Clear Sky Science · en
Reliable repurposing of the antibody interactome inside the cell
Turning Antibodies into Tiny Tools Inside Our Cells
Antibodies are famous as the bloodstream’s precision weapons, homing in on viruses and other threats. But most of the action in many diseases, including Alzheimer’s, Parkinson’s and some cancers, happens inside cells, where regular antibodies rarely reach or work well. This study shows how to systematically rebuild existing antibodies into smaller “intrabodies” that function reliably inside living human cells, opening the door to new diagnostics, research tools and potentially future therapies that operate directly at the heart of disease processes.
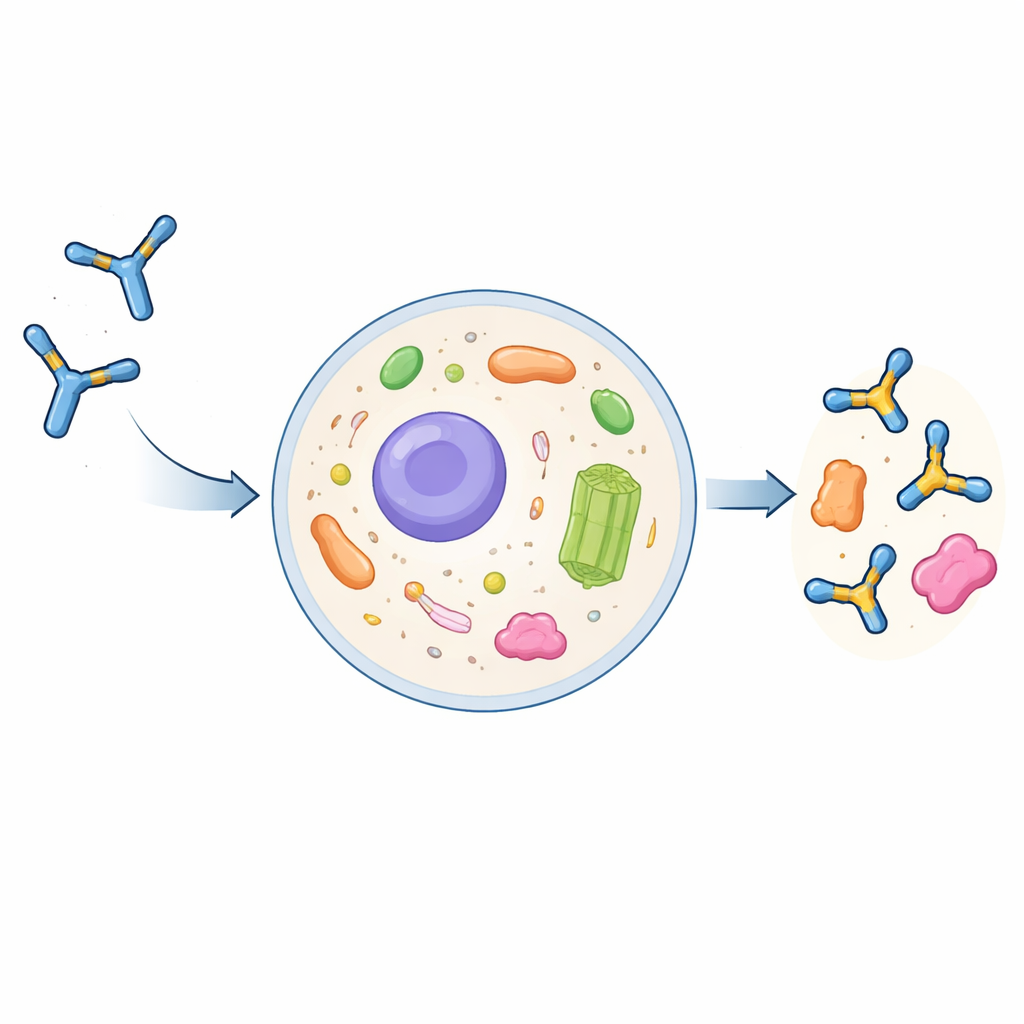
Why Regular Antibodies Struggle Inside Cells
Full-sized antibodies are large, Y‑shaped proteins evolved to operate outside cells. They are assembled and chemically modified in a cell’s secretory pathway and then released into the bloodstream. If they are taken back up by cells, they usually end up in degradation compartments rather than in the fluid interior of the cell, the cytoplasm. Scientists can trim antibodies down to their essential recognition parts, called single‑chain variable fragments (scFvs), and express these as intrabodies inside cells. However, these intrabodies often clump together, become insoluble, and lose their ability to bind their targets. Until now, there has been no general, dependable recipe for turning a conventional antibody into a well‑behaved intrabody.
