Clear Sky Science · en
The redox driven Na+-pumping mechanism in Vibrio cholerae NADH-quinone oxidoreductase relies on dynamic conformational changes
How Cholera’s Power Plant Became a Drug Target
The bacterium that causes cholera, Vibrio cholerae, survives and multiplies by running a tiny molecular power plant in its cell membrane. This study reveals, in unprecedented detail, how one of its key engines—an enzyme called Na⁺-NQR—changes shape as it works to pump sodium ions out of the cell. Understanding this molecular machinery not only satisfies basic curiosity about how life converts food into usable energy, but also points to new ways to design antibiotics that shut down harmful bacteria without affecting human cells.
A Sodium-Powered Engine in Bacteria
Many marine and disease-causing bacteria rely on Na⁺-NQR as the first step in their respiratory chain, the series of reactions that extracts energy from nutrients. Na⁺-NQR sits in the inner membrane and takes electrons from a fuel molecule called NADH, passing them along a chain of colored “helper” molecules (called cofactors) to quinone, another small molecule embedded in the membrane. As electrons flow, the enzyme uses that energy to push sodium ions (Na⁺) from the inside of the cell to the outside, creating a sodium gradient. This gradient is like a charged battery: it powers the bacterial flagellar motor for swimming, helps make ATP (the cell’s energy currency), and drives nutrient uptake and drug efflux. Because Na⁺-NQR exists only in bacteria and is very different from the related enzyme in our mitochondria, it is an attractive target for highly selective antibiotics.
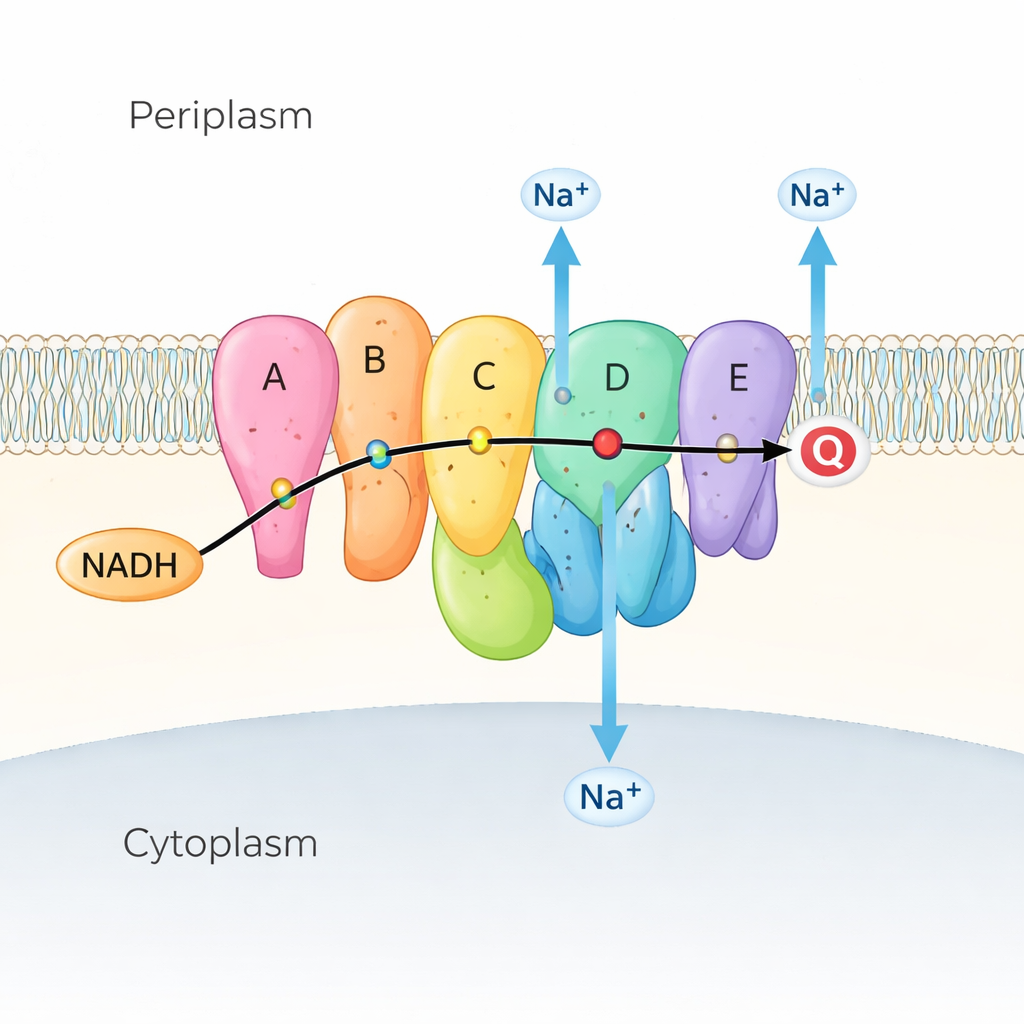
Watching the Machine Move
Previous X-ray and cryo–electron microscopy snapshots showed where Na⁺-NQR’s subunits and cofactors sit, but also revealed a puzzle: some critical electron jumps were too far apart for electrons to move efficiently unless the protein shifted its shape. The authors tackled this by capturing Na⁺-NQR in many slightly different conditions. They used mutants that remove specific cofactors, drugs that stall the reaction at particular stages, and solutions with and without sodium. With high-resolution cryo-EM and advanced image analysis, they could separate and reconstruct multiple conformations of the same enzyme, effectively turning static snapshots into a movie of its working cycle.
