Clear Sky Science · en
FidlTrack: high-fidelity structure-aware single particle tracking resolves intracellular molecular motion in organelles sensing APP processing
Watching Single Molecules Inside Living Cells
Inside every cell, countless molecules are constantly on the move, bumping, binding, and breaking apart. Modern microscopes can follow single molecules one by one, promising a detailed view of how life works at its smallest scales. But when many molecules move quickly in tight, winding spaces such as the endoplasmic reticulum or mitochondria, their paths become tangled and easy to misread. This paper introduces FidlTrack, a new way to clean up that tangle so researchers can trust what they see when they track single molecules in real time.
Why Tracking Tiny Movements Is So Hard
Single particle tracking works by following the same bright dot across many movie frames. On a relatively flat cell surface, where molecules move slowly and are spaced apart, this is manageable. Deep inside the cell, however, molecules diffuse far more quickly and crowd into narrow, looping tubes and sheets. Between frames, the same dot might have moved a long distance, and several identical dots may lie within reach. Traditional software must “guess” which dot in the next frame is the same molecule, and those guesses can easily be wrong. Worse, correct and incorrect paths can look very similar, so it is hard to know which parts of the data to trust.
Designing Experiments for Honest Data
The researchers first built a realistic simulator that generates “ground truth” movies of diffusing molecules, where the true paths are known. They used these synthetic datasets to systematically test how key factors affect tracking reliability: how fast molecules move, how many appear in each image, how often images are collected, and how far a molecule is allowed to jump between frames. From this, they created maps that show, for any given situation, which settings maximize the fraction of correctly reconstructed movements. These maps reveal that for slow-moving molecules at moderate densities, tracking can be very reliable, but for fast-moving molecules at high densities, errors grow quickly and adding more molecules no longer improves the useful information.
