Clear Sky Science · en
PRICE: direct and robust detection of microRNAs at single-nucleotide resolution
Why tiny genetic messages matter
Inside every cell, there is a swarm of tiny RNA molecules called microRNAs that help fine‑tune which genes are turned on or off. A change of just one “letter” in these short molecules can tilt the balance toward health or disease, including cancers. Doctors and researchers would love to read these tiny differences directly from blood or tissue, but today’s methods are often slow, expensive, and not precise enough to reliably spot a single‑letter change. This paper introduces a new laboratory technique, called PRICE, that promises to read these microscopic typos in microRNAs more simply and accurately, opening doors for earlier cancer detection and better tracking of how tumors behave.
A new way to sort look‑alike molecules
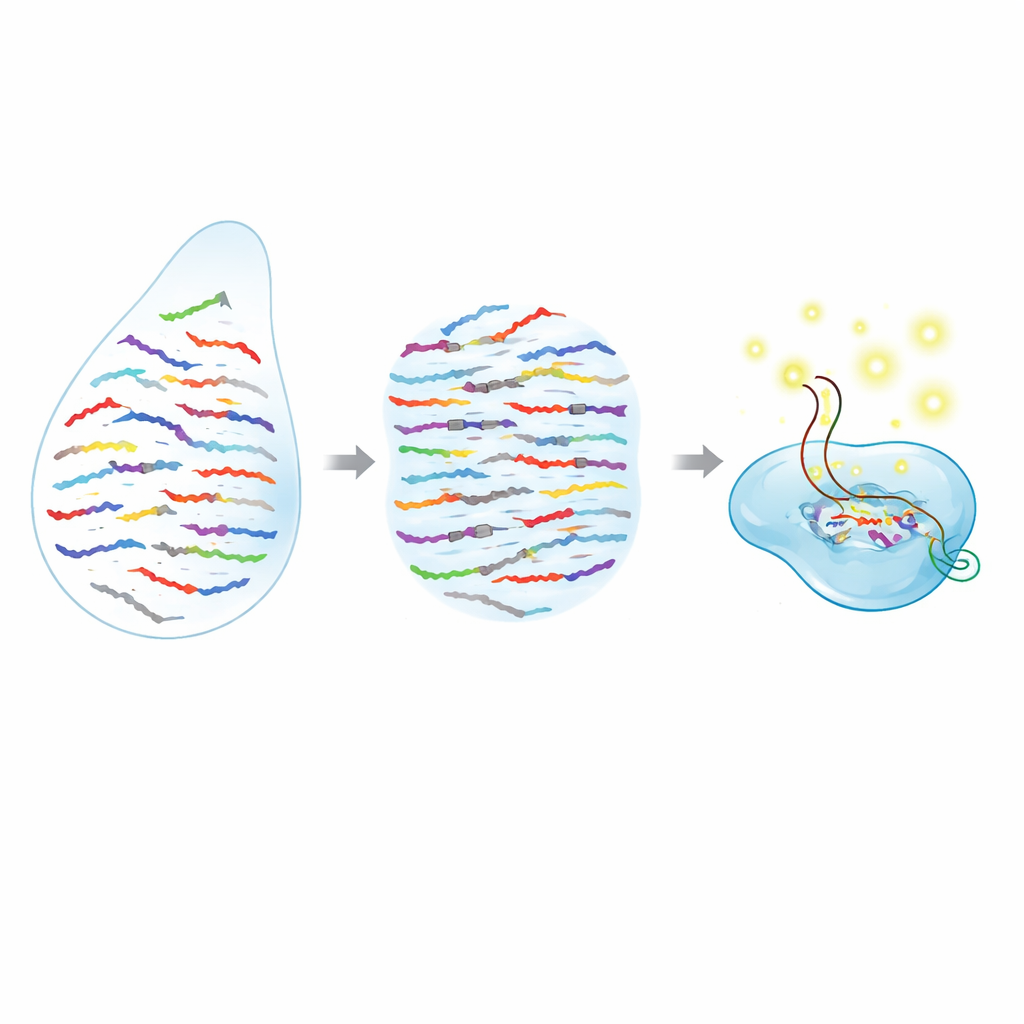
MicroRNAs are extremely short, which makes them hard to tell apart when their sequences differ by only one building block. Many current tools, such as PCR and RNA sequencing, do a good job counting microRNAs overall, but struggle when two versions are almost identical, or they require complex sample preparation and costly instruments. The authors combine two powerful ideas to overcome this: a programmable molecular “scissor” from the CRISPR family, and synthetic DNA‑like pieces called peptide nucleic acids (PNAs). The CRISPR enzyme Cas13a can be told which RNA to recognize, and once it finds its target it begins cutting nearby reporter molecules, producing a bright fluorescent signal. PNAs, meanwhile, are short strands that bind especially tightly to matching RNA sequences but lose their grip sharply if even a single letter is wrong.
