Clear Sky Science · en
Radical footprinting in mammalian whole blood
Seeing Proteins at Work in Real Blood
Proteins in our blood constantly change shape as they carry out vital jobs like fighting infection, moving iron, and responding to disease. Until now, most tools for examining these tiny shape shifts have worked only in simplified lab samples or isolated cells. This study shows, for the first time, that scientists can read protein shapes directly in intact mammalian blood, opening a path to watch disease processes as they really happen in the body.
A New Way to Map Protein Shapes
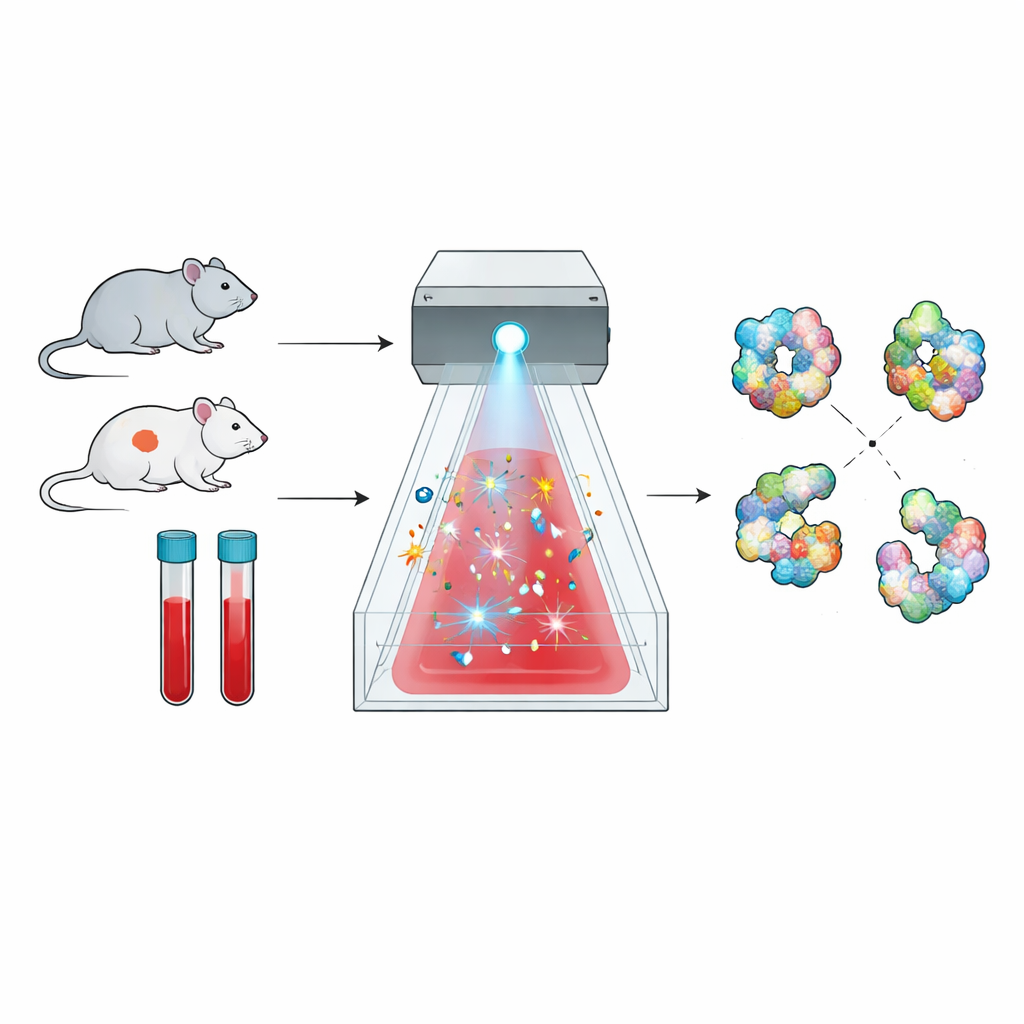
Proteins are not rigid beads; they fold into intricate three-dimensional forms, and those forms determine what they can do. The method explored here, called radical protein footprinting, takes advantage of this fact. Brief bursts of highly reactive molecules act like a chemical “camera flash,” striking only the parts of a protein that are exposed to the surrounding liquid. Later, mass spectrometry is used to count where those chemical hits occurred, producing a kind of surface map, or footprint, of each protein’s shape. Changes in that footprint between healthy and diseased states reveal subtle shifts in how the protein is folded or how it interacts with partners.
Making the Technique Work in Whole Blood
Applying this approach directly to blood has been a long-standing challenge. Blood strongly absorbs ultraviolet light and is packed with enzymes, such as catalase, that quickly destroy the usual radical-generating chemicals before they can mark the proteins. The researchers solved this by switching to sodium persulfate, which can be split by a bright broadband flash of light into powerful sulfate radicals. Using a commercial system called FOX, they showed that persulfate can be activated reliably and that a built-in dosimetry readout can track how many radicals are produced, allowing careful control of the “exposure” each sample receives.
Protecting Cells While Capturing Detail
Because this method is meant to study proteins in something close to their natural state, it was crucial that the blood cells themselves were not destroyed. Tests on mouse blood showed that adding concentrated persulfate caused only minor, reversible changes to red blood cell shape and less than two percent cell rupture, similar to simple salt solutions. The team also developed an improved “quench” mixture that rapidly soaks up leftover reactive species, preventing spurious damage from slower side reactions after the flash. Together, these refinements allowed them to label proteins in intact mouse blood while preserving cell structure and keeping background noise very low.
