Clear Sky Science · en
Engineering plasmids with synthetic origins of replication
Why Rewriting Microbial DNA Matters
Many of the tools behind modern biotechnology, from insulin production to cutting-edge gene circuits, rely on small circles of DNA called plasmids. These genetic workhorses shuttle engineered genes into bacteria, but their basic design has barely changed in decades. This study shows that the very heart of plasmids—the sequences that tell cells when and how often to copy them—can be completely reimagined. By rebuilding this "copy engine" from the ground up, the authors create plasmids that are easier to tune, combine, and program, opening the door to more flexible diagnostics, biomanufacturing, and synthetic biology.
Old Tools with Hidden Limits
Plasmids naturally evolved as mobile DNA, helping bacteria share traits such as antibiotic resistance or new ways to digest food sources. In the lab, engineers borrow these plasmids to carry useful genes, but most rely on a few classic designs discovered in the 1980s. Those old designs hide a tangle of overlapping genetic parts that control how many copies of a plasmid sit inside each cell and whether different plasmids can peacefully coexist. Because these parts are fused together, changing one feature can unpredictably break another. As a result, scientists are stuck with a short menu of plasmids that have fixed copy numbers and limited compatibility, restricting how complex engineered systems can become.
Rebuilding the Copy Engine from Scratch
The researchers focused on a widely used origin of replication from the pMB1 plasmid family. In its natural form, this origin uses a finely balanced conversation between two RNAs—one that starts DNA copying and another that shuts it down—to keep plasmid numbers in check. The team first "refactored" this system: they pulled apart overlapping genes, disabled a buried promoter, and placed key pieces on separate, clean genetic cassettes. This alone showed that the origin’s core job could be preserved while simplifying its layout, turning a messy, evolution-shaped part into something more like a modular machine with accessible components.
Swapping in Synthetic Control Knobs
With the original control logic exposed, the authors then replaced it with fully synthetic regulators. They connected the replication primer to engineered RNA switches that act like dimmer knobs: small controlling RNAs can flip these switches to either allow or block production of the primer and, in turn, plasmid copying. By choosing different switch variants and pairing them with promoters of varying strength, they tuned plasmid copy numbers across more than two orders of magnitude. They also explored different physical layouts of the control cassettes on the DNA, discovering arrangements that improved stability and allowed them to shrink the replication region to a compact, functional core while adding natural helper systems that prevent problematic plasmid tangles.
