Clear Sky Science · en
An experimentally-informed polymer model reveals high resolution organization of genomic loci
How DNA Folding Shapes Cell Identity
Every cell in your body carries essentially the same DNA, yet brain cells, skin cells, and stem cells behave very differently. A key reason is how that DNA is folded and packed inside the nucleus. This study introduces a new way to "see" that folding in remarkable detail, connecting the physical arrangement of DNA to whether important genes are switched on or off. By blending experiments with physics-based computer simulations, the authors reveal hidden clumps of genetic material that appear to act as basic building blocks of genome organization.
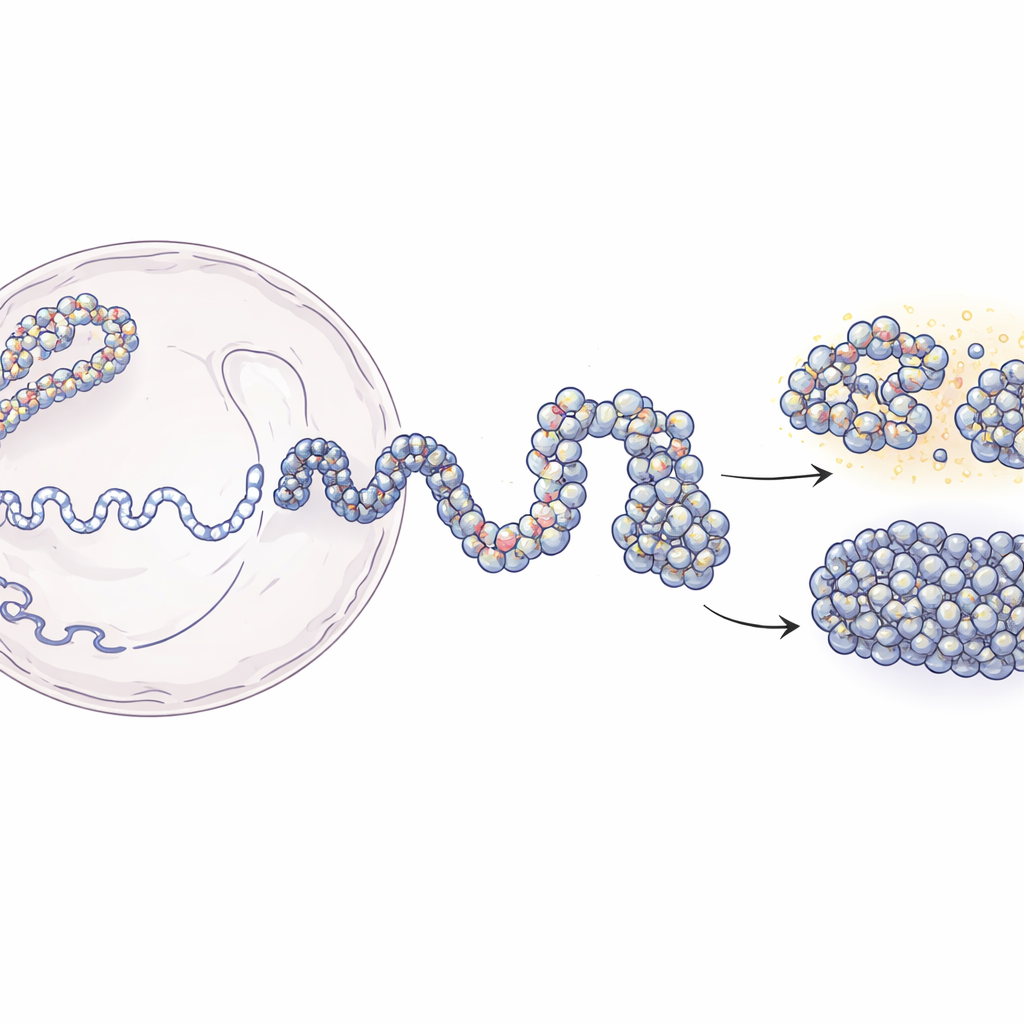
From Long DNA Threads to 3D Genome Maps
Inside the nucleus, DNA is wrapped around protein spools called nucleosomes, forming a beads-on-a-string structure known as chromatin. Modern techniques like Hi-C and Micro-C can tell us which pieces of DNA are near each other in 3D, but they usually offer blurry, population-averaged snapshots. On the other hand, experiments that locate individual nucleosomes provide sharp local detail but little sense of the larger structure. This work bridges that gap. The authors start from low-resolution contact maps that report how often distant DNA segments touch, then combine them with experimental maps of nucleosome positions. Using principles from polymer physics, they build simulated 3D ensembles of chromatin that match experimental data yet resolve structures down to a few dozen DNA letters.
