Clear Sky Science · en
Boosting the detection of enhancer-promoter loops via normalization methods for chromatin interaction data
Seeing the Hidden Loops in Our DNA
Inside every cell, long strands of DNA fold and loop in three dimensions, bringing distant stretches of the genome into close contact. Some of these loops physically connect on–off switches called enhancers with the genes they control, shaping how cells develop and how diseases like cancer arise. This article introduces a new computational method, Raichu, that makes these subtle regulatory loops much easier to detect in genome-wide experiments, opening a sharper window onto how DNA folding and gene activity are linked.
Why Tiny DNA Loops Matter
Our genomes are not arranged like straight strings of letters. Instead, they crumple into a complex 3D structure inside the nucleus. In this folded state, far-apart regions of DNA can touch, forming chromatin loops. Some loops act as scaffolds that help organize whole neighborhoods of the genome. Others are regulatory loops that connect gene promoters with distant enhancers that boost their activity. Disruption of these regulatory loops has been tied to developmental disorders and cancers, so scientists are eager to map them in detail.
The Challenge of Reading 3D Genome Maps
Techniques such as Hi-C and related methods capture millions to billions of DNA–DNA contacts and summarize them as heatmaps, where each pixel reflects how often two genome regions meet. However, these maps are riddled with technical quirks: some regions are easier to read than others because of sequence composition, how well they can be aligned, or how they are cut in the experiment. To clean things up, researchers use normalization methods like ICE and KR that balance the maps so every region appears to have equal visibility. While these tools sharpen large features such as broad domains and strong structural loops, the authors show that they unintentionally wash out the weaker, but biologically crucial, enhancer–promoter loops.
A New Way to Clean the Signal
Raichu takes a different approach to cleaning chromatin contact data. Instead of forcing every region of the genome to look equally visible, it models each interaction as the sum of three pieces: a general falloff in contact frequency with increasing distance along the chromosome, a bias specific to each location, and a remaining signal specific to that particular contact. Using an optimization algorithm, Raichu estimates the bias pattern that best explains the observed data while keeping the overall distance-dependent trend fixed. The contacts are then adjusted by dividing out these bias values. This preserves the natural global decay of interactions while selectively correcting technical distortions, leaving behind clearer signals of real, specific DNA contacts.
Uncovering Thousands of Missed Regulatory Loops
When the authors applied Raichu to deeply sequenced human and mouse datasets, it revealed nearly twice as many chromatin loops as standard methods, while still recovering almost all previously known loops. The extra loops detected by Raichu were strongly enriched for biochemical marks of active enhancers and promoters and for binding of transcription factors that control gene activity. Many of these loops were independently supported by other 3D genome technologies and even by high-resolution imaging, confirming that they reflect real physical proximity in the nucleus. Importantly, Raichu maintained this advantage even when the data were thinned to simulate lower sequencing depths, when studying specialized mapping methods like Micro-C and region capture Micro-C, and even when pooling small numbers of single cells.
