Clear Sky Science · en
Impact of solvent forces and broken symmetry on the assembly of designed proteins at a liquid-solid interface
Why this tiny world on a surface matters
From solar cells to medical sensors, many future technologies will rely on building precise structures where biological molecules meet solid materials. This study looks at how custom-designed protein "rods" arrange themselves on mineral surfaces in salt water. The surprising lesson is that the water right at the surface, and subtle asymmetries in the crystal beneath it, can completely change how these proteins line up—sometimes producing patterns that classic theory says should not exist.
Designing proteins to fit a crystal
The researchers work with a man-made protein shaped like a short, stiff rod. Its surface chemistry and spacing of charged groups were carefully designed to match the pattern of potassium ions on a common mineral called mica. In principle, this should make each protein rod prefer three equivalent directions on the crystal surface, like aligning along three spokes of a wheel. Earlier work showed that, even with such careful design, the proteins formed multiple unexpected patterns instead of just the one engineers had in mind. That inconsistency hinted that some important force was missing from current design rules.
Two nearly identical surfaces, two very different outcomes
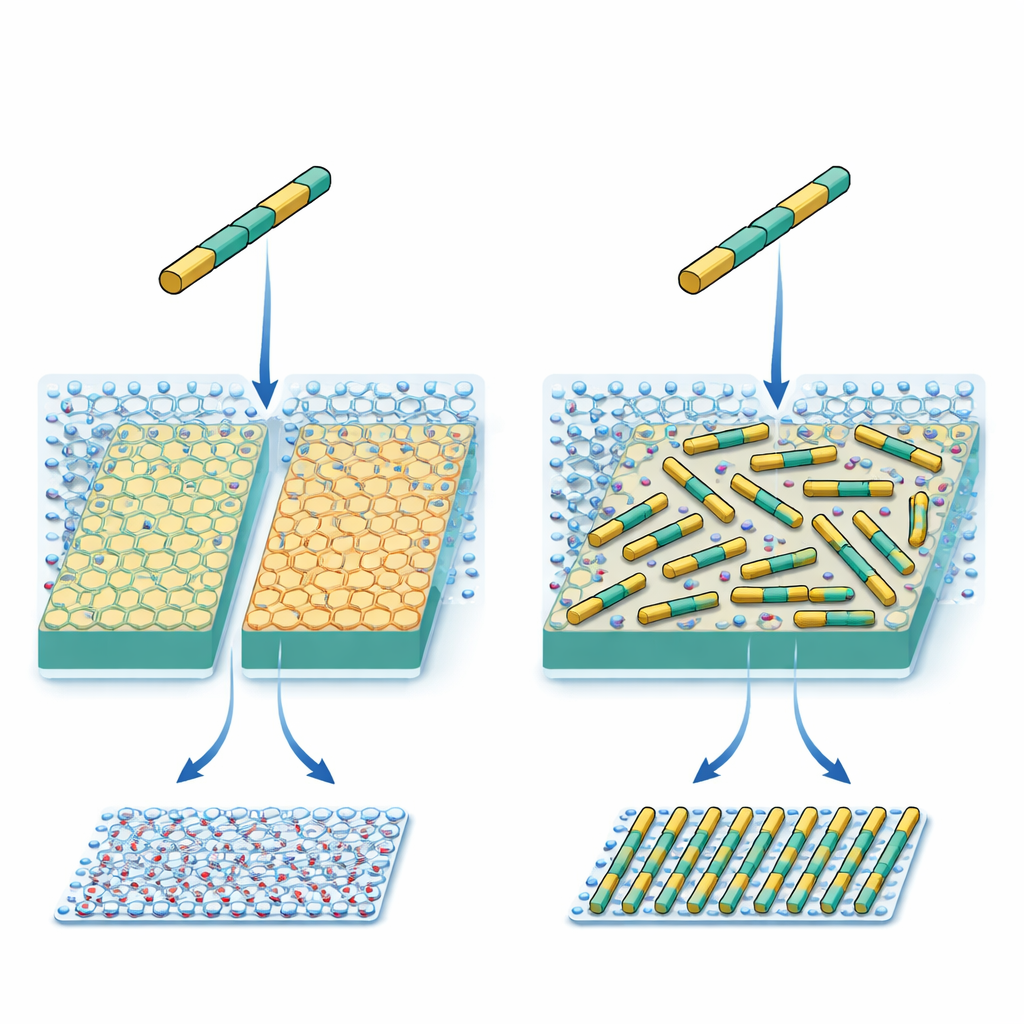
To track what was going on, the team used high-speed atomic force microscopy, which can watch individual protein rods move and assemble on a surface in real time. They compared two closely related forms of mica. Both have the same exposed potassium lattice, but their internal atomic structures differ slightly, which in turn changes how water organizes in layers just above the surface. At modest salt levels, the rods formed a dense but locally disordered carpet on both surfaces, with only small patches of alignment. When the salt concentration was raised very high, however, the behavior split: on one mica type, the rods remained disordered along three directions, while on the other they spontaneously formed long, parallel, evenly spaced rows across the entire surface.
Water layers and broken symmetry
Computer modeling of the minerals and surrounding water helped explain this split personality. On the more symmetric form of mica, the first and second layers of water molecules preserve a regular, hexagonal pattern. On the less symmetric form, internal atoms and buried groups within the crystal break that three-way symmetry, and this broken pattern is transmitted into the nearby water layers, which develop stripe-like regions. The proteins do not touch bare crystal alone; they interact with this structured water as well. As a result, one orientation of the rods becomes slightly more favorable than the other two, even though the designed protein–crystal match would suggest all three should be equal.