A Two-Step Strategy to Rebuild Chromatin
The modeling approach unfolds in two main stages. First, the team uses Hi-C data to generate many possible large-scale shapes of a 200,000-letter stretch of DNA, treating the chromatin as a flexible chain where 5,000-letter segments are gently guided to make or avoid contacts as observed in the experiments. These coarse structures capture the overall folding pattern that proteins in the cell help to create. In the second stage, each large bead is replaced by a much finer chain made of individual nucleosomes and the short DNA linkers between them. The positions of these nucleosomes come from an enzyme-based mapping method (MNase-seq) that reveals where they typically sit along the genome. The fine-grained chains are then allowed to fold while still respecting the broader architecture. When the researchers "blur" their high-resolution models back to the experimental resolutions, they reproduce both Hi-C and Micro-C contact maps with high accuracy.
Discovering Nucleosome Blobs as Structural Units
When the authors zoomed in on their simulated structures, a striking pattern emerged: nucleosomes were not arranged evenly, but instead gathered into irregular clusters, which the authors call nucleosome blobs. These blobs resemble the clumpy structures seen previously in super-resolution microscope images of real cells. By analyzing many thousands of simulated snapshots, the team showed that these blobs are elongated, not spherical, and typically contain several nucleosomes packed together. Crucially, contacts inside these blobs correspond closely to the domain-like interaction blocks seen in experimental data, indicating that blobs are not random accidents but fundamental 3D units of chromatin organization. The simulations even predict additional subtle domain boundaries that are hard to detect experimentally, suggesting that this physical model can uncover fine-scale features hidden in noisy data.
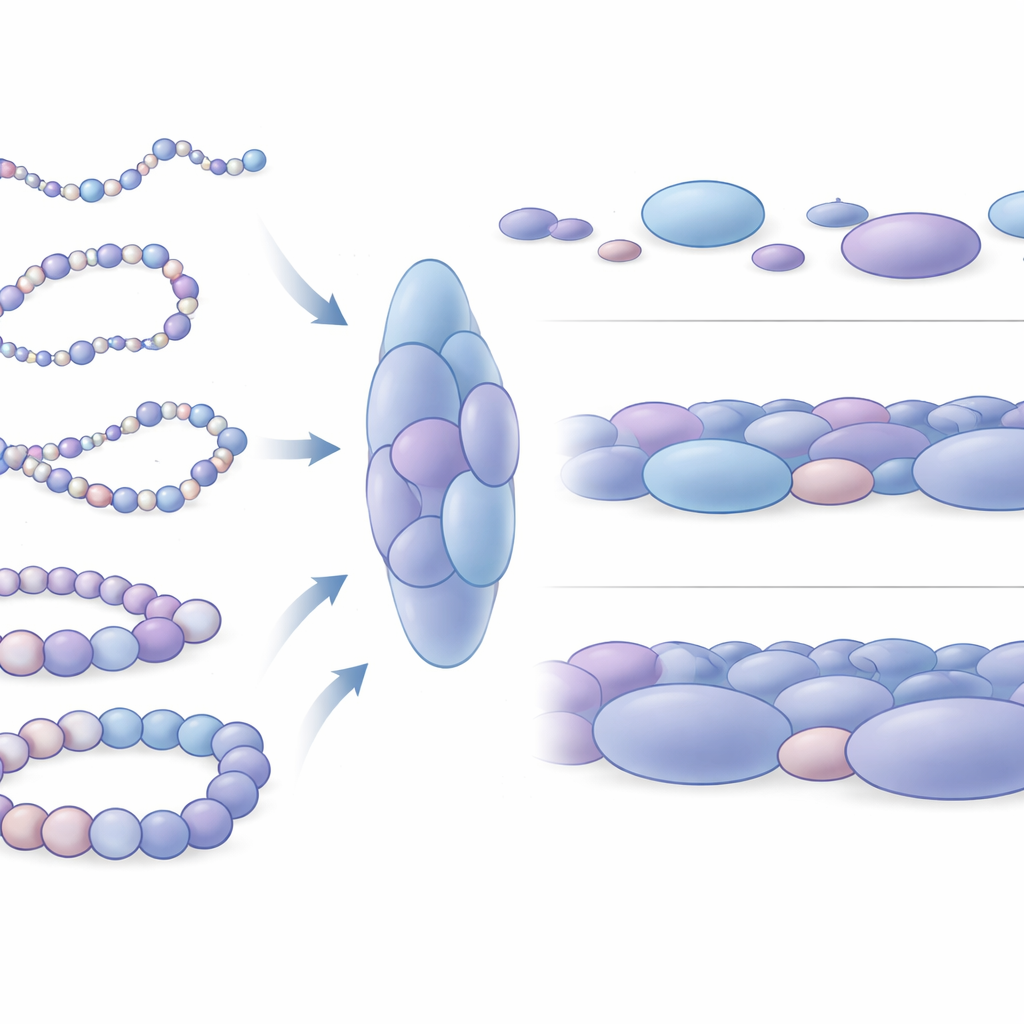
How Packing Differences Reflect Gene Activity