How the PRICE method works
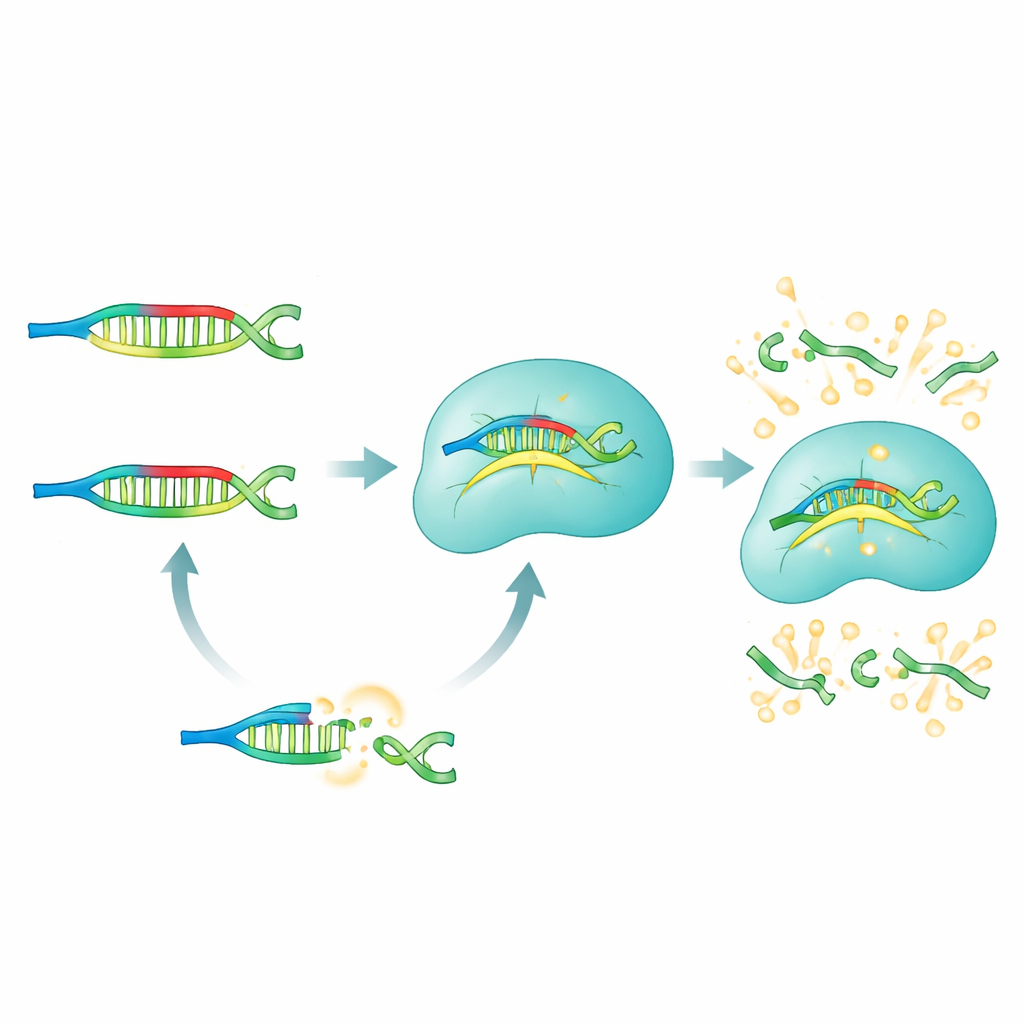
In the PRICE system, PNAs act like selective blockers that silence everything researchers don’t want to see. For a given microRNA of interest, the team designs a panel of PNA blockers that perfectly match all of the “wrong” versions—those with unwanted single‑letter changes—while deliberately mismatching the chosen target. When a biological sample (from serum, cells, or tissue) is mixed with these PNAs, the non‑target microRNAs are tightly bound and effectively masked. The true target is left unbound and free to interact with the Cas13a complex, which has been pre‑programmed with a guide RNA that recognizes that microRNA. Only this unblocked target can switch Cas13a into its active cutting mode, which in turn snips apart fluorescent reporter molecules and generates a strong, easily measured glow.
Fine‑tuning single‑letter precision
The authors systematically explored how to design PNA blockers that are strong enough to grab the wrong microRNA, yet weak enough to ignore the desired one that differs by only one letter. They found that two temperature‑like design rules are crucial: the PNA should form a stable pair with the unwanted mutant at or above body temperature, but form a much weaker pair with the target that falls apart well below that temperature. Using these rules, supported by a simple predictive model, they built PNAs that cut false signals from mutant microRNAs by more than 70 percent while leaving the true signal nearly untouched. Importantly, the added PNAs did not reduce the overall sensitivity: PRICE could still detect microRNAs at concentrations of only a few tens of femtomolar, comparable to the best amplification‑based tests but without the extra copying steps.
Putting the test to work in cancer samples
