Clear Sky Science · en
Protein C-terminal variations impact proteostasis
How the Tail End of Proteins Shapes Health
Every protein in our cells ends with a tiny “tail” of just a few building blocks. These tails may look like an afterthought, but this study reveals they quietly decide which proteins linger and which are quickly removed. Because protein levels underlie cancer, genetic disorders, and how cells respond to stress, understanding these tails offers new ways to read disease risk and even design future treatments.
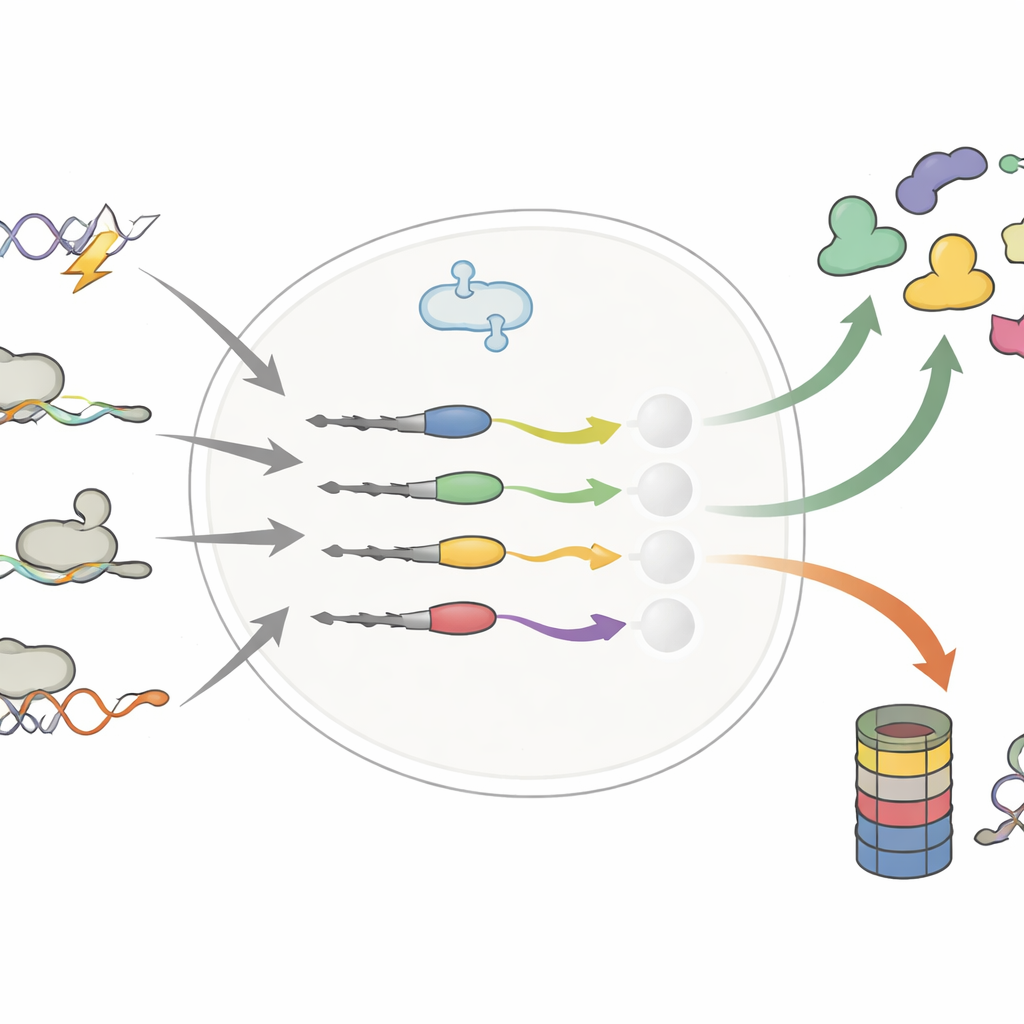
Small Changes at the Protein Tail, Big Shifts in Lifetime
The authors focused on the far end of proteins—the C-terminus—where even a few extra or swapped building blocks can appear through DNA mutations, alternative RNA splicing, or ribosomes reading past the usual stop sign. Earlier work suggested that such “erroneous” tails mainly act as disposal tags, sending faulty proteins to the cell’s shredder. By systematically comparing thousands of normal and altered tails, this study overturns that simple view. The team shows that altered tails can either shorten or lengthen a protein’s life, sometimes stabilizing cancer drivers or destabilizing tumor blockers.
When Genetic Errors Help or Hurt Proteins
One major source of tail changes is nonstop mutations, which erase a normal stop signal so the ribosome continues into a stretch of RNA that usually remains silent. The researchers tested over 3,000 disease-linked nonstop variants using a dual-color reporter that measures how strongly each tail protects or weakens a model protein. On average, mutant tails still made proteins less stable than their normal counterparts, but a substantial fraction did the opposite: roughly one in ten nonstop mutations stabilized proteins, while almost a third destabilized them. These shifts affected well-known oncoproteins and tumor suppressors, implying that what seemed like subtle mutations can in fact tip the balance of cell growth and death.
Built-In Variety from Splicing and Readthrough
Not all tail differences are accidents. Our genes routinely produce multiple protein versions through alternative splicing and programmed stop-codon readthrough. By mining human gene catalogs, the authors found that nearly two-thirds of genes make proteins with more than one tail design, and many differ only in the final few building blocks. Testing selected examples showed that these near-identical versions often have clearly different half-lives. Some readthrough-generated tails made proteins more stable, others less so. This suggests that cells deliberately use tail variation as a fine-tuning knob, adjusting how long specific protein forms persist in different tissues or conditions.
