Clear Sky Science · en
Evolutionary repurposing of a metabolic thiolase complex enables antibiotic biosynthesis
How everyday microbes reinvent their chemistry
Many of the antibiotics and medicines we rely on are quietly crafted by bacteria. These tiny chemists use enzymes—molecular machines—to build complex defensive compounds. This study uncovers how a core metabolic enzyme complex, once used for basic cell housekeeping, has been evolutionarily rewired in oral bacteria to manufacture a potent antibiotic. Understanding this transformation not only reveals how nature invents new chemistry but also offers clues for designing next-generation drugs and biocatalysts.
From basic metabolism to chemical weaponry
All living cells depend on a set of conserved enzymes that run their central metabolism, such as breaking down fats and building essential molecules. One such group, called the thiolase superfamily, normally helps handle small energy-rich molecules like acetyl-CoA and supports vital pathways for making lipids and other cell components. The authors discovered that in several oral Streptococcus species, a three-part thiolase-based complex—made of proteins related to HMGS, ACAT, and a third helper protein—has abandoned its ancestral metabolic role. Instead, this complex now completes the biosynthesis of an antibiotic called reutericyclin A, which helps these bacteria compete in the crowded environment of the mouth.
A new kind of enzyme reaction on an antibiotic scaffold
By reconstituting the Streptococcus complex in the lab and feeding it a chemically synthesized precursor of reutericyclin A, the researchers showed that the complex performs an unusual carbon–carbon bond-forming reaction known as Friedel–Crafts C-acylation. Rather than attaching an acetyl group to oxygen, as many enzymes do, it installs this group directly onto a carbon atom of a pyrrolidine-2,4-dione ring—an important part of the reutericyclin scaffold. The complex accepts both natural acetyl donors such as acetyl-CoA and several artificial donors, then can even run the reaction in reverse, breaking the bond it created. This flexibility expands the known repertoire of biological acyltransferases and hints at how such enzymes might be adapted to build diverse chemical products.
Repurposing a conserved protein scaffold