Finding the Design Rules for Soluble Intrabodies
The researchers started by measuring how well 45 different intrabodies stayed dissolved when produced in human cells. They systematically compared many protein features—such as overall hydrophobicity, predicted structure quality and the properties of the binding loops—to see which ones best explained whether an intrabody remained soluble. One simple factor stood out: overall electrical charge at near‑physiological pH. Intrabodies with a strongly negative net charge were much more likely to remain soluble, whereas those with neutral or positive charge tended to aggregate. Other sophisticated predictors, including structure confidence scores from the AlphaFold system, did not correlate with solubility. This allowed the team to build a practical, charge‑based model that can estimate how much of an intrabody will be found in the soluble fraction of a cell extract.
Re‑Engineering Linkers and Domains for Better Behavior
Armed with this insight, the team focused on features they could easily tune without disturbing how an intrabody recognizes its target. A key element is the short peptide “linker” that connects the two variable domains. Most existing designs use neutral linkers that do not alter charge. By swapping in negatively charged amino acids at these positions, the researchers could drive the overall charge of the intrabody down, dramatically improving solubility while preserving target binding. They also discovered that the order of the variable domains (whether the heavy part or the light part comes first) and specific framework positions outside the binding loops influence how easily the protein folds and stays dissolved. Choosing a domain order that protects more fragile regions and introducing carefully placed charge‑swapping mutations further boosted performance.
Using Artificial Intelligence to Build Intrabodies at Scale
To move beyond one‑by‑one trial‑and‑error, the authors combined their simple charge rules with advanced protein design software based on deep learning. Starting from known antibody backbones, they used an "inverse folding" approach to suggest new sequences that would still adopt the desired shape but carry surface features—especially charge patterns—that favor stability and solubility. They restricted changes at the key binding loops to preserve recognition, while allowing the algorithm to redesign surrounding surfaces. Testing showed that many of these AI‑guided intrabodies were not only highly soluble but also more thermally stable than human‑designed variants, and they retained the ability to bind disease‑relevant targets such as misfolded SOD1, p53, α‑synuclein and other proteins linked to neurodegeneration.
