Clear Sky Science · en
RiboBright reveals cell-type-specific differences in ribosome organization and movement
Seeing the Cell’s Protein Factories Light Up
Every second, each of our cells churns out thousands of new proteins using tiny machines called ribosomes. Until now, watching where these machines sit and how they move inside living cells has been surprisingly difficult. This study introduces RiboBright, a small fluorescent molecule that latches onto ribosomes and lights them up, allowing scientists to track these protein factories one cell at a time. The work reveals that ribosomes are organized and used very differently across cell types, especially as stem cells choose their future fates.
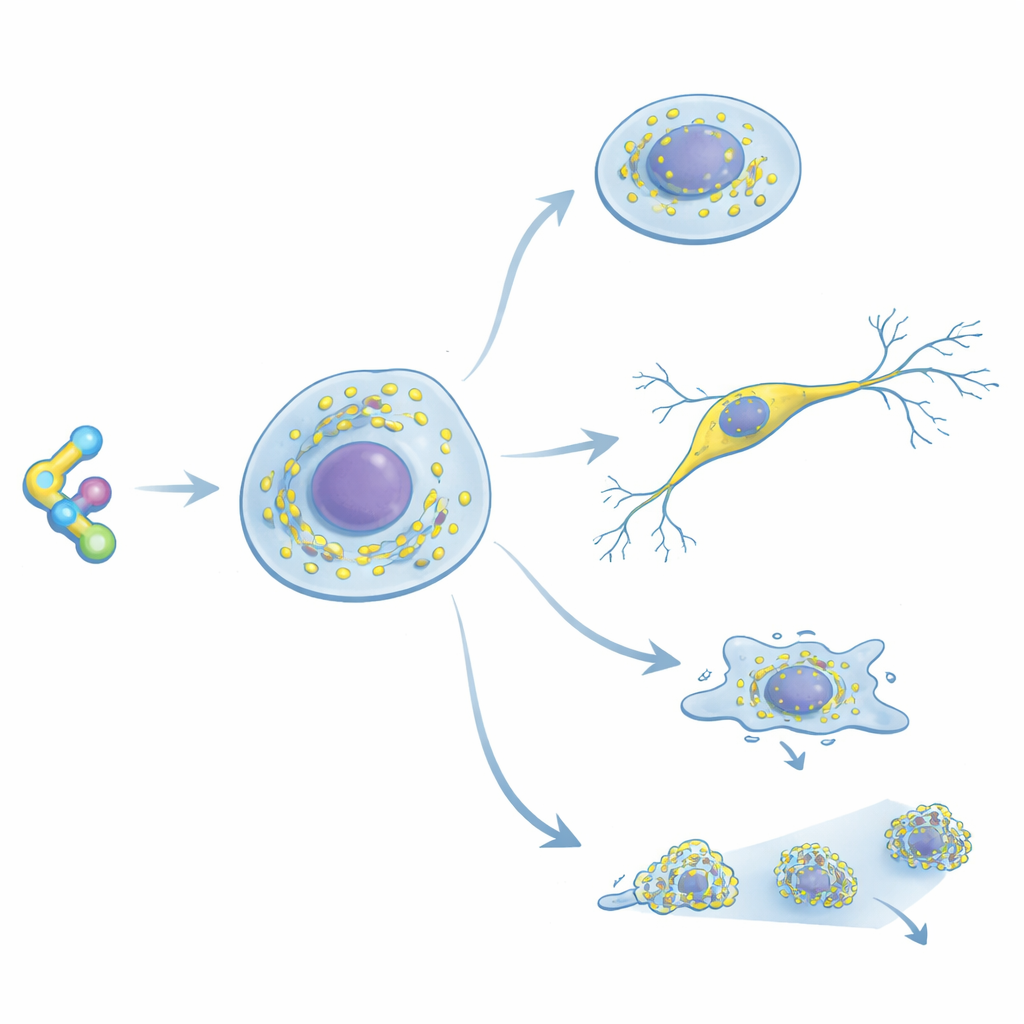
A New Way to Highlight Ribosomes
The researchers built RiboBright by modifying cycloheximide, a classic drug known for binding strongly and selectively to ribosomes in animals and humans. They attached a special “molecular rotor” dye at a position on the drug that does not weaken its grip on the ribosome. This dye barely glows when it spins freely in solution, but becomes bright when its motion is restricted—such as when the probe wedges into the ribosome. Tests in test tubes showed that the modified molecules still shut down protein production at nearly the same doses as the original drug, confirming they bind well. In living cells, one version in particular produced sharp, bright spots, indicating it had the right balance between tight ribosome binding and strong light emission; this optimized probe was named RiboBright.