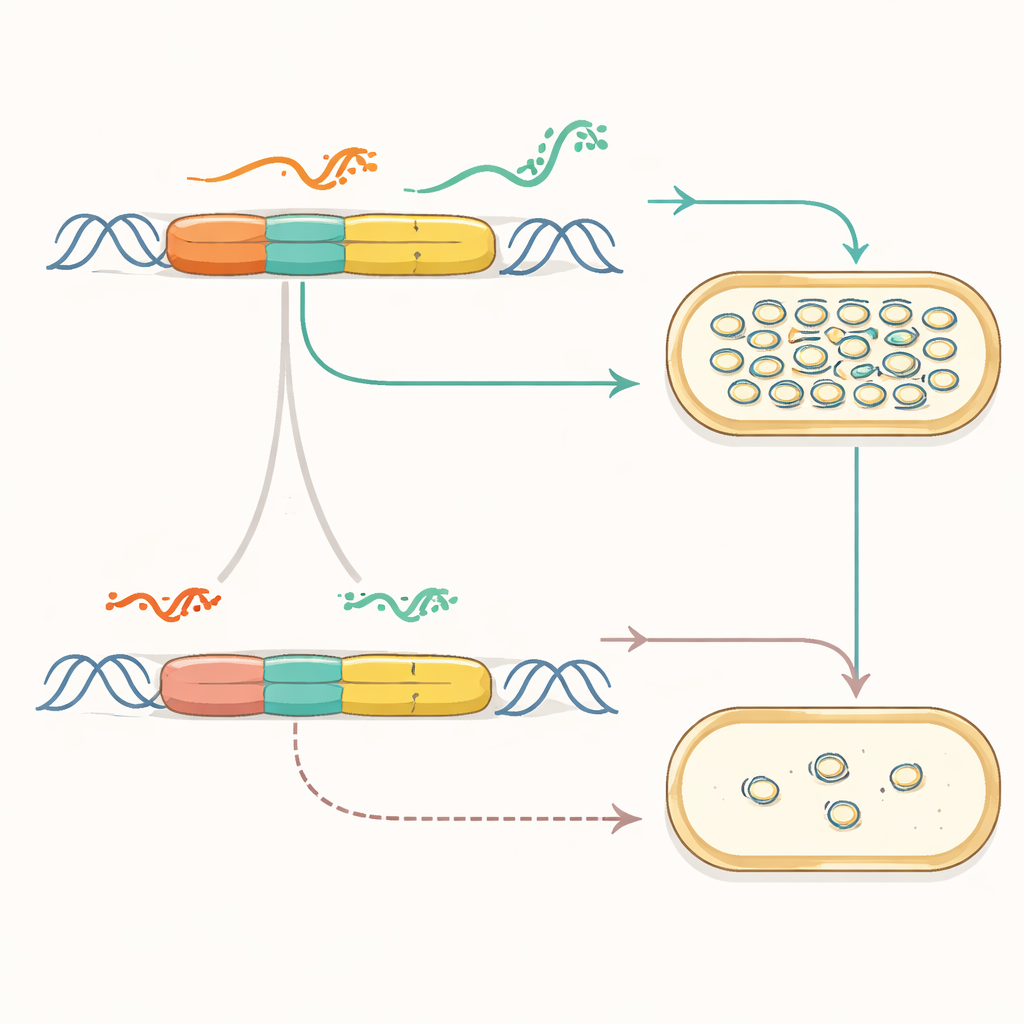
Turning Chemical Signals into DNA Counts
Once the copy engine was modular, the team showed it could be wired to sense the outside world. They connected the synthetic origin to inducible promoters and RNA structures called riboswitches, which change shape in response to small molecules. In these new plasmids, adding a chemical like IPTG or cumate caused the plasmid copy number—and thus a reporter signal—to rise or fall. Multiple signals could be combined so that one chemical drove copying while another reined it in. The researchers even built pairs of plasmids, each responding to a different chemical, and tracked their changing copy numbers in the same cell using DNA sequencing, effectively turning plasmid counts into a barcode of environmental history.
Many Custom Plasmids in One Cell
A key test of this approach was whether several independently controlled plasmids could coexist. By drawing on a library of orthogonal RNA regulators, the team built six different plasmids, each with its own synthetic origin and antibiotic marker, and introduced all six at once into E. coli. Whole-plasmid sequencing over several days confirmed that all six remained present, even though their relative abundances shifted. Attempts to do the same with six conventional plasmids failed, underscoring how the refactored design and added stability systems make the new plasmids more compatible and robust when crowded together in a single cell.
What This Means for Future Biotechnology
To a non-specialist, the takeaway is straightforward: the authors have turned plasmids from rigid, one-size-fits-all tools into a customizable platform. Their synthetic origins of replication act like plug-and-play engines whose speed, sensitivity, and inputs can be chosen on demand. This makes it possible to build bacteria that record chemical exposures as changes in DNA copy number, test many genetic pathways in parallel by distributing them across multiple plasmids, or finely balance growth and production in industrial strains. While some stability trade-offs remain, this work shows that the core mechanics of plasmid replication are no longer off-limits to engineering, opening new space for innovation in synthetic biology.
Citation: Liu, B., Seet, Z.R.D., Peng, X. et al. Engineering plasmids with synthetic origins of replication. Nat Commun 17, 2255 (2026). https://doi.org/10.1038/s41467-026-68907-1
Keywords: synthetic plasmids, origin of replication, RNA regulators, copy number control, synthetic biology