Clear Sky Science · en
TANGO: Analysis and curation of particles in cryo-electron tomography
Seeing Cells in 3D, Then Making Sense of the Crowd
Cryo-electron tomography lets scientists freeze living cells in action and capture them in three dimensions, almost like pausing a movie at atomic detail. But once thousands of tiny molecular "specks" have been mapped inside a cell, a new problem appears: how do we understand who is sitting next to whom, who forms teams, and where the important patterns are hiding in this crowded molecular city? This study introduces TANGO, a software framework that turns raw 3D particle maps into readable stories about how molecules are arranged and work together.
From Dots in Ice to a Map of Molecular Neighbors
Cryo-electron tomography starts with tilting a frozen sample in the microscope to record many 2D images. These are combined into a 3D volume, called a tomogram, where individual molecular complexes appear as fuzzy spots. Existing methods already use these data to sharpen the shapes of molecules by averaging many copies, but they largely ignore an equally rich treasure: the exact positions and orientations of every particle. TANGO is built to exploit this overlooked information. It treats all these particles as points in space, each with a direction, and analyzes how they are positioned and oriented relative to one another. By doing so, it moves beyond simply asking "what does this molecule look like?" to "how are these molecules organized together in the cell?" 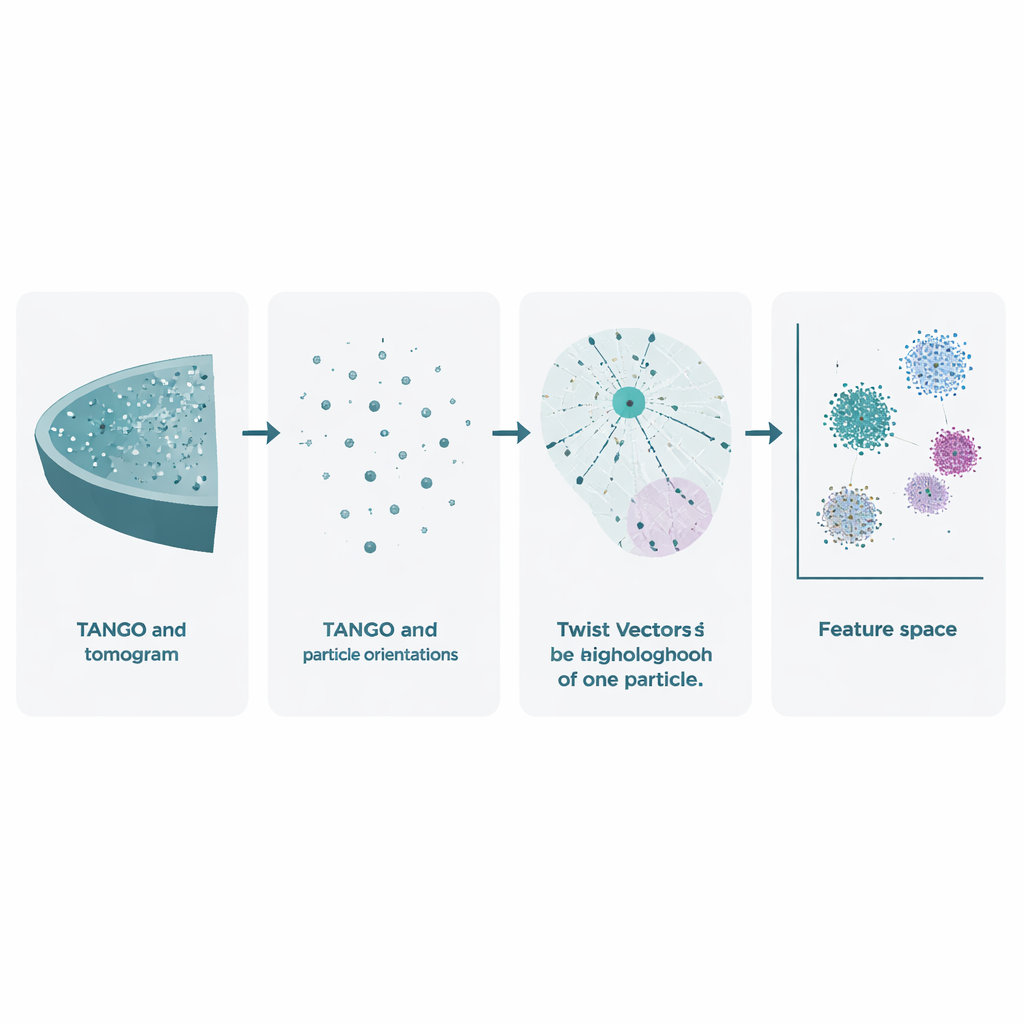
Capturing Twists and Turns with a New Descriptor
At the heart of TANGO is the idea of "twist vectors". For any given particle, the software looks at all its neighbors within a chosen radius and records two things: where each neighbor lies in 3D space and how it is rotated relative to the particle at the center. These combined position-and-angle relationships are encoded as concise numerical descriptions known as twist descriptors. Because TANGO always reorients neighborhoods into a common reference frame, these descriptors are insensitive to how the whole sample is rotated in the microscope. This makes it possible to compare local neighborhoods across different tomograms and experiments in a consistent way.
Cleaning Up Noisy Data and Rebuilding Molecular Assemblies
Real experimental data are messy: automatic picking methods can include many false particles and lose track of which small pieces belong to which large structure. TANGO tackles this by turning the network of twist relationships into a graph, where particles are nodes and neighborhood links are edges. By analyzing how nodes connect, TANGO can group particles back into their correct parent assemblies and discard outliers that do not fit the expected geometry. The authors show that this approach accurately recovers the ring-shaped architecture of nuclear pores in the cell’s nuclear envelope, the tubular arrangement of microtubules, and the roughly spherical shells of immature HIV virus-like particles. In each case, TANGO both cleans the particle lists and restores which pieces belong together, often matching or improving on careful manual curation.