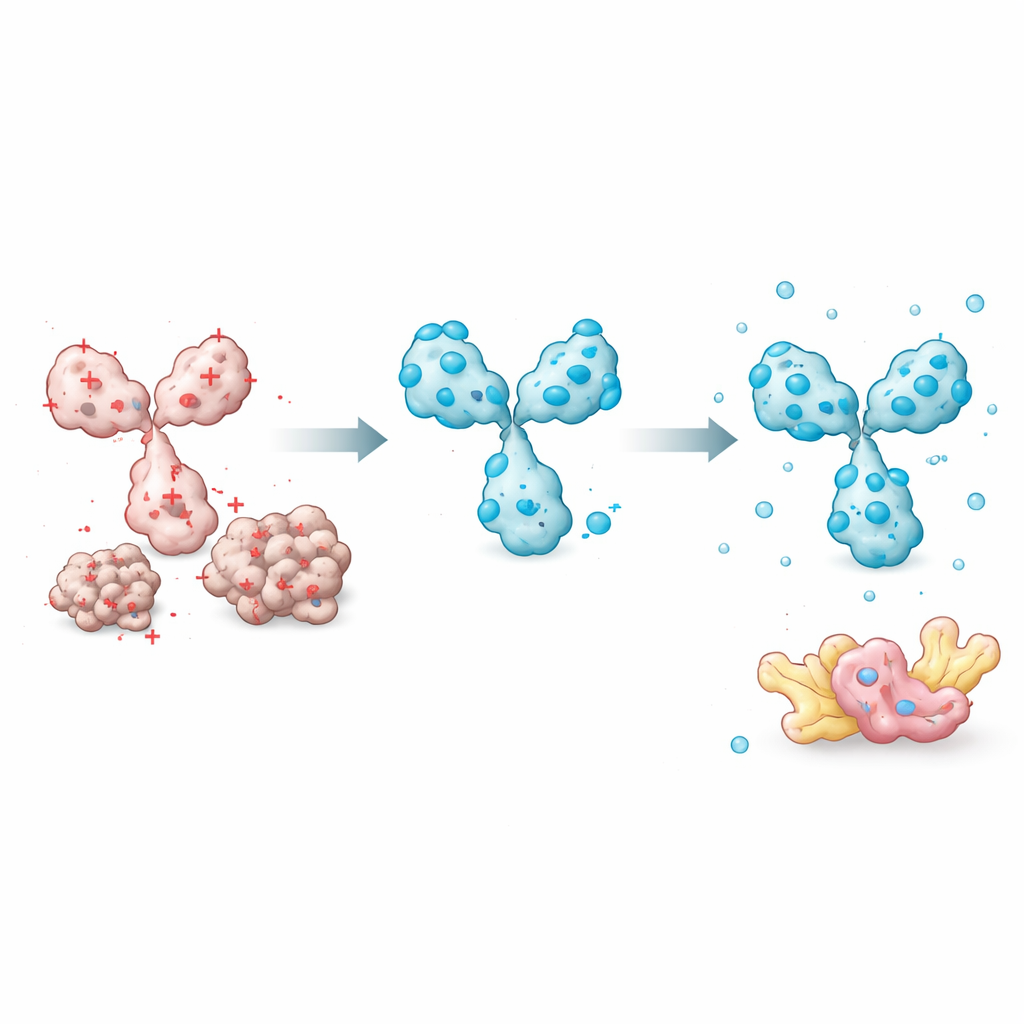
Building a Library of Inside‑Cell Protein Catchers
Applying these rules and tools at large scale, the team computationally reformatted 672 antibodies into intrabodies predicted to work well inside cells. These engineered molecules cover more than 60 different intracellular targets, including proteins involved in Alzheimer’s, Parkinson’s and Huntington’s diseases, as well as cancer‑related factors and post‑translational modifications like phosphorylation and acetylation. In experiments, selected intrabodies successfully fished out specific disease‑linked protein forms, such as mutant SOD1 or particular shapes of α‑synuclein, while ignoring normal versions. All sequences are released openly, and the authors provide a web tool that lets others estimate intrabody solubility from sequence using the same charge‑based model.
What This Means for Future Medicines and Research
In practical terms, this work turns the vast existing collection of antibodies—many already studied in the clinic—into a toolkit that can be reliably adapted for use inside cells. By following a straightforward set of design rules, researchers can now create intrabodies that are likely to work on the first attempt, saving time and cost. For a layperson, the key message is that scientists have found a simple way to "rewire" our best protein detectors so that they function in the crowded interior of living cells, where many diseases start. This could speed the development of new ways to track, study and eventually treat conditions driven by misfolded or malfunctioning proteins.
Citation: O’Shea, C.M., Shahzad, R., Aghasoleimani, K. et al. Reliable repurposing of the antibody interactome inside the cell. Nat Commun 17, 2222 (2026). https://doi.org/10.1038/s41467-026-69057-0
Keywords: intrabodies, antibody engineering, protein solubility, neurodegenerative disease, artificial intelligence in biology