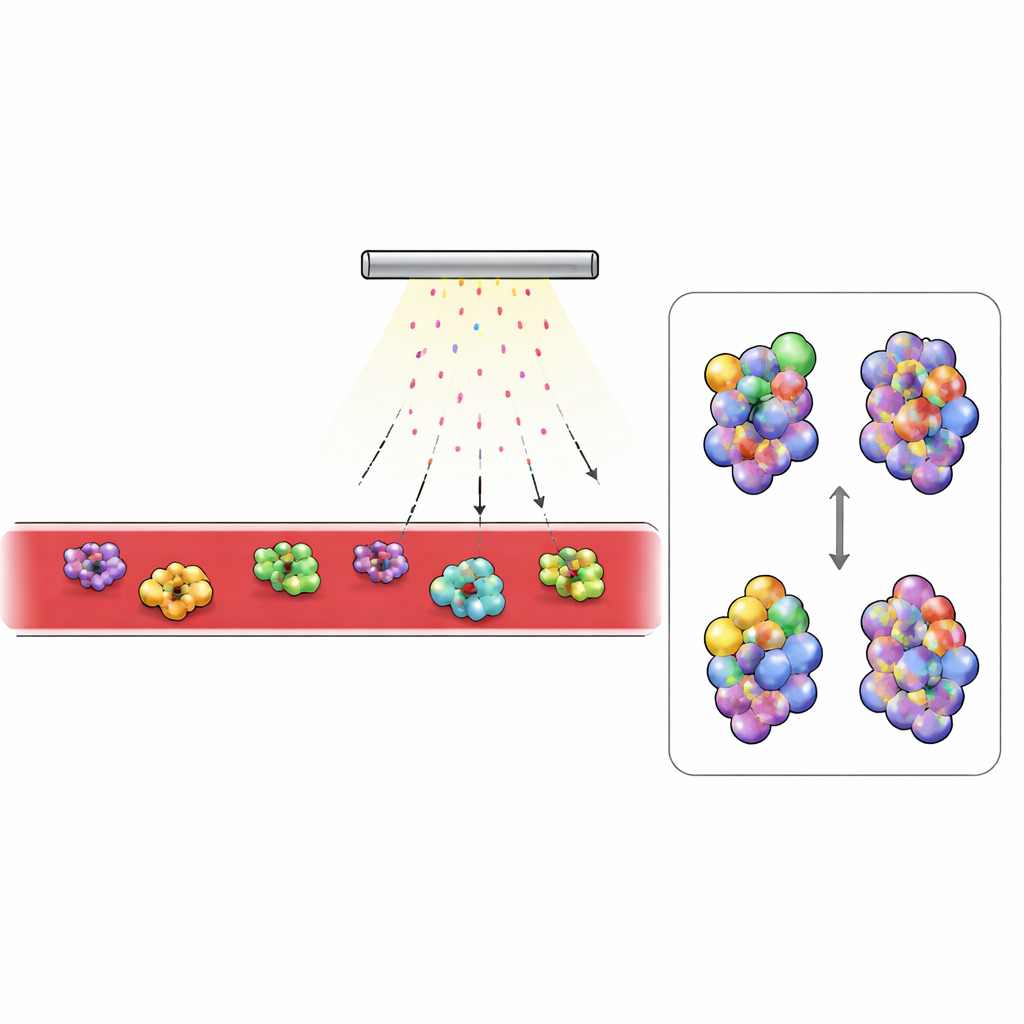
What Changes in Diabetic Blood
With this platform in hand, the researchers compared blood from healthy mice and from a common model of type 2 diabetes. They focused on the most abundant proteins detected, especially those circulating outside cells. Extracellular proteins showed much heavier labeling than those inside cells, reflecting limited penetration of persulfate into cell interiors. Two blood proteins stood out: complement C3, a key component of the immune system, and transferrin, which transports iron. In diabetic mice, regions of C3 that become buried when the protein switches into its active form were less labeled, while normally hidden regions were more exposed. This pattern matched the known structural change when C3 is converted to its active fragment, C3b, and blood tests confirmed that diabetic animals had far more activated C3. In transferrin, areas near the iron-binding site were more protected in diabetic blood, consistent with the protein carrying more iron. Independent measurements showed higher serum iron and a greater fraction of transferrin saturated with iron in diabetic mice.
Why This Matters for Health and Medicine
To a non-specialist, the essential message is that the authors have created a way to “feel” the shapes of many proteins at once directly in real blood, then use those shape patterns to infer how disease is altering the body’s chemistry. In this mouse model of type 2 diabetes, the method reveals overactive complement signaling and heavier iron loading on transferrin—changes that classical tests might miss or see only indirectly. Because the approach works in small blood samples and uses a standardized light source, it could eventually help identify early structural warning signs of disease, track how biological drugs behave in the bloodstream, and guide the design of more effective treatments.
Citation: Zhao, M., Tobin, L., Misra, S.K. et al. Radical footprinting in mammalian whole blood. Nat Commun 17, 2470 (2026). https://doi.org/10.1038/s41467-026-68982-4
Keywords: structural proteomics, blood proteins, type 2 diabetes, complement system, iron metabolism