Spotting and Cutting Out Dubious Steps
Next, the team tackled a subtle but powerful idea: ambiguity. A movement is called ambiguous when more than one possible next dot lies within range, forcing the algorithm to choose among several plausible links. Using their simulations, the authors showed that a large share of tracking mistakes arise from these ambiguous steps. They defined an Ambiguity Score that counts how often such situations occur, and then explored what happens if all ambiguous steps are removed after tracking. This pruning sacrifices some data and shortens paths, but it sharply boosts the overall reliability of what remains and improves estimates of how quickly molecules move. When applied to real movies of a marker in the endoplasmic reticulum, ambiguity was higher in crowded regions near the cell nucleus, and removing ambiguous links cleaned up those areas without harming simpler regions.
Letting Cell Architecture Guide the Tracking
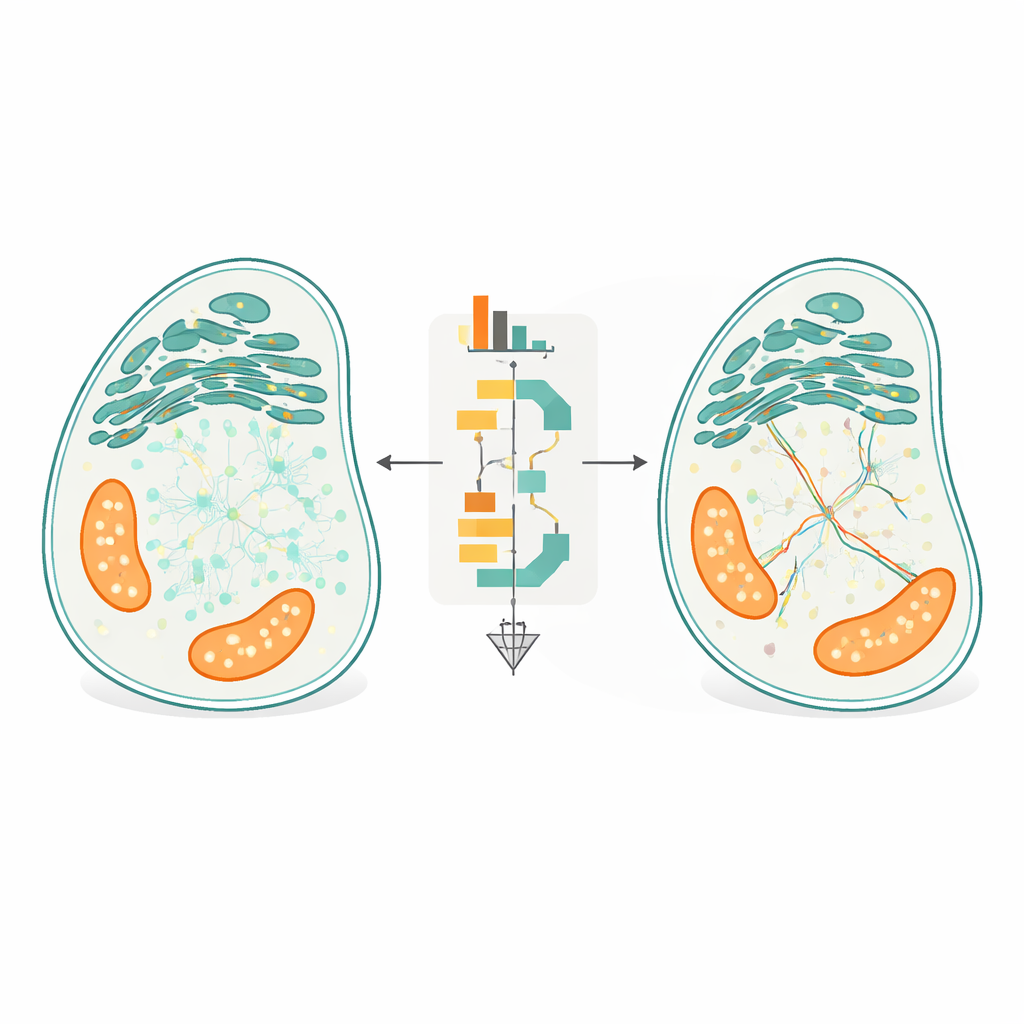
The central innovation of FidlTrack is “structure-aware” tracking. Instead of treating each dot as moving in empty space, the method uses images of the cell’s internal architecture—such as the outlines of the endoplasmic reticulum, mitochondria, or slender neuronal projections—to constrain where molecules can realistically travel. The organelle image is converted into a graph of connected pixels, and distances are measured along this graph rather than straight lines through space. Links that would require a molecule to jump across a gap between two separate tubules can thus be flagged as impossible and rejected. In simulations of tightly packed tubes and in real movies of neuronal processes and organelles, this structural awareness reduced ambiguous linkages by up to half and increased the amount of trustworthy, non-ambiguous motion data several-fold.
