Clear Sky Science · en
Origin of small chromosome A08 and genome evolution of Arachis species
Why Peanut DNA Matters
Peanuts are a staple snack and cooking oil around the world, but behind every kernel lies a surprisingly complex genetic story. Wild peanut relatives in South America carry natural resistance to pests and diseases that could make crops hardier and more sustainable. To fully tap that potential, scientists need to understand how peanut genomes are built and how they have changed over millions of years. This study uncovers the origin of a peculiar tiny chromosome in peanuts and maps out how different wild species are related, offering a genetic roadmap for future breeding.
Following the Family Tree of Peanuts
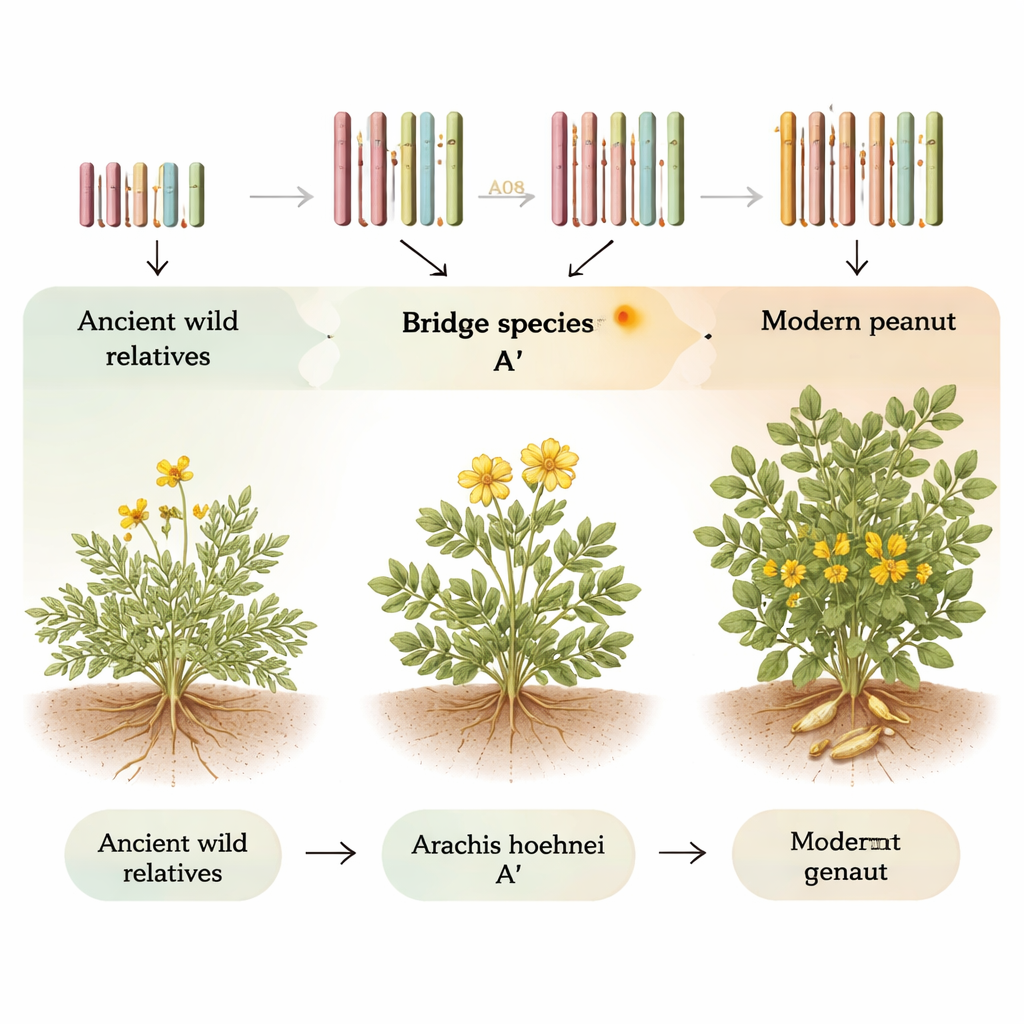
The peanut we eat today is actually a recent newcomer in evolutionary terms. It formed when two wild species with slightly different sets of chromosomes merged and doubled their DNA, creating a plant with four copies of each chromosome instead of two. Earlier work had shown that species called Arachis duranensis and Arachis ipaensis donated these two genome halves, known as the A and B genomes. But the wider family, which includes more than 80 wild species, still had an unclear family tree, especially for less-studied genome types labeled F, K, and H. One puzzling feature was a uniquely small chromosome known as A08 that appears only in A-type genomes and stands out like a runt among taller siblings.
