Clear Sky Science · en
Virus glycoprotein nanodisc platform for vaccine analytics
Turning Virus Proteins into Measurable Targets
Modern vaccines increasingly rely on precisely engineered versions of the proteins that stud the surface of viruses. These proteins are the main targets for protective antibodies, yet they are notoriously hard to study in their natural, membrane-bound form. This paper describes a new laboratory platform that places full-length viral surface proteins into tiny, flat lipid particles called nanodiscs. This setup lets scientists measure how well vaccine candidates mimic real viruses, how antibodies latch onto them, and how to redesign these vaccines for better protection.
Why Recreating the Viral Surface Matters
Many of the most important viral proteins sit anchored in a fatty membrane, just as they do on an actual virus. Traditional lab approaches trim off the membrane part to make these proteins easier to produce and purify in solution. But that shortcut removes regions near the membrane that are home to especially valuable antibody targets, including a segment of HIV known as the membrane-proximal external region (MPER). These near-membrane regions can look and behave differently when they are not embedded in a realistic lipid environment, which means soluble protein fragments may give an incomplete or misleading picture of how a vaccine will perform in the body.
Building a Versatile Nanodisc Platform
The researchers created a streamlined, five-day workflow to harvest full-length viral surface proteins from human cells, keep their natural membrane-spanning segments intact, and reinsert them into a controlled lipid disc. They first expressed engineered HIV and Ebola glycoproteins on cell surfaces, then gently extracted them with detergent and captured them on a purification matrix. While the proteins were still attached, they mixed in defined lipids and a belt-like scaffold protein that self-assembles into a small, disc-shaped membrane. As detergent was removed, the viral proteins settled into these nanodiscs. The resulting preparations were highly pure, stable for months in the refrigerator, and carried glycan (sugar) coatings closely matching those found on native viral spikes. 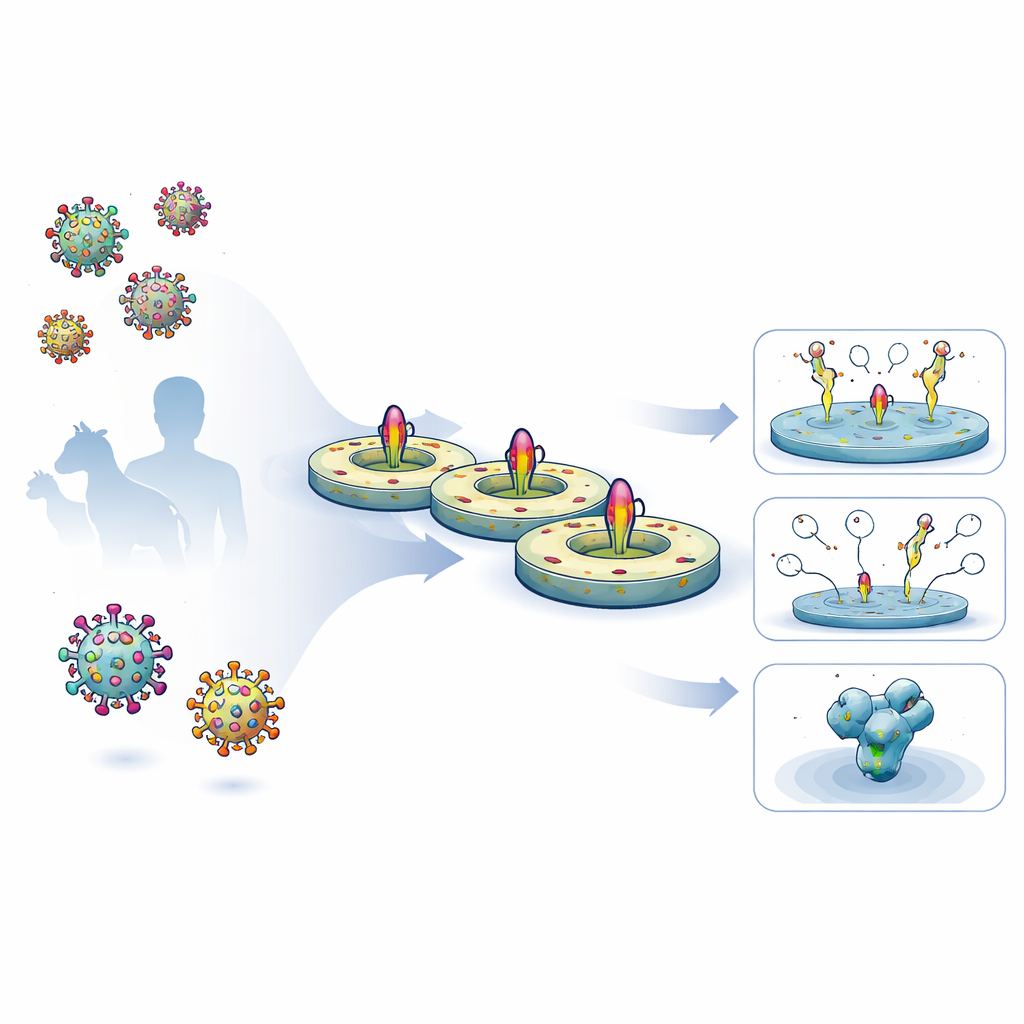
Reading Antibody Binding and Immune Responses
