Clear Sky Science · en
Characterising processing conditions that artifactually bias human brain tissue transcriptomes
Why brain handling after death matters
Many of the most important insights into Alzheimer’s disease, Parkinson’s disease and other brain disorders come from studying brain tissue donated after death. But the way this tissue is handled in the hours after a person dies can subtly, and sometimes dramatically, change which genes appear active. This study asks a deceptively simple question with big consequences: when we read the molecular “messages” in autopsy brain tissue, how much of what we see reflects the person’s biology, and how much is a side effect of time and temperature after death?
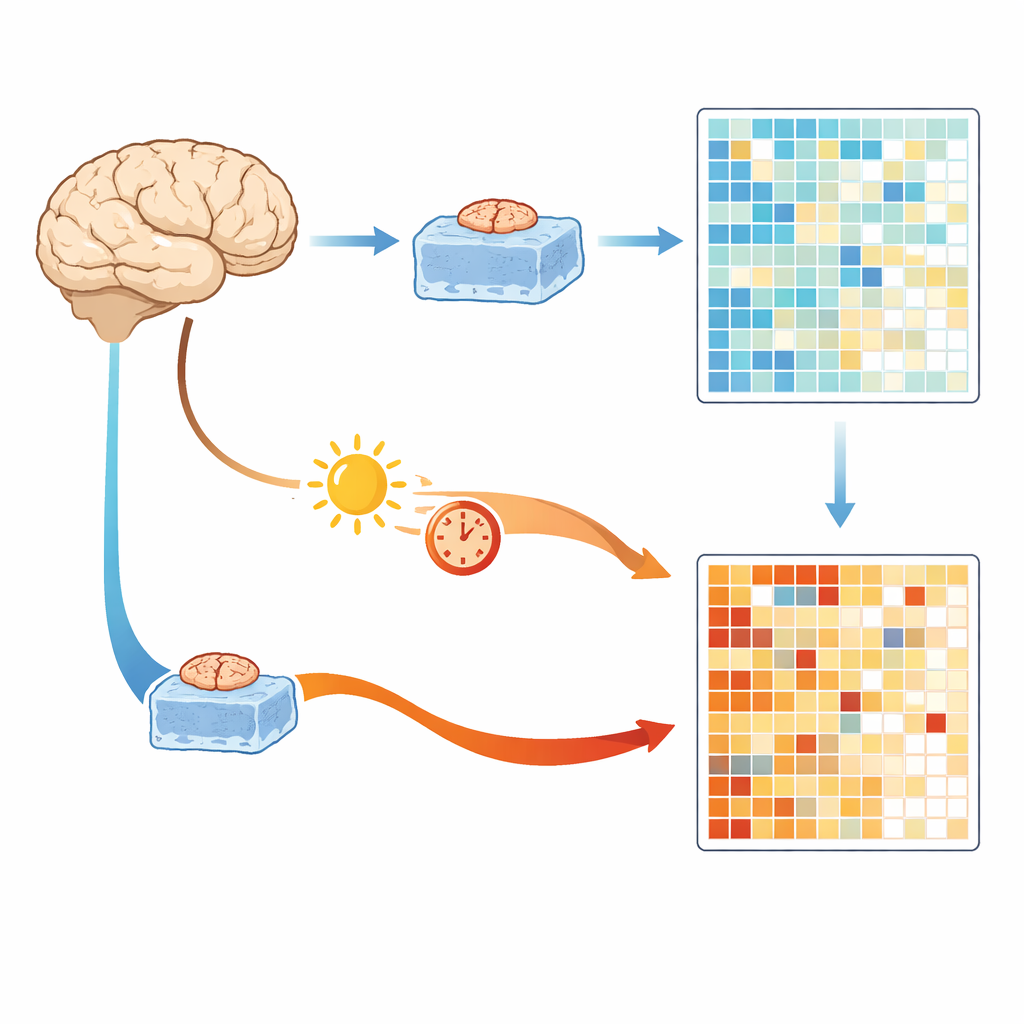
Comparing fast-frozen surgery tissue to autopsy brains
The researchers began with a rare advantage: access to small pieces of healthy-looking brain tissue removed during surgery for tumors, which can be cooled and frozen within about half an hour. These samples provide a close snapshot of gene activity in the living brain. The team compared them with brain tissue from large autopsy banks collected after either a short delay of about six hours or a long delay of about 36 hours. All samples were processed and sequenced in the same way to avoid technical differences. Across thousands of genes, the main factor that separated samples was not the donor’s age or sex, but whether the tissue came from rapidly frozen surgery samples or delayed autopsy samples.
Hidden stress signals and the rise of artifact genes
Both short- and long-delay autopsy tissues showed strong changes in gene activity compared with the immediate surgery tissue. Many of the increased genes were linked to stress responses, energy production in mitochondria and inflammatory pathways. The authors call this shared cluster of genes “Brain Artifact Genes,” or BAGs, because they appear to be turned on by conditions after death rather than by disease itself. Even a relatively short six‑hour delay was enough to produce thousands of changes, including genes involved in communication between nerve cells, suggesting that some apparent “disease signals” in past studies may partly reflect how long the brain sat before being preserved.