Painting Chromosomes to Reveal Hidden Patterns
To sort out who is related to whom, the researchers used a method likened to painting chromosomes. They designed thousands of short DNA tags that latch onto specific regions of each chromosome and glow under a microscope in different colors. By applying these “paints” to 17 peanut and wild Arachis species, they could match microscopic chromosomes to their digital counterparts in genome sequences and group them into 10 consistent sets across species. This karyotype map revealed where big chunks of DNA had been flipped, swapped, or duplicated as species diverged over time. It also showed that one wild species, Arachis hoehnei, has chromosomes that do not quite fit the classic A or B types and carries a larger version of the small chromosome’s ancestor.
A Bridge Genome and the Birth of a Tiny Chromosome
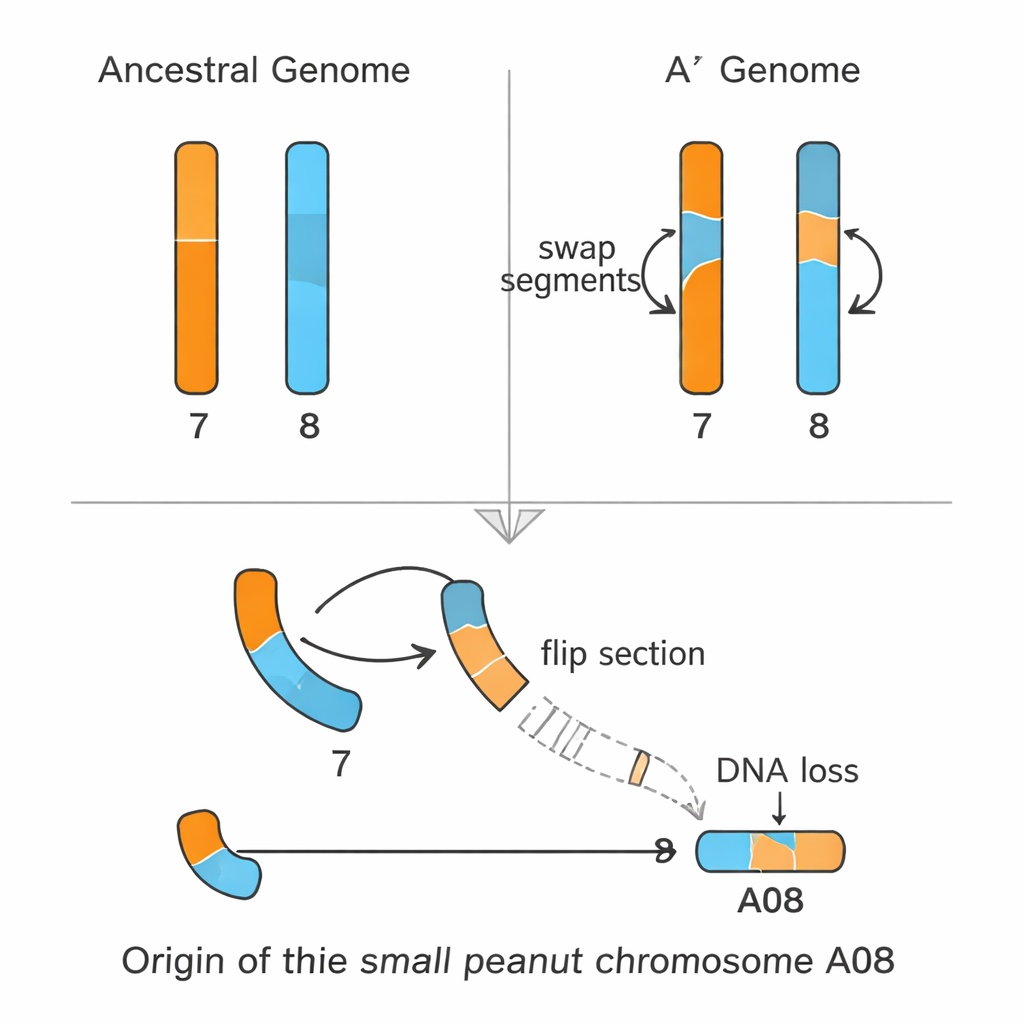
The team then built a complete, gap-free genome sequence for A. hoehnei from end to end on all 10 chromosomes—an achievement called a telomere‑to‑telomere assembly. Comparing this genome to cultivated peanut and other relatives showed that A. hoehnei forms a genetic “bridge” between the A and B genomes. Its genome was therefore designated A′ (A‑prime): closely related to the A genome but distinct. By aligning the A′ chromosomes with those of modern A and B genomes, the researchers reconstructed how the odd small chromosome A08 arose. First, the ancestors of chromosomes 7 and 8 exchanged segments to form new versions in the A′ genome. Later, in the A genome lineage, two large stretches of the future A08 were flipped in orientation (inversions) and more than 50 million DNA letters—rich in repeated sequences and about 500 genes—were lost. What remained is the much shorter A08 found in today’s A‑genome peanuts.
Junk DNA, Repair Systems, and Disease Resistance
The A′ genome turned out to be the largest among the wild peanut genomes studied, packed with repetitive DNA elements that copy and move themselves around. These sequences, once dismissed as “junk,” have clearly helped reshape chromosomes and expand genome size. Many of the structural changes that distinguish A, B, and A′ genomes trace back to such mobile elements. Gene-family analyses showed that A. hoehnei carries extra copies of genes involved in DNA repair, suggesting