Clear Sky Science · en
Structural basis of pausing during transcription initiation in mycobacterium tuberculosis
How a Germ’s Gene Switches Can Stall
Mycobacterium tuberculosis, the bacterium that causes tuberculosis, must constantly turn genes on and off to survive in the body’s harsh environment. The work described in this article peeks into one of the earliest steps of this genetic control, capturing a fleeting “pause” that happens just as the cell starts to read DNA into RNA. Understanding this pause helps explain how the microbe decides whether to fully switch a gene on or abort the attempt—and may eventually reveal new weak points for antibiotics.
Starting the Message From DNA
In bacteria, a large protein machine called RNA polymerase moves along DNA to make RNA, the first step toward producing proteins. To begin, it partners with helper proteins known as sigma factors that guide it to specific starting sites on DNA and help open a small “bubble” in the double helix. For Mycobacterium tuberculosis, one such helper, called sigma E, is especially important when the bacterium is under stress outside its inner cell membrane. Another factor, CarD, helps stabilize the open DNA bubble. Together, these players form an initiation complex that makes the first few building blocks of RNA before either committing to a full-length message or giving up and releasing short RNA fragments.
A Hidden Pause at Six or Seven Letters Long
Previous biochemical and single-molecule studies had hinted that RNA polymerase often pauses very early, when the new RNA strand reaches only six or seven building blocks in length. But those pauses are extremely brief and hard to see directly, so no high-resolution structures were available. In this study, the authors used cryo-electron microscopy to freeze and visualize multiple versions of the initiation machinery from M. tuberculosis. They engineered DNA–RNA scaffolds that trap the complex when the RNA is six or seven units long and solved structures for three main states: two “regular” initiation complexes and a distinct paused initiation complex.
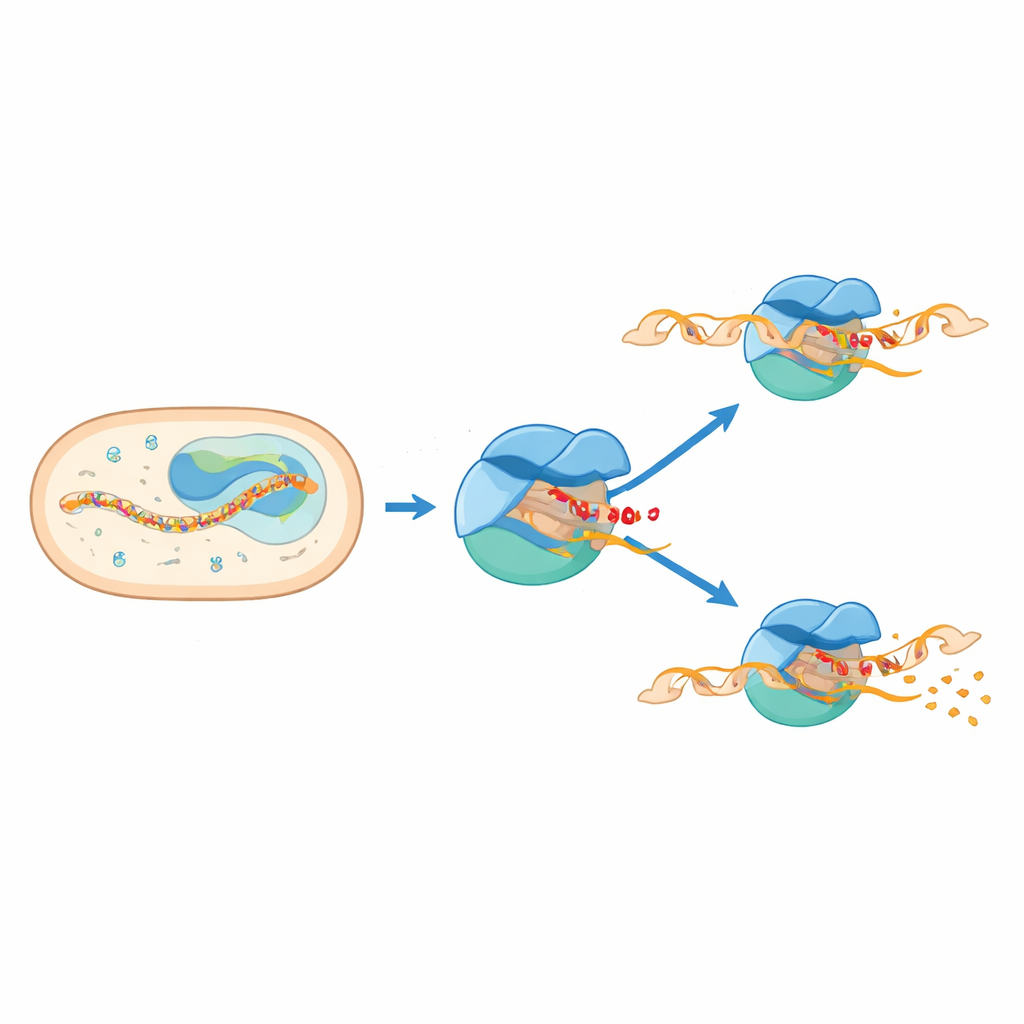
When Two Parts of the Machine Collide
The structures reveal that the pause arises when two parts of the machinery try to occupy the same space. A flexible segment of sigma E, known as the 3.2 region, juts into the channel where the growing RNA must pass to exit the enzyme. As the RNA lengthens to six or seven units, its front end collides with this protruding loop. That clash forces a swiveling motion of a large module within RNA polymerase and distorts a key structural element called the bridge helix. At the same time, DNA just upstream of the active site is pulled inward and partially unwound, producing a “scrunched” and kinked bubble. These coupled motions lock the complex into a paused, half-advanced state that stores mechanical stress.