Proving the Probe Targets Real Ribosomes
To be sure RiboBright truly marks ribosomes, the team combined several complementary approaches. When cells were pre-treated with either cycloheximide itself or another, unrelated ribosome-blocking compound, the RiboBright signal nearly vanished, implying all three compete for the same binding pocket. Chemical probing of ribosomal RNA further showed that RiboBright protects the same nucleotide that cycloheximide shields, consistent with an identical docking site. High-resolution microscopy revealed that RiboBright signal overlaps strongly with markers for the large ribosomal subunit and with structures such as the endoplasmic reticulum and mitochondria, where many ribosomes reside. At the same time, some bright spots float free in the cytoplasm, likely representing mobile ribosomes not attached to membranes.
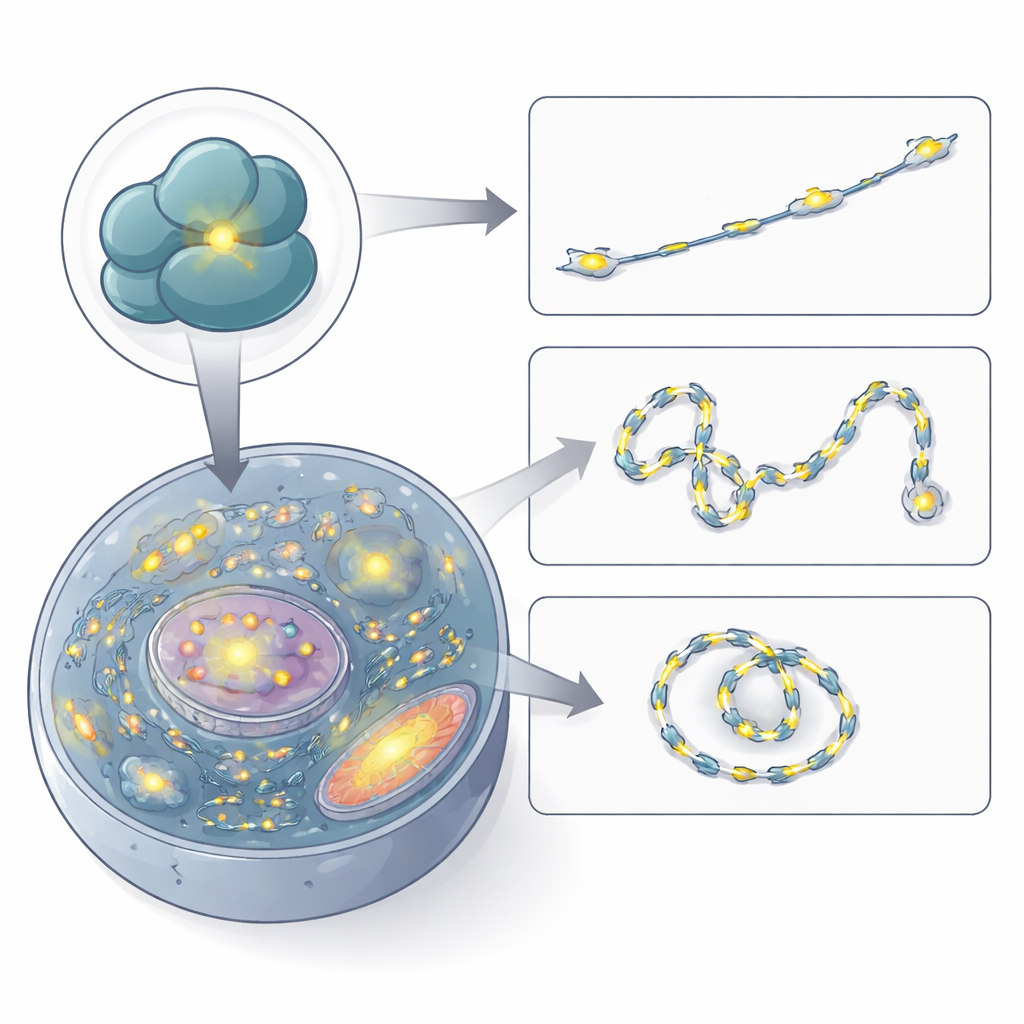
Watching Ribosomes Move in Living Cells
With confidence in its specificity, the authors used RiboBright to film ribosomes in action. By recording images every few hundred milliseconds, they traced the paths of individual fluorescent specks across the cell. Many spots barely moved or stayed confined to small regions, suggesting ribosomes translating in place on anchored messenger RNAs or on the endoplasmic reticulum. Others wandered in a random, diffusive manner, and a smaller fraction showed clear directed motion, hinting at active transport along cellular tracks. The speeds they measured match earlier estimates for messenger RNAs and ribosome clusters, confirming that RiboBright can capture realistic ribosome dynamics in real time.
Different Cell Types, Different Ribosome Strategies
RiboBright also revealed how strongly ribosome content and organization depend on cell identity. Across ten human and mouse cell lines, including stem cells, cancer cells, and noncancerous cells, the pattern and brightness of ribosome spots varied dramatically. Some cells contained large, bright clusters that may correspond to regions of intense protein production, while others displayed mostly small, dispersed specks. Surprisingly, the sheer number of ribosomes in a cell did not reliably predict how much protein it was making: in most cell types, protein output and ribosome content showed little correlation. Embryonic stem cells stood out by having low per-ribosome productivity and relatively uniform translation between individual cells, suggesting that they maintain large reserves of underused ribosomes while keeping protein production noise low.
