Clear Sky Science · en
Architectural and evolutionary features of TE-derived TSSs shape tissue-specific promoter activity in the human genome
Hidden switches in our DNA
Nearly half of the human genome is made of mobile DNA fragments, long dismissed as “junk” or genetic freeloaders. This study shows that thousands of these pieces actually act as hidden on–off switches, helping turn genes on in specific tissues such as the brain, lungs, or testis. Understanding how these ancient genetic hitchhikers are reused as control knobs for our genes sheds light on human evolution, disease, and what makes different cell types unique.
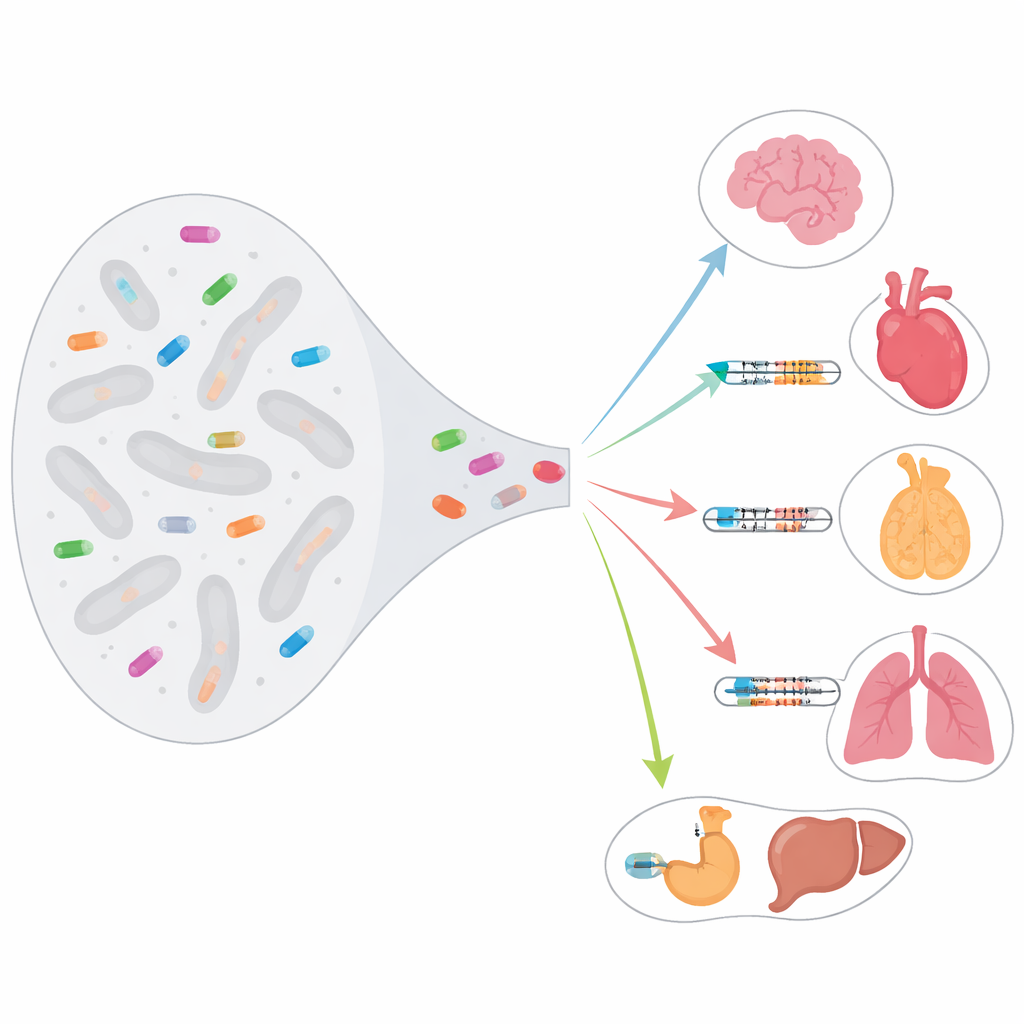
Jumping genes turned into control knobs
Transposable elements are stretches of DNA that once copied and pasted themselves around the genome. Over time, most were silenced by the cell to prevent damage. Yet some insertions have been repurposed as starting gates for gene activity, known as transcription start sites. The authors used a precise technique called RAMPAGE, applied to 115 human samples from 87 tissues and 28 cell types, to map more than 26,000 such start sites that lie within transposable elements. These sites are not random background noise: many are wired into normal gene circuits and help determine when and where genes are switched on.
Tissue-specific switches across the body
By comparing samples across the body, the team found that transposable-element–based start sites are highly tissue specific. More than half appear in only one sample, and genes that use them tend to be turned on strongly but only in certain tissues. In brain, these switches are linked to genes involved in synapses; in lung, to immune defense; in testis, to genes that shape the cell’s contractile machinery; and in spleen, to immune and transport functions. In many genes, these sites are not minor extras: in roughly one quarter of affected genes, they provide at least half of the gene’s promoter activity, effectively acting as the main ignition point for transcription.