With nanodisc-embedded proteins in hand, the team tested how well different antibodies could bind under several surface plasmon resonance (SPR) setups, a technique that tracks binding in real time. Using HIV nanodiscs designed to expose the MPER region, they showed that a potent broadly neutralizing antibody called 10E8 bound about 70 times more tightly to the engineered construct than to an earlier design, mostly because it let go far more slowly. Control mutations that disrupted the MPER region eliminated 10E8 binding altogether, confirming that the platform can sensitively report how design changes affect key epitopes. The same nanodiscs worked as probes in flow cytometry, where they helped fish out B cells from immunized mice and monkeys that recognized the full transmembrane form of the HIV protein, including parts that are absent from standard soluble probes. 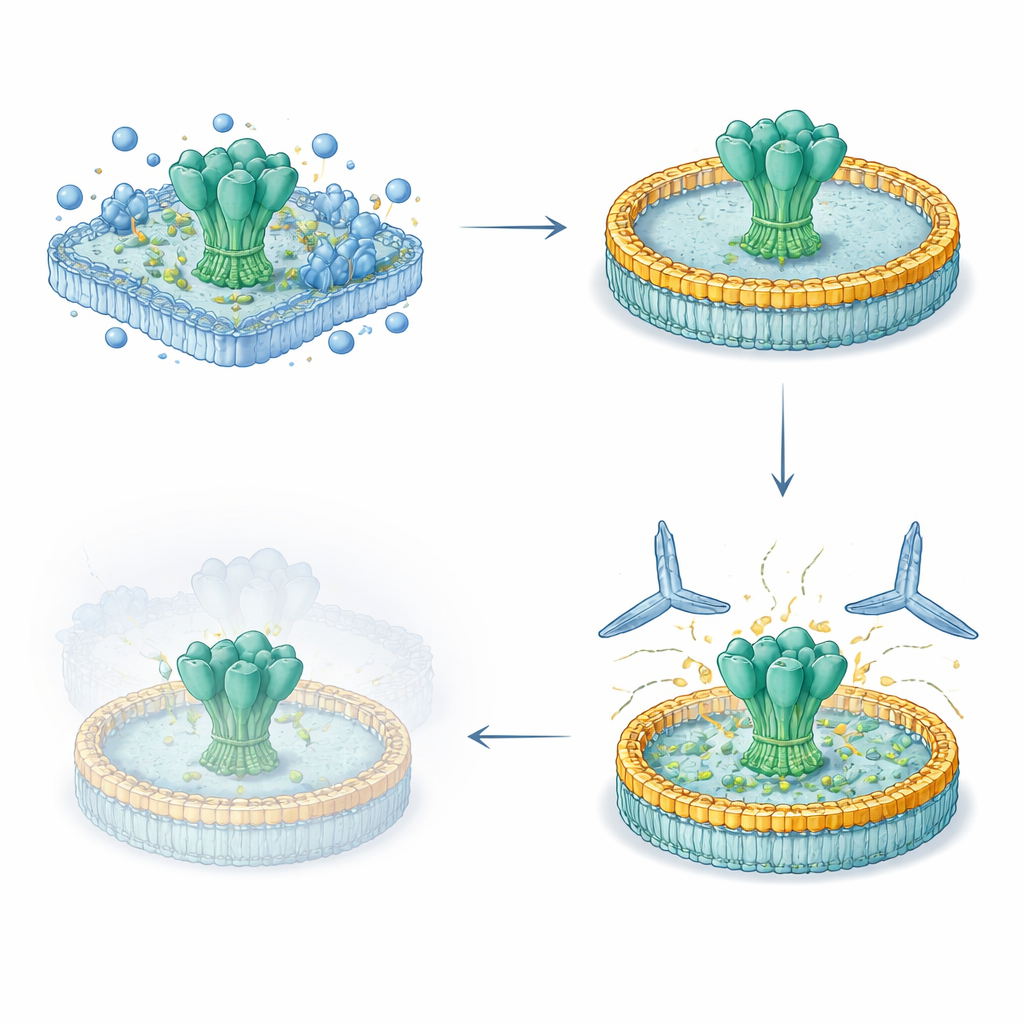
Seeing the Full Target in Atomic Detail
To understand exactly how antibodies recognize the MPER in its native setting, the authors used cryogenic electron microscopy to image the engineered HIV nanodiscs bound to three different broadly neutralizing antibodies. They obtained a 3.5-angstrom structure of one antibody, 10E8, engaging the MPER segment while the rest of the HIV spike remained anchored in the lipid disc. This revealed a continuous network of contacts between 10E8 and both the MPER and nearby portions of the outer protein, including a deep pocket at the junction between two subunits. By comparing this structure with previous images of 10E8 bound only to a short peptide, the team showed that the membrane environment and the full-length spike allow extra contacts and dynamic movements that were invisible in simpler models, and that specific amino acids in the virus strongly influence how easily 10E8 can neutralize it.
Broad Uses for Smarter Vaccines
In practical terms, this nanodisc platform gives vaccine designers a way to evaluate transmembrane vaccine candidates with the same powerful analytical tools long reserved for soluble protein fragments. It works across different viruses, supports detailed binding measurements, enables precise sorting and sequencing of vaccine-induced B cells, and yields high-resolution structures that capture realistic membrane-proximal epitopes. For a lay reader, the takeaway is that scientists now have a more lifelike testbed for viral surface proteins, one that can show which vaccine designs truly look and act like the real virus and how to tweak them for stronger, broader antibody responses. This should speed the development of next-generation vaccines not only for HIV but for many other enveloped viruses.
Citation: Rantalainen, K., Liguori, A., Ozorowski, G. et al. Virus glycoprotein nanodisc platform for vaccine analytics. Nat Commun 17, 2561 (2026). https://doi.org/10.1038/s41467-026-68985-1
Keywords: nanodiscs, HIV vaccines, viral glycoproteins, broadly neutralizing antibodies, cryo-EM