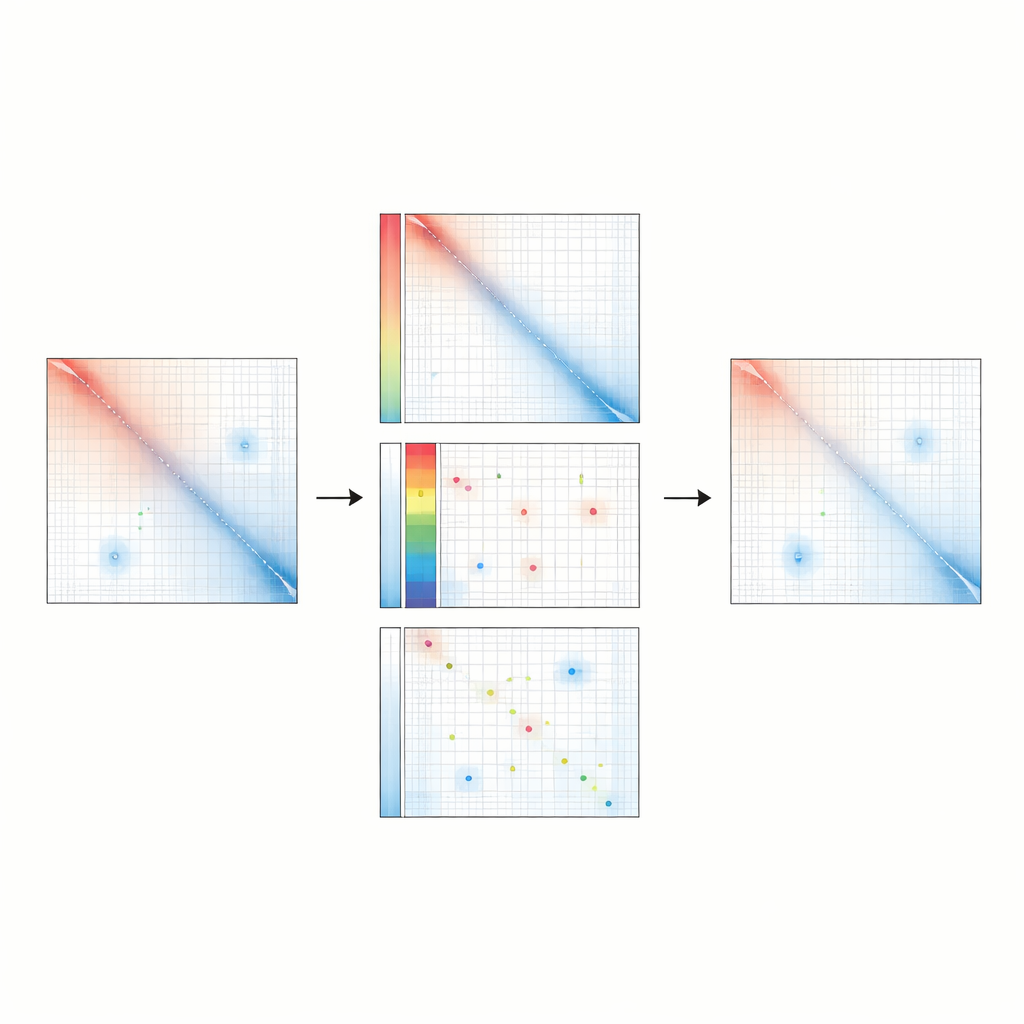
Revealing Subtle Changes and Evolutionary Patterns
Because Raichu is especially sensitive to regulatory loops, it is better at spotting meaningful differences between conditions that might otherwise appear similar. In an engineered human cell model carrying a leukemia risk variant, Raichu uncovered new loops that emerged only in the risk state, linking key genes to their enhancers and aligning with shifts in gene activity. Across mouse and human neural progenitor cells, Raichu detected thousands of enhancer–promoter loops that were conserved between species and often connected distant enhancers to genes involved in brain development. These findings suggest that many important regulatory contacts have been hiding in plain sight, masked by earlier normalization approaches.
What This Means for Future Genomic Research
To a lay reader, the key message is that how we process big genomic datasets can dramatically influence what biology we see. By rethinking the cleanup step for 3D genome maps, Raichu restores faint but important signals that link gene switches to their targets. This makes it easier to trace how DNA folding controls gene activity in health and disease, from single cells to whole tissues and across species. As more studies adopt Raichu, researchers can expect richer maps of enhancer–promoter communication and a clearer picture of how changes in genome architecture contribute to development, cancer, and other complex conditions.
Citation: Wang, X., Shi, D., Xue, F. et al. Boosting the detection of enhancer-promoter loops via normalization methods for chromatin interaction data. Nat Commun 17, 2299 (2026). https://doi.org/10.1038/s41467-026-69082-z
Keywords: 3D genome organization, enhancer promoter loops, Hi-C data analysis, chromatin normalization, gene regulation