Testing the roles of time, temperature and cell type
To tease apart which post-mortem factors mattered most, the team took pieces of the surgery-derived tissue and deliberately held them at either refrigerator temperature or room temperature for different lengths of time before freezing. They then measured gene activity again. Tissue kept briefly in the fridge looked most like the immediately frozen samples, while longer times and warmer temperatures caused stronger and more widespread BAG activation. By analyzing individual cell nuclei, the researchers also found that different brain cell types responded at different stages: glutamatergic neurons were the earliest “responders” after several hours at room temperature, while oligodendrocytes and microglia showed the strongest artifact signatures after about a day. This means that measurements of specific cell populations can be skewed in time-dependent ways.
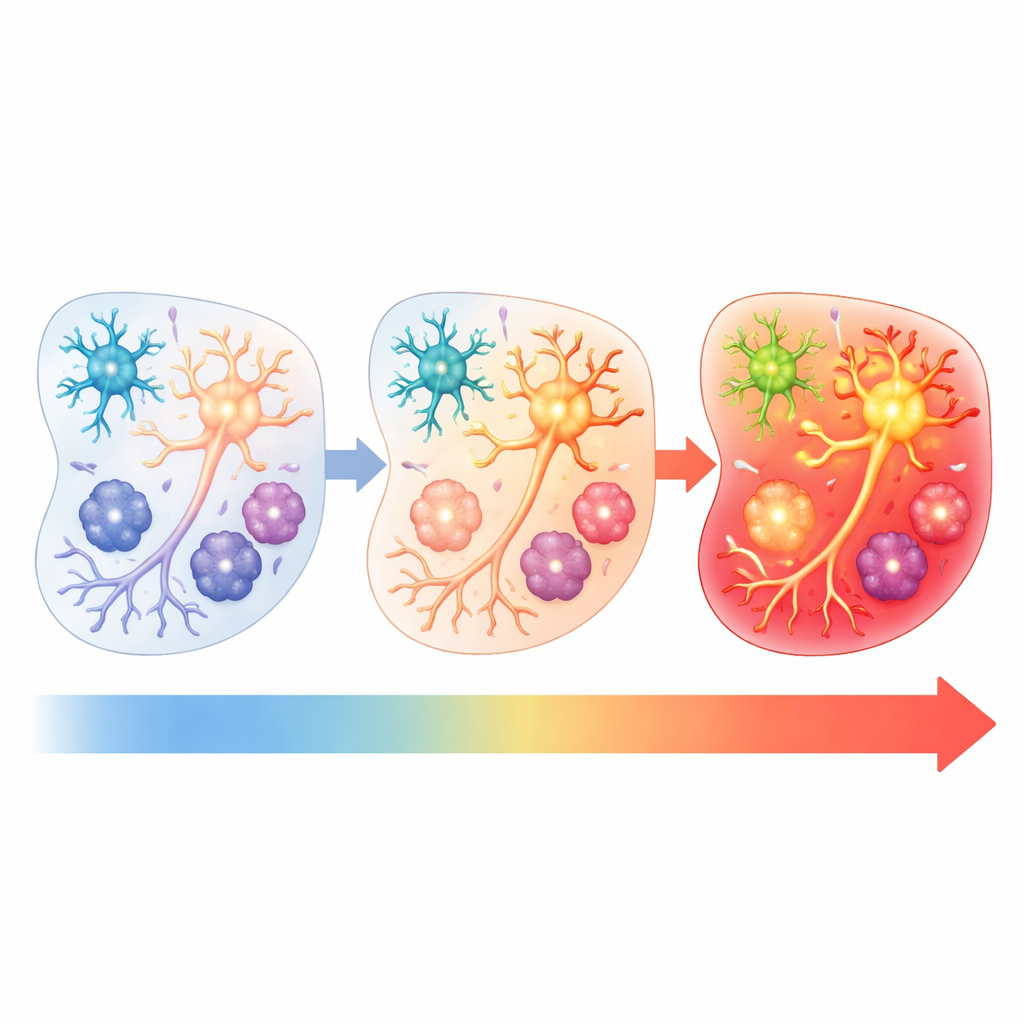
Building a molecular quality score with machine learning
Because no brain bank can perfectly control every detail of post-mortem handling, the authors turned to machine learning to create a practical quality-check tool. Using gene activity patterns from tissue exposed to known combinations of time and temperature, they trained a model to recognize three broad “domains” of processing conditions. From thousands of genes, the model distilled a smaller signature they call TTRUTH (Time and Temperature Response genes Underlying Transcriptional Heterogeneity). The resulting TTRUTH score estimates how strongly any given brain sample carries time‑ and temperature‑related artifacts. When applied to multiple independent autopsy datasets from other studies, most samples fell into a domain consistent with moderate artifact exposure, while a minority showed patterns more like ideal handling or severe stress, highlighting real-world variability across donors and centers.
What this means for brain research
For non-specialists, the take‑home message is that brain tissue “remembers” how it was treated after death, and these memories can masquerade as signs of disease. This work provides a roadmap and an openly available online tool that allows researchers to score their own datasets for hidden handling effects, distinguish biological signals from technical noise and better group samples for analysis. Ultimately, by recognizing and correcting for these artifacts, scientists can draw more reliable conclusions about how the human brain works in health and illness—and make more confident progress toward new treatments.
Citation: Yaqubi, M., Thomas, M., Talbot-Martin, J. et al. Characterising processing conditions that artifactually bias human brain tissue transcriptomes. Nat Commun 17, 2848 (2026). https://doi.org/10.1038/s41467-026-68872-9
Keywords: brain biobanks, post-mortem tissue, gene expression, RNA sequencing, machine learning