To show that PRICE is more than a theoretical gadget, the team focused on the let‑7 family of microRNAs, which is heavily involved in cancer biology. They demonstrated that PRICE can pick out target microRNAs from mixtures containing many near‑identical family members and can even detect rare variants that make up as little as 5 percent of a sample. The researchers then turned to real clinical material: liver cancer cell lines, tissue biopsies, and patient serum. In both tumor tissue and blood, PRICE consistently detected lower levels of specific let‑7 microRNAs in patients compared to non‑cancer controls, in line with earlier studies. When directly compared with standard RT‑qPCR, PRICE showed at least similar and often better ability to distinguish cancer from non‑cancer cases, especially in low‑abundance serum samples where PCR results became noisy.
Why this matters for future diagnostics
At its core, this work shows that it is possible to directly read out single‑letter differences in tiny regulatory RNAs in a practical, scalable way. By letting PNAs lock down unwanted look‑alike sequences and handing the final readout to a CRISPR enzyme, PRICE offers a flexible toolbox for designing tests that can be retargeted simply by swapping the blocker panel and guide RNA. Beyond liver cancer and the let‑7 family, the same strategy could be extended to many other diseases where small RNA changes matter, and even to DNA variants when paired with related CRISPR enzymes. The authors also demonstrate a compact, multi‑channel reader that can run PRICE assays outside of large central labs, hinting at future point‑of‑care tests. If developed further, PRICE could become a practical way to monitor subtle genetic shifts that signal cancer and other diseases long before symptoms appear.
Citation: Wang, B., Zhou, S., Zhang, X. et al. PRICE: direct and robust detection of microRNAs at single-nucleotide resolution. Nat Commun 17, 2647 (2026). https://doi.org/10.1038/s41467-026-69181-x
Keywords: microRNA diagnostics, CRISPR Cas13a, peptide nucleic acid, single nucleotide variant, liver cancer biomarkers