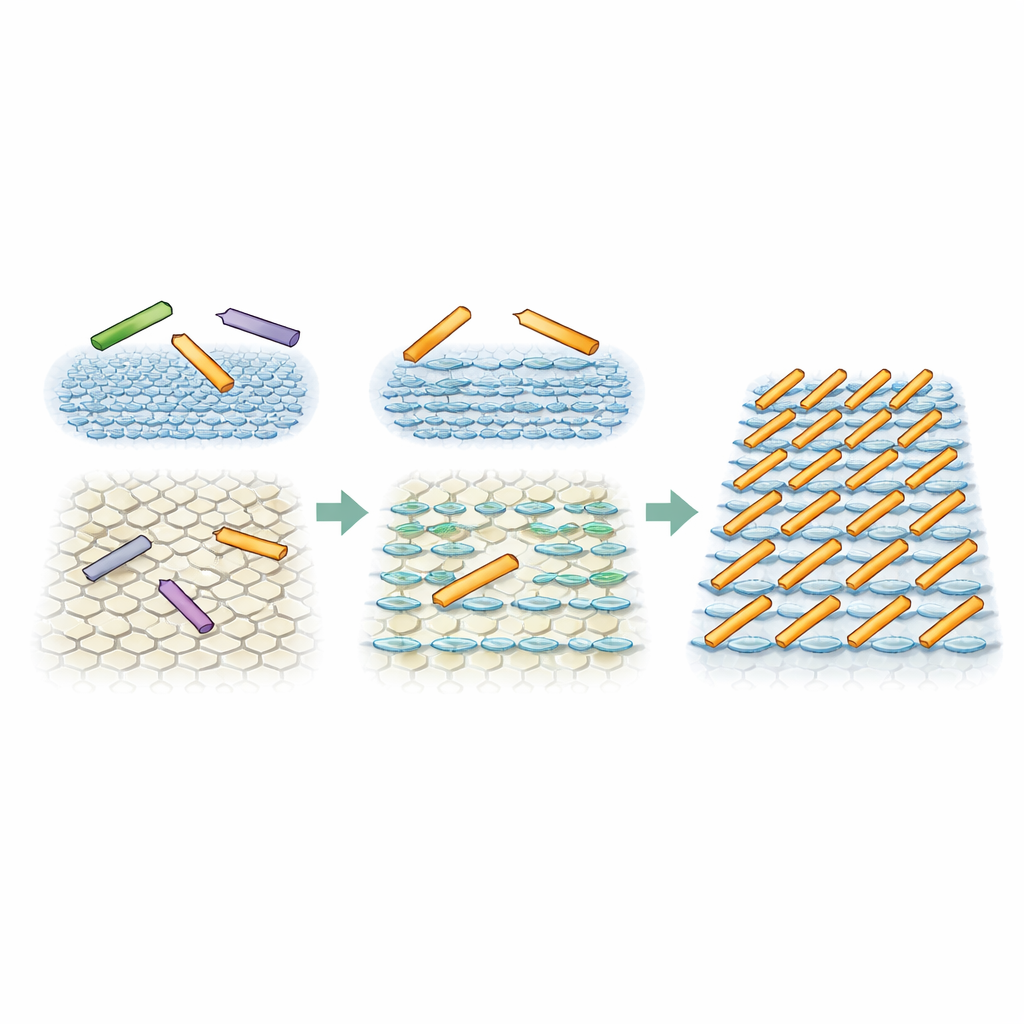
Simulations reveal an unexpected phase
To test whether a subtle directional bias from the surface environment could really explain the observed patterns, the researchers ran Monte Carlo simulations of simple hard rectangles representing the protein rods. In one set of simulations, all three orientations were equally likely, mimicking the fully symmetric surface. In that case, the rods stayed in a high-density but disordered state, with only temporary or limited alignment, just as seen on one mica type. In a second set, one direction was made only modestly more favorable—about twice as likely as the other two—standing in for the influence of the striped water layers. Under these conditions, and when the rods could move enough, the system naturally evolved into a state with parallel, evenly spaced rows. This is a so‑called smectic phase, which long-standing theory says should not appear for non‑interacting rods in two dimensions, yet the small directional push from the interface made it stable.
Rethinking how we design bio‑inspired materials
Put simply, this work shows that finely tuned protein–surface chemistry is not enough to predict how designed proteins will arrange themselves on real materials. The water sitting between the protein and the solid, and the way the crystal subtly breaks symmetry, can steer assembly into patterns that textbook models would never anticipate. By combining high-speed microscopy, machine learning to quantify order, and physics-based simulations, the study outlines a way to bring these hidden solvent and symmetry effects into future protein design tools. For anyone hoping to engineer reliable, hybrid bio‑inorganic materials, the message is clear: you must design not just for the protein and the surface, but for the structured layer of water that connects them.
Citation: Yadav Schmid, S., Helfrecht, B., Stegmann, A. et al. Impact of solvent forces and broken symmetry on the assembly of designed proteins at a liquid-solid interface. Nat Commun 17, 2446 (2026). https://doi.org/10.1038/s41467-026-69170-0
Keywords: protein self-assembly, liquid-solid interfaces, interfacial water structure, smectic ordering, bioinspired materials