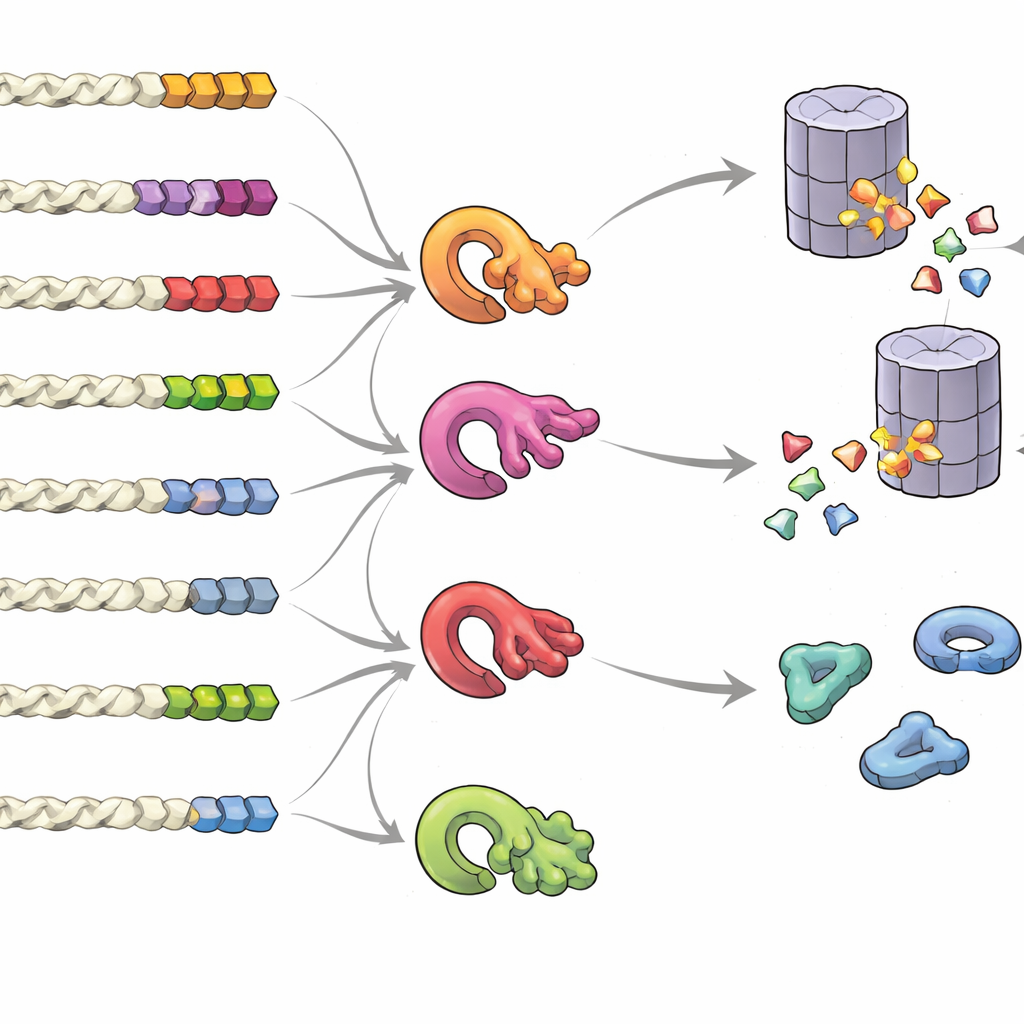
What Makes a Tail a “Keep Me” or “Trash Me” Signal
To understand what features of a tail sway protein fate, the team created libraries of random tails and measured their impact in bulk. They found that it is not just one fixed “motif” but overall composition, position, and clustering of certain building blocks that matter. Tails rich in small or oily components, or the rarer cysteine and tryptophan, tended to invite degradation, especially when these residues clustered. In contrast, tails enriched in acidic components favored stability. Looking across many species, normal protein tails showed clear signs of evolutionary pressure to avoid risky patterns, while error-prone readthrough or frameshift tails were biased toward destabilizing compositions. The group also discovered that multiple quality-control enzymes—different ubiquitin ligases—specialize in recognizing distinct hydrophobic tail patterns, forming an intricate network that decides which proteins are destroyed.
Why This Matters for Medicine and Biotechnology
The work reframes protein tails from passive endpoints into active regulators of protein abundance. For a layperson, this means that many diseases may arise not only from broken proteins but also from subtle changes in how long proteins stick around, driven by tiny differences at their ends. It also warns that therapies that deliberately promote readthrough of stop signals, used to treat certain genetic disorders, could unintentionally alter the lifetimes of many normal proteins. At the same time, the findings suggest powerful new strategies: by engineering tail sequences or nudging cells to favor particular splice forms or readthrough events, scientists may one day dial protein levels up or down with remarkable precision.
Citation: Chu, CY., Hsu, SY., Yeh, CW. et al. Protein C-terminal variations impact proteostasis. Nat Commun 17, 2288 (2026). https://doi.org/10.1038/s41467-026-68979-z
Keywords: protein stability, C-terminal tail, protein degradation, genetic mutations, proteostasis