Ribosomes During Early Cell Fate Decisions
The team then followed mouse embryonic stem cells as they began to differentiate into two early lineages: an ectoderm-like branch and an extraembryonic endoderm (XEN)-like branch. Using RiboBright together with cell-surface markers, they found that both emerging lineages acquired slightly more total ribosome content than undifferentiated cells, but arranged these ribosomes differently. XEN-like cells, which develop long protrusions, showed slower and more confined ribosome movement, consistent with localized protein synthesis in specialized regions. In contrast, ectoderm-like cells displayed more mobile ribosomes and, after about two days, developed striking “translational hubs” where both ribosome signal and new protein synthesis peaked. Over the first 72 hours of differentiation, overall protein production and ribosome levels modestly declined, but the remaining ribosomes became slightly more efficient, especially in differentiating cells.
What This Means for Understanding Cells
Together, these findings paint a richer picture of ribosomes as dynamic, context-dependent machines rather than uniform workhorses. RiboBright gives researchers a practical tool to visualize and count ribosomes across many cell types, in both fixed and living samples, and to relate ribosome position and motion to how much protein a cell is actually making. For non-specialists, the key takeaway is that cells tune not only how many protein factories they build, but also where they place them and how actively they run them, in ways that depend on cell type and developmental state. This new fluorescent probe makes those hidden logistics visible, opening the door to studying how ribosome behavior contributes to development, disease, and responses to therapy.
Citation: Poulladofonou, G., Grandi, C., Hu, X. et al. RiboBright reveals cell-type-specific differences in ribosome organization and movement. Nat Commun 17, 2734 (2026). https://doi.org/10.1038/s41467-026-68947-7
Keywords: ribosomes, fluorescent probes, single-cell imaging, stem cell differentiation, protein synthesis