A Flexible Arm and a Shuttling Clamp
One key finding involves a flexible cytoplasmic subunit, NqrF, which holds two electron-carrying cofactors. The team identified three positions for its “ferredoxin-like” domain—nicknamed the “up,” “middle,” and “down” states. In the rare but crucial “down” state, this small domain swings into a pocket formed by two membrane subunits, NqrD and NqrE, bringing its iron–sulfur cluster close enough to pass an electron onward. Meanwhile, a periplasmic (outer) subunit called NqrC behaves like a movable clamp. In one conformation (“stable”), it nestles against another membrane subunit, NqrB, positioned to pass electrons toward the final quinone acceptor. In a second (“shifted”) conformation, NqrC moves toward NqrD/E, bringing its flavin cofactor closer to accept an electron from their iron–sulfur center. These movements show that NqrF and NqrC act as dynamic couriers, bridging large gaps in the cofactor chain.
How Shape Changes Pump Sodium
The heart of the sodium pump lies in the paired NqrD and NqrE subunits, which form a central bundle of membrane-spanning helices. Using both structures and atomic-level molecular dynamics simulations, the authors show that when the iron–sulfur cluster in NqrD/E is reduced (gains an electron), it creates a negatively charged site that attracts a Na⁺ ion and a few water molecules from the cytoplasmic side, forming a transient binding pocket. Hydrophobic amino acids above and below this pocket behave like inner and outer gates, controlling access to the ion. When NqrD/E switch from an “inward-open” to an “outward-open” shape, the Na⁺ moves toward the periplasmic side and is eventually released as the iron–sulfur cluster is reoxidized. Simulations show Na⁺ remaining partially hydrated but never seeing a continuous water-filled tunnel, suggesting the enzyme is tightly sealed against unwanted proton leaks.