Spotting Subtle Defects and Patterns in Lattices
Many viruses and cellular structures form repeating lattices, like molecular tiling on curved surfaces. TANGO uses its twist descriptors to measure how regular these patterns are and to detect places where they bend or break. A key ingredient is an "angular score" that compares orientations while respecting built-in symmetries, such as the six-fold symmetry of hexamers. Applied to mature HIV capsids, TANGO detects pentamers—five-part units needed to close the conical shell—hidden among large fields of hexamers. In immature HIV lattices, it separates well-ordered regions from distorted ones and links low angular scores to irregular, broken areas of the shell. Similar analyses on synthetic chromatin and ribosome data reveal stacked nucleosomes, helical DNA–protein arrangements, and recurring pairs of ribosomes that resemble previously described translation states. 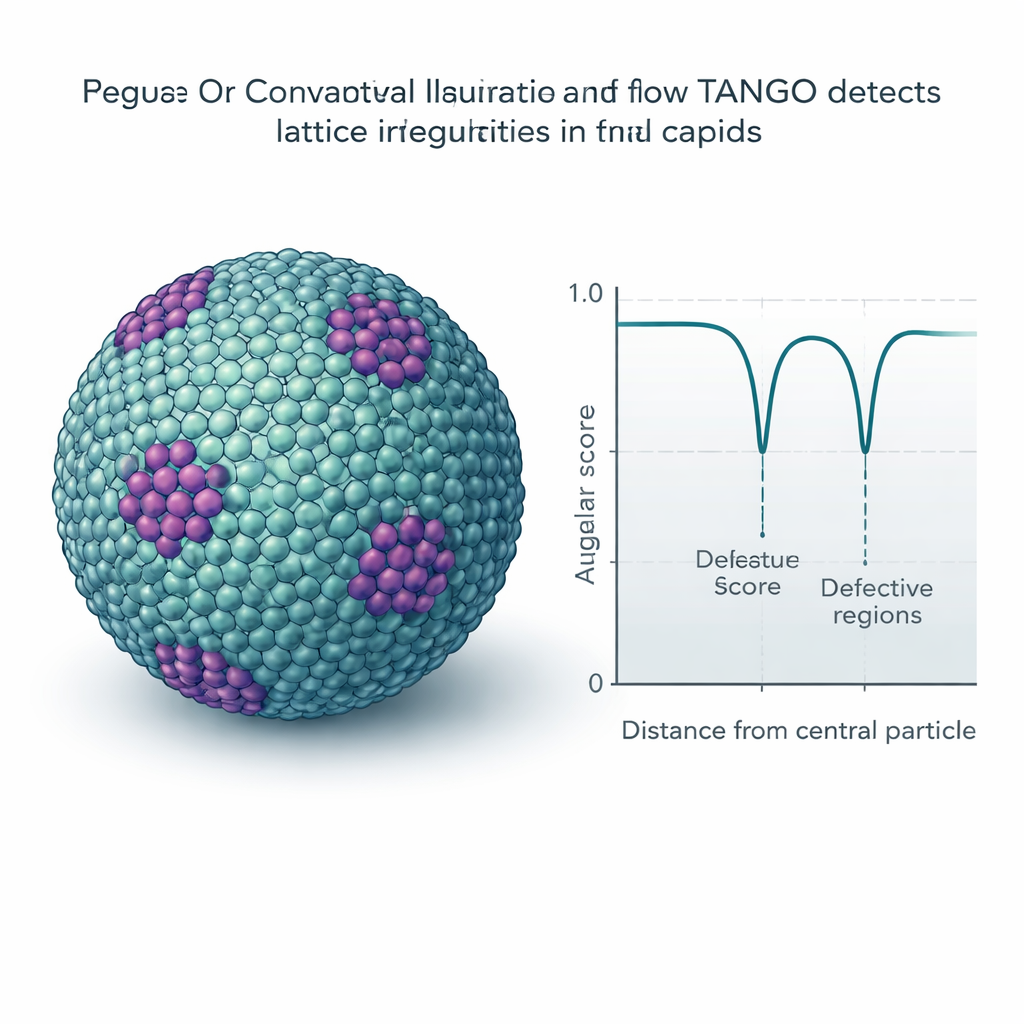
A Flexible Toolbox for Exploring Cellular Architecture
TANGO is implemented as open-source Python software and comes with a graphical interface so that users can test different neighborhood shapes, filters, and features without heavy programming. Because it is modular, researchers can plug in their own geometric measures or pattern descriptors and immediately use them within the same workflow. For newcomers, this lowers the barrier to exploring spatial organization; for experts, it provides a framework that can grow with new ideas and datasets.
Why This Matters for Understanding Living Cells
In simple terms, this work gives biologists a way to move from static pictures of individual molecules to dynamic maps of how those molecules are arranged and cooperate inside cells. By encoding the "who is near whom" and "how they are oriented" relationships into robust numerical features, TANGO turns noisy 3D microscopy data into patterns that can be clustered, compared, and tested statistically. This can reveal hidden assemblies, pinpoint defects in viral shells, and uncover rare molecular arrangements linked to disease or drug action. As cryo-electron tomography becomes more widely used, tools like TANGO will help transform dense clouds of particles into clear insights about the inner choreography of life.
Citation: Schreiber, M., Turoňová, B. TANGO: Analysis and curation of particles in cryo-electron tomography. Nat Commun 17, 1557 (2026). https://doi.org/10.1038/s41467-026-69195-5
Keywords: cryo-electron tomography, spatial organization, molecular lattices, viral capsids, ribosomes