A Shape-Shifting Helper Steadies the Bubble
CarD, the accessory factor, turns out to play a more versatile role than previously appreciated. In the non-paused initiation complex, CarD wedges into the minor groove of one DNA strand at the bubble edge, helping keep the bubble open so RNA polymerase can start synthesis. In the paused complex, CarD shifts to embrace the opposite, non-template strand of DNA, working together with sigma E to clamp the scrunched bubble more tightly. Biochemical tests show that CarD boosts sigma E–dependent transcription and influences how many RNA molecules accumulate at the six-to-seven-unit stage. These findings suggest that CarD both stabilizes the initial open state and helps the paused complex avoid collapsing prematurely.
A Checkpoint Between Failure and Commitment
By comparing high-resolution structures formed on partially pre-opened DNA with lower-resolution structures assembled on fully matched DNA, the authors show that the same basic paused shapes appear under more natural conditions. They propose that the collision between the growing RNA and the sigma 3.2 loop creates an energy-loaded checkpoint at six to seven RNA units. From this poised state, the complex can follow two main paths: it can release the short RNA and reset, an abortive attempt, or it can use the stored stress to break free from the promoter region and enter stable, long-term RNA production. CarD’s repositioning and the loosening grip of the sigma factor over upstream DNA appear to guide this decision.
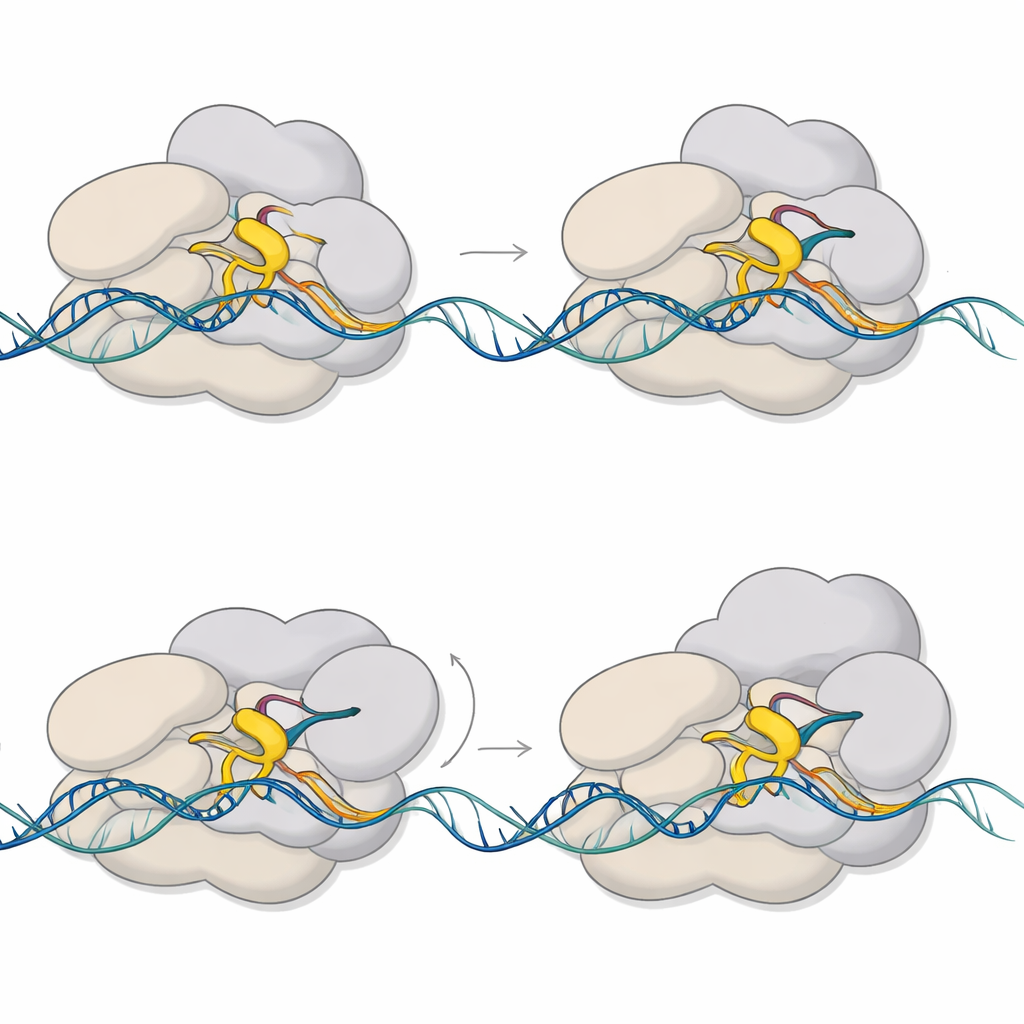
Why This Matters for Tuberculosis
For a lay reader, the take-home message is that the tuberculosis bacterium does not simply switch genes on like a light. Instead, it hesitates at a tiny RNA length, using a brief mechanical pause as a decision point between aborting or committing to full gene activation. This work captures that pause in atomic detail, showing how a small moving loop, a swiveling core, and a reshaped DNA bubble together create a controllable checkpoint. Because sigma E and CarD are central to how M. tuberculosis responds to stress, these insights could eventually help researchers design drugs that jam this early step and blunt the bacterium’s ability to adapt and survive.
Citation: Zheng, L., Xu, K. Structural basis of pausing during transcription initiation in mycobacterium tuberculosis. Nat Commun 17, 2197 (2026). https://doi.org/10.1038/s41467-026-69104-w
Keywords: transcription initiation, RNA polymerase pausing, Mycobacterium tuberculosis, sigma factors, cryo-EM