Using high-resolution cryo-electron microscopy, the team solved the three-dimensional structure of the Streptococcus complex, termed SmaATase. Despite its new function, SmaATase retains a striking architectural resemblance to a related complex in archaea that participates in the mevalonate pathway for building isoprenoid lipids. Both are large multimeric assemblies built from repeating modules and feature a shared cavity where the cofactor CoA can bind and move between active sites. However, key catalytic residues that are essential for the original metabolic reaction are missing or altered in the bacterial version. Biochemical tests confirm that SmaATase can no longer make the ancestral product HMG-CoA and instead uses acetyl-CoA purely as a donor for its new small-molecule substrates.
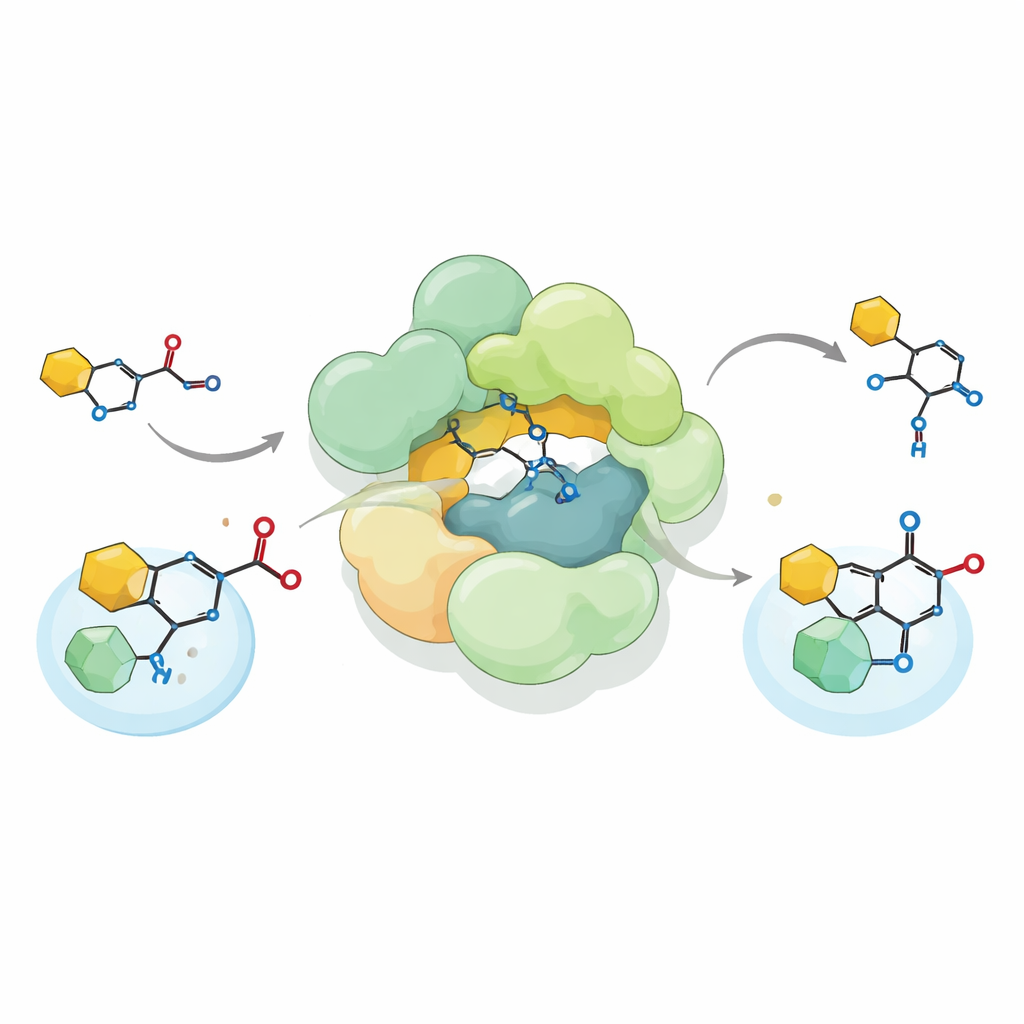
An evolutionary trail from metabolism to antibiotics
To understand how such a repurposing might have unfolded, the authors compared sequences and activities of related complexes across many bacteria and archaea. They identified over a thousand gene clusters containing the same trio of enzyme types. Some still support classical thiolase chemistry; others, like those in Pseudomonas, fuel different small-molecule antibiotics. A representative complex from Pseudomonas fluorescens appears to sit partway along this evolutionary path: its HMGS-like domain is disabled, but its ACAT-like unit still performs a more conventional reaction on CoA-linked substrates. In SmaATase and its Pseudomonas counterpart that makes the crop-protective compound DAPG, further changes around the active site have tailored the enzymes to recognize nonstandard acceptors such as aromatic rings or tetramic acid cores.
Why a tiny acetyl group makes a big difference
Ultimately, the biological payoff of this molecular makeover is clear. When the team tested the acetylated antibiotics reutericyclin A and DAPG against a panel of Gram-positive bacteria, these molecules showed strong killing activity, whereas their unacetylated precursors were far weaker or almost inactive. That small acetyl group, placed in just the right position by the repurposed enzyme complex, turns a modest scaffold into a powerful weapon, enabling Streptococcus in the mouth or Pseudomonas around plant roots to outcompete their neighbors. This work showcases how evolution can recycle existing enzyme frameworks with minimal but strategic mutations, turning routine metabolic machinery into specialized tools for chemical defense—and offering a blueprint for engineers hoping to design new carbon–carbon bond-forming catalysts for drug discovery and synthetic biology.
Citation: Liao, G., Sun, R., Shen, Z. et al. Evolutionary repurposing of a metabolic thiolase complex enables antibiotic biosynthesis. Nat Commun 17, 2196 (2026). https://doi.org/10.1038/s41467-026-68910-6
Keywords: enzyme evolution, natural product biosynthesis, antibiotic chemistry, metabolic repurposing, Friedel-Crafts acylation