Revealing Hidden Cell Behaviors and Disease-Relevant Events
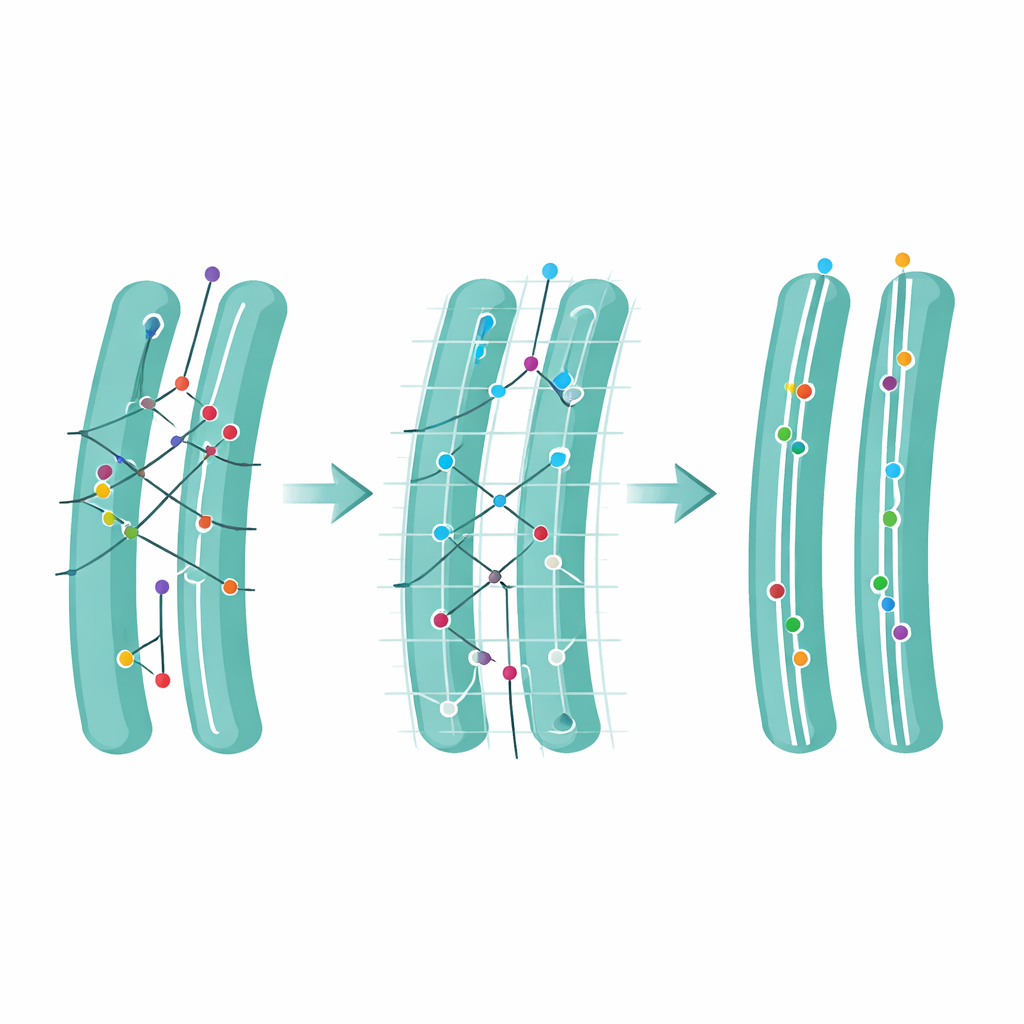
Armed with these tools—optimized settings, ambiguity filtering, and structure-awareness—the authors revisited several biological questions that had previously been just out of reach. In the endoplasmic reticulum, they could cleanly track how proteins move as they engage with exit sites that funnel cargo toward the Golgi apparatus, distinguishing brief “fly-bys” from longer stays. They captured rare moments when the Alzheimer’s-related protein APP is cut by the enzyme BACE1, visible as a sudden switch from slow, membrane-tethered motion to faster free diffusion. They also followed engineered antibody-like molecules in the ER and, from changes in their motion, inferred when they were bound to their target versus freely drifting. Across these diverse cases, FidlTrack recovered more reliable trajectories and sharpened differences that standard tracking either blurred or underestimated.
What This Means for Future Cell Biology
For non-specialists, the key message is that not all single-molecule tracks are created equal: some are trustworthy, others are misleading, and until now it has been hard to tell them apart. FidlTrack provides a practical, open-source way to both measure how reliable a given dataset is and to improve that reliability by tuning experimental settings, cutting out ambiguous steps, and using the cell’s own geometry as a guide. This makes it possible to watch molecules navigate the cell’s intricate interior with greater confidence and to detect rare or subtle events, from protein sorting to disease-relevant processing, that were previously buried in noise.
Citation: Parutto, P., Yuan, Y., Davì, V. et al. FidlTrack: high-fidelity structure-aware single particle tracking resolves intracellular molecular motion in organelles sensing APP processing. Nat Commun 17, 2639 (2026). https://doi.org/10.1038/s41467-026-69067-y
Keywords: single particle tracking, intracellular dynamics, organelle structure, protein motion, Alzheimer’s disease