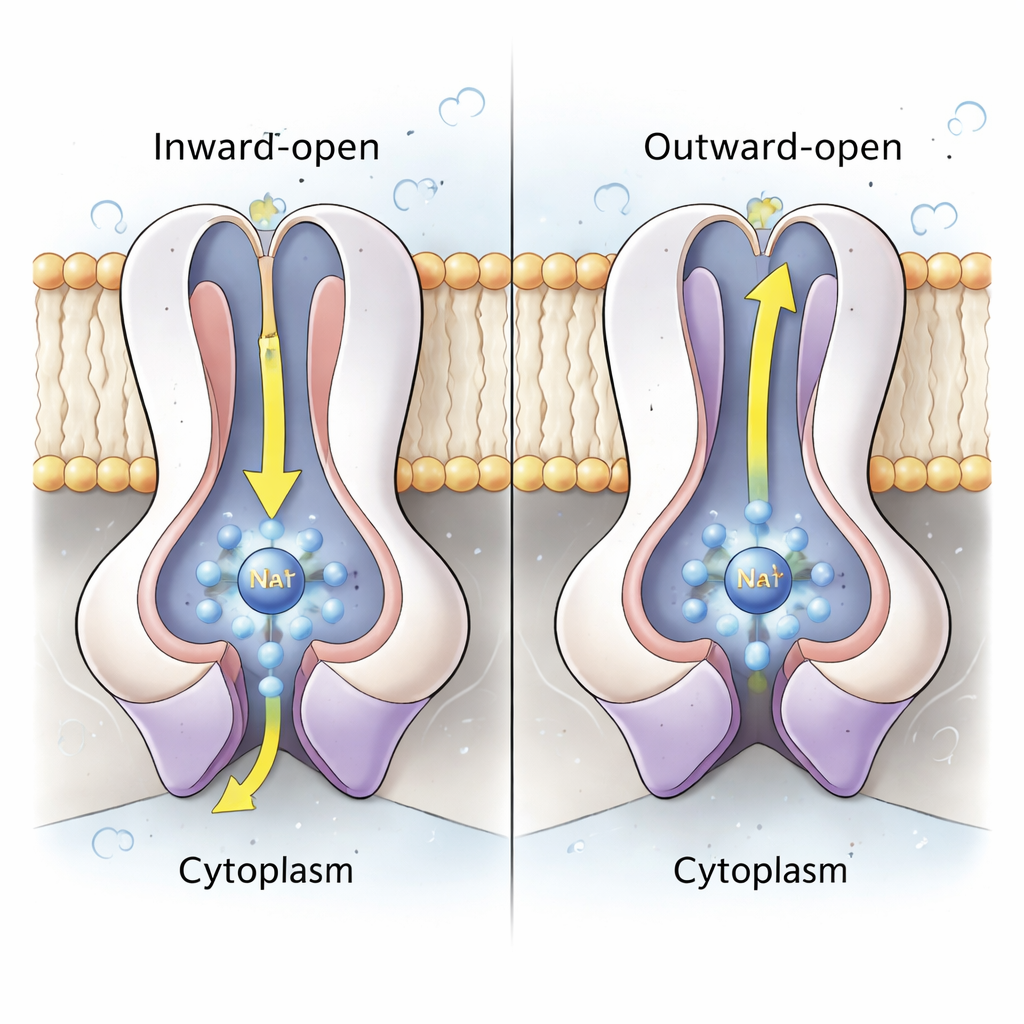
A Coordinated Cycle That Locks in Direction
By combining all their structural states and simulations, the researchers propose a six-step cycle. NADH first binds and donates electrons to NqrF, which then passes one electron to NqrD/E, triggering Na⁺ uptake from the cytoplasm and loading the central pocket. The resulting conformational switch of NqrD/E to the outward-open form both expels Na⁺ to the periplasm and encourages NqrC to move into position to accept the electron. Subsequent steps transfer electrons along to the final flavin and riboflavin cofactors and finally to quinone, which must receive two electrons to become quinol. Importantly, the same shape changes that move Na⁺ also alternately loosen and stiffen NqrF and NqrC, making forward electron flow likely only when Na⁺ is being pumped outward and discouraging backward slippage. This coupling helps the enzyme pump Na⁺ against an existing gradient, keeping the bacterial “battery” charged.
Why This Matters for Medicine and Biology
To a non-specialist, this work shows how life builds exquisitely choreographed machines out of proteins, where tiny changes in charge trigger large, useful motions. For infectious disease research, it clarifies that Na⁺ translocation in Na⁺-NQR occurs mainly through the NqrD/E core, not through the previously suspected NqrB subunit. That insight refines where future drugs should bind to best disrupt the engine that powers Vibrio cholerae and other pathogens, while sparing human enzymes. More broadly, the study provides a structural blueprint for how redox reactions can be wired to ion pumping—a design principle that nature reuses in many forms of bioenergetic machinery.
Citation: Ishikawa-Fukuda, M., Seki, T., Kishikawa, Ji. et al. The redox driven Na+-pumping mechanism in Vibrio cholerae NADH-quinone oxidoreductase relies on dynamic conformational changes. Nat Commun 17, 1394 (2026). https://doi.org/10.1038/s41467-026-69182-w
Keywords: sodium pump, bacterial respiration, Na+-NQR, cryo-EM, antibiotic targets