The researchers then asked how these blobs differ around genes that are active versus those that are silent. They focused on four well-studied stretches of human DNA, including two genes that help keep stem cells in a flexible, undifferentiated state (Nanog and Lin28A) and two developmental control genes (HoxB4 and HoxA13) that are shut down in the same cells. Around the inactive genes, blobs were on average larger and more tightly packed, with nucleosomes forming more fully closed local arrangements. In contrast, blobs near active genes were smaller, somewhat looser, and more varied. On a broader scale, the DNA around active genes sampled many more distinct shapes and was mechanically more bendable, while the regions around silent genes behaved like stiffer segments of chromatin. This mechanical difference likely affects how easily distant regulatory DNA elements can meet and cooperate with gene switches.
Why This Matters for Understanding Gene Control
Taken together, the findings paint a picture in which the genome is built from dynamic clusters of nucleosomes whose size, shape, and spacing help determine whether nearby genes are accessible or locked down. The new model connects experimental contact data, nucleosome maps, and physical principles into a single framework that explains how stem-cell genes can remain flexible and interactive while developmental genes stay sequestered in stiffer, more compact neighborhoods. For non-specialists, the key idea is that gene activity is not governed by DNA sequence alone; it also depends on how that DNA is folded into three-dimensional structures. By revealing nucleosome blobs as basic building blocks of that folding, this work offers a powerful route to link microscopic genome architecture to large-scale processes like development, cell identity, and disease.
Citation: Mittal, R., Heermann, D.W. & Bhattacherjee, A. An experimentally-informed polymer model reveals high resolution organization of genomic loci. Nat Commun 17, 2338 (2026). https://doi.org/10.1038/s41467-026-68928-w
Keywords: chromatin folding, nucleosome clusters, 3D genome organization, gene regulation, polymer modeling