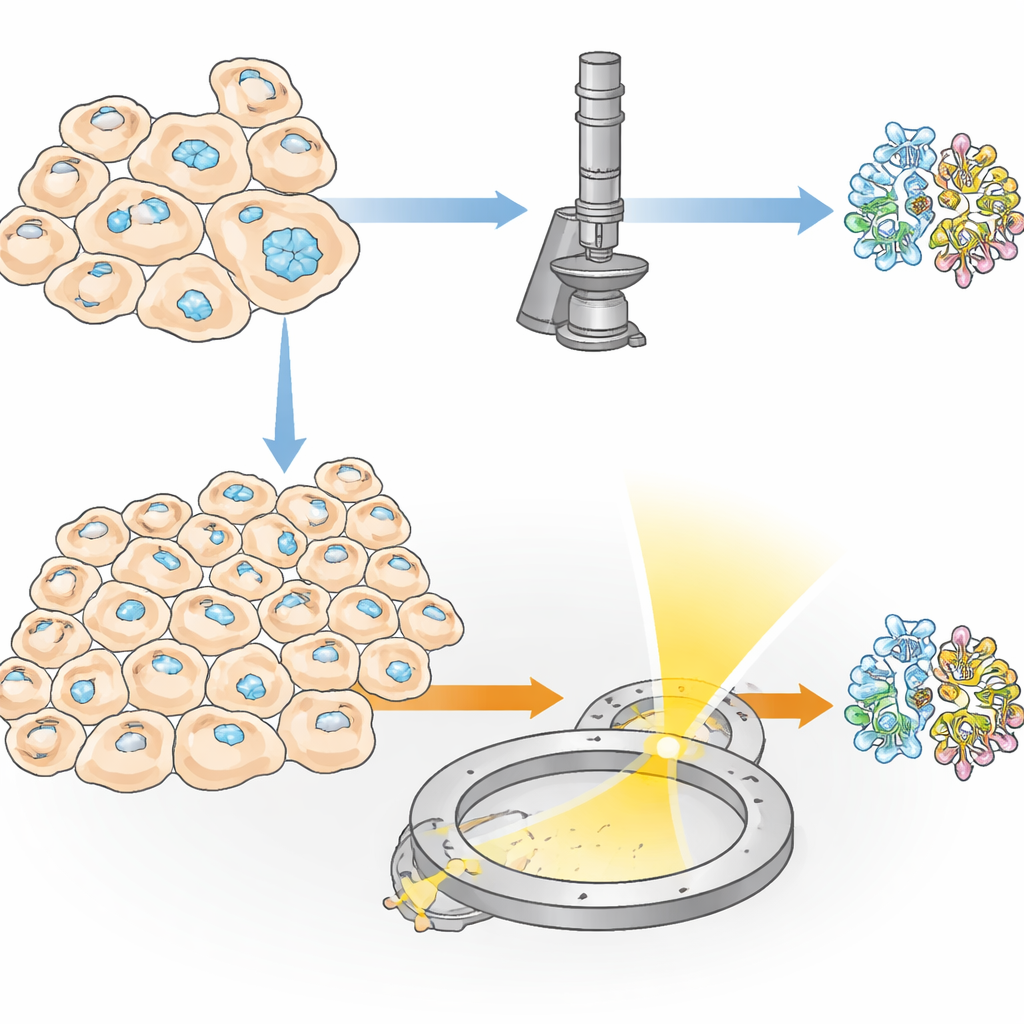
Some proteins naturally gather into tiny crystals inside living cells, serving purposes like storage, protection, or helping cells respond to stress. Researchers can also coax cells to produce such crystals by engineering them to make large amounts of a chosen protein. This “in‑cell” crystallization has two big advantages: the protein never leaves its natural‑like environment, and delicate features—such as sugar decorations or small bound molecules—can be preserved in ways that often fail in standard test‑tube crystallization. However, one major obstacle has remained: in many experiments, only a tiny fraction of cells actually form crystals, so traditional X‑ray methods need tens of thousands of crystals and, therefore, enormous numbers of cells.
A New Route: Electrons Instead of X‑Rays
The authors introduce a method they call IncelluloED, which marries in‑cell crystallization with three‑dimensional electron diffraction. Electrons interact far more strongly with matter than X‑rays do, meaning that useful data can be collected from crystals that are both smaller and fewer in number. The team chose a fungal protein called HEX‑1, which normally forms hexagonal crystals that help plug tiny pores between fungal cells under stress. By producing this protein inside insect cells, they created regular microscopic crystals that served as a test case for the new pipeline.