it evolved a strong system for keeping this restless genome stable. The species also harbors unique genes and gene versions linked to stress and disease responses. When the team exposed A. hoehnei to web blotch, a serious leaf disease, dozens of genes involved in plant–pathogen interactions and protective compounds switched on, including a defense‑related PR10 protein with an insertion not seen in cultivated peanut.
Building New Peanuts for the Future
To test how compatible these genomes are, the researchers crossed a cultivated peanut variety with A. hoehnei. The initial hybrid had poor fertility, but after doubling its chromosomes they produced a hexaploid line carrying A, B, and A′ genome sets. Though this synthetic peanut was still less vigorous than modern varieties, it showed that genes from the A′ genome can be combined with cultivated peanut, opening a path to transfer disease‑resistance traits into future crops. Putting all the evidence together, the authors propose an evolutionary model in which an ancestral genome split into several lines, giving rise to the F, H, B, K, A′, and eventually modern A genomes. Along this path, large DNA rearrangements and mobile elements acted as powerful engines of change.
What This Means for Farmers and Breeders
For non‑specialists, the key takeaway is that the peanut genome is not a static blueprint but a living record of flips, swaps, and losses of DNA. The strange small chromosome A08 is the end product of these events, and understanding its history reveals how wild species are connected and where valuable traits reside. By pinning chromosomes to precise DNA sequences and decoding the bridge genome A′, this study equips breeders with detailed maps for bringing disease resistance and other useful traits from wild relatives into cultivated peanut. Over time, that knowledge could translate into hardier crops, more reliable yields, and reduced reliance on chemical treatments, all rooted in a deeper grasp of peanut’s evolutionary journey.
Citation: Du, P., Fu, L., Chen, G. et al. Origin of small chromosome A08 and genome evolution of Arachis species. Nat Commun 17, 2029 (2026). https://doi.org/10.1038/s41467-026-68884-5
Keywords: peanut genome evolution, Arachis hoehnei A prime genome, small chromosome A08, structural variation in plants, wild peanut disease resistance