Special DNA designs for precise control
The study also reveals that these start sites share a distinctive architectural style. Compared with standard human promoters, they more often form narrow peaks of initiation, meaning transcription begins at a very precise base rather than being spread across a broad region. They are enriched for a classic “TATA box” pattern just upstream of the start, and they tend to sit in DNA regions that lack CpG islands, a feature typically associated with switch-like, tissue-specific control. Biochemical assays confirm that many of these sites can drive strong gene activity on their own, particularly when they serve as the unique or dominant promoter for a gene.
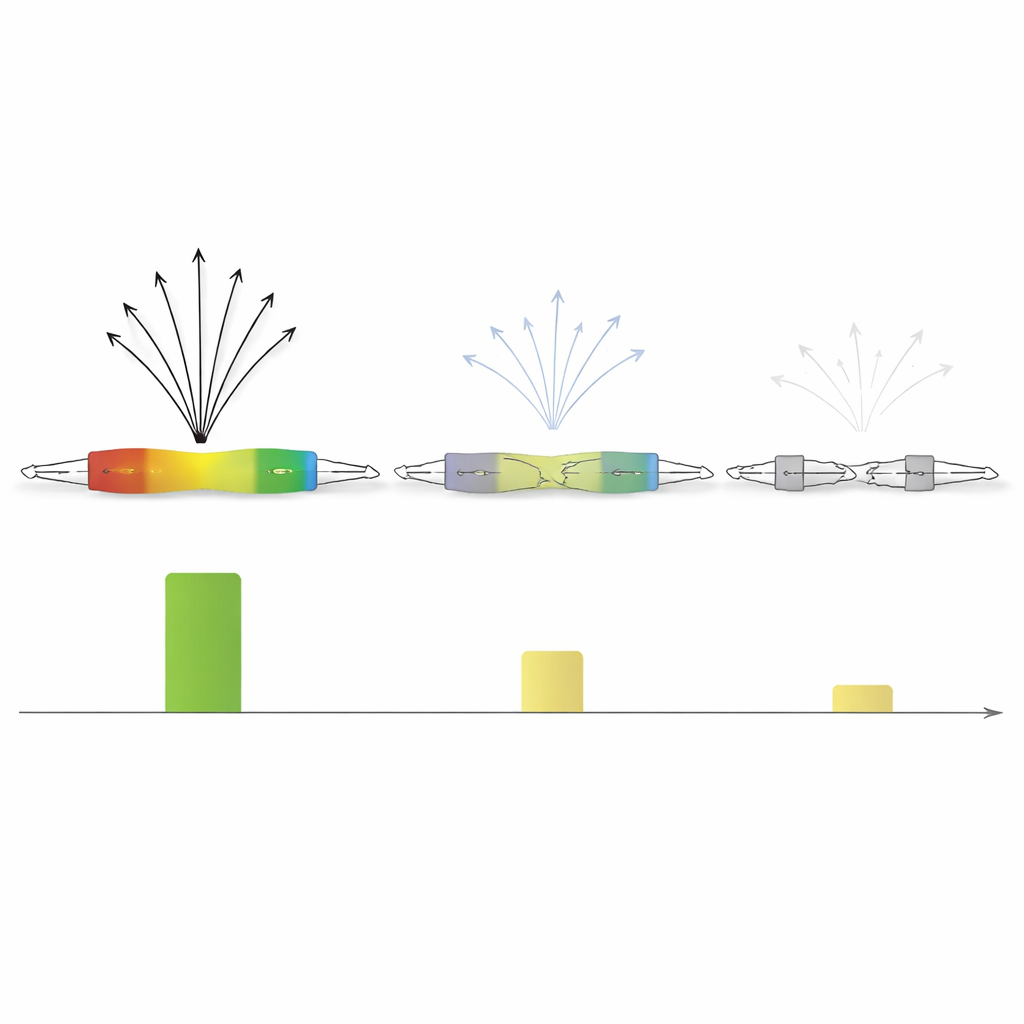
Younger elements, sharper switches
Because transposable elements entered the genome at different times, the authors could treat them as a kind of evolutionary time series. They found that younger families of elements, especially those unique to primates and great apes, often retain intact promoter-like motifs. These younger copies show strong intrinsic activity and highly focused starting points for transcription. Older families, by contrast, accumulate mutations and structural breakage. Their transcription start sites become weaker and more scattered, losing the sharp focus seen in their younger counterparts and relying more heavily on surrounding genomic context.
How mobile DNA helped shape human traits
By tying tissue-specific gene control to the age and structure of mobile DNA fragments, this work suggests that bursts of transposable element activity left behind a toolkit of potential switches that evolution could recruit. Young, still-intact elements provide powerful, precise control points, some of which have been wired into pathways for brain function, immunity, metabolism, and reproduction, particularly in primates and hominids. In simple terms, bits of once-roaming DNA have been recycled into finely tuned switches that help define what our cells do, and may have contributed to traits that distinguish humans and our close relatives.
Citation: Zhang, Y., Fan, Y., Wu, H. et al. Architectural and evolutionary features of TE-derived TSSs shape tissue-specific promoter activity in the human genome. Nat Commun 17, 2219 (2026). https://doi.org/10.1038/s41467-026-68934-y
Keywords: transposable elements, gene regulation, promoters, tissue specificity, human evolution