Turning One Crystal into a Detailed Map
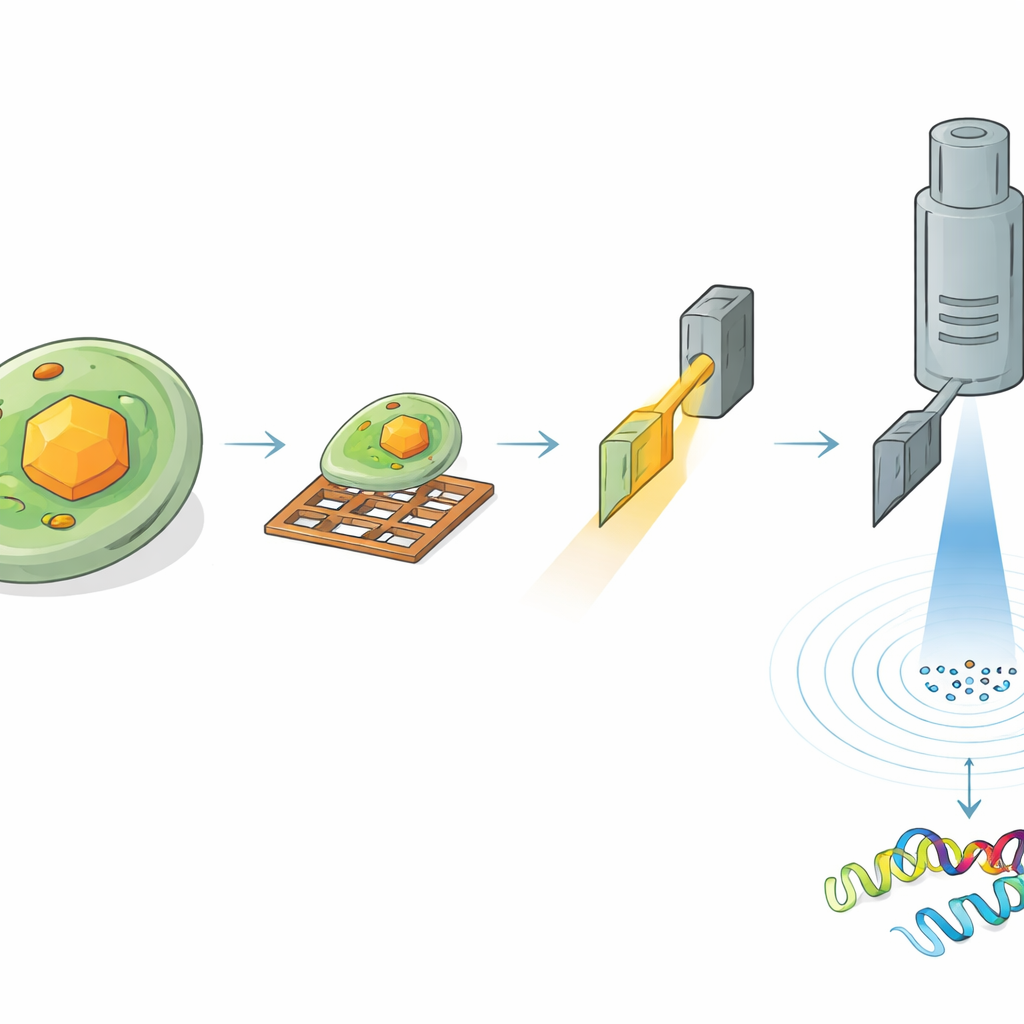
To read the structure of HEX‑1 inside a cell, the researchers had to find and carefully thin down the right region of the sample. First, they froze the crystal‑bearing cells on tiny metal grids and coated the surface with a thin platinum layer. Using cryogenic light microscopy, they scanned large areas of the grid to locate cells containing crystals and measured the crystals’ three‑dimensional positions beneath the surface. Next, they moved the same sample into a specialized instrument that combines a scanning electron microscope with a focused ion beam. Guided by the earlier light images, they milled away surrounding material to carve out an ultra‑thin slice, or lamella, passing through a chosen crystal, just a few hundred nanometers thick—ideal for electrons to pass through.
Electrons Reveal Atomic Detail from Microscopic Volumes
The prepared lamellae were then transferred to a high‑end electron microscope operating at cryogenic temperatures. As the crystal slice was slowly rotated in the microscope, a finely controlled electron beam passed through it, creating a series of diffraction patterns—delicate spot arrangements that encode the positions of atoms. From a crystal volume of roughly 1.6 cubic micrometers, the team reconstructed the full 3D structure of HEX‑1 at 1.9 ångström resolution, sharp enough to model most of the protein’s side chains. Even smaller volumes of about 0.8 cubic micrometers yielded a nearly identical structure at only slightly lower resolution. Importantly, the resulting models closely matched those obtained using a conventional serial X‑ray approach that required more than 60,000 crystals and a total crystal volume roughly seven million times larger.
How This Changes the Structural Biology Game
Side‑by‑side comparisons showed that the structure determined from a single in‑cell electron‑diffraction crystal is essentially the same as the one averaged from tens of thousands of crystals with X‑rays. Any differences were minor and mostly confined to flexible loops, where natural motion is expected. The researchers also demonstrated that the electron doses used were low enough to avoid serious radiation damage, and that every crystal they processed produced high‑quality data. Although preparing the thin lamellae still requires skill and time, the necessary instruments—cryo‑light microscopes, focused ion beam systems, and cryo‑electron microscopes—are now widespread in many research centers.
From Many Cells to a Single‑Cell Structural Lab
This work shows that it is now possible to determine an atomic‑level protein structure from just one crystal inside one cell, without ever purifying the protein. IncelluloED could be especially powerful when only a few cells form crystals, or when proteins are hard to isolate without losing important partners or chemical groups. As the workflow becomes more automated and is extended to other proteins, it may enable researchers to explore how structures vary from cell to cell, to study disease‑related changes in their native setting, and even to support drug discovery directly in living cells. In effect, the study brings the vision of a “single‑cell structural laboratory” much closer to reality.
Citation: Bílá, Š., Pinkas, D., Khakurel, K. et al. Single-cell structural biology with intracellular electron crystallography. Nat Commun 17, 2109 (2026). https://doi.org/10.1038/s41467-026-69205-6
Keywords: electron diffraction, in cellulo crystallography, single-cell structural biology, protein structure, cryo-EM