Clear Sky Science · en
Coordinated gene family evolution shapes the genome of dimorphic Mucorales
One Fungus, Two Bodies
Some fungi can live a double life, switching between a single-celled “yeast” form and a branching “mold” form. This shape-shifting ability helps them survive changing environments and, in some cases, invade human tissues. The study summarized here uncovers how one group of these fungi, the Mucorales, reorganizes and uses its genes to support both lifestyles within a single genome.
Why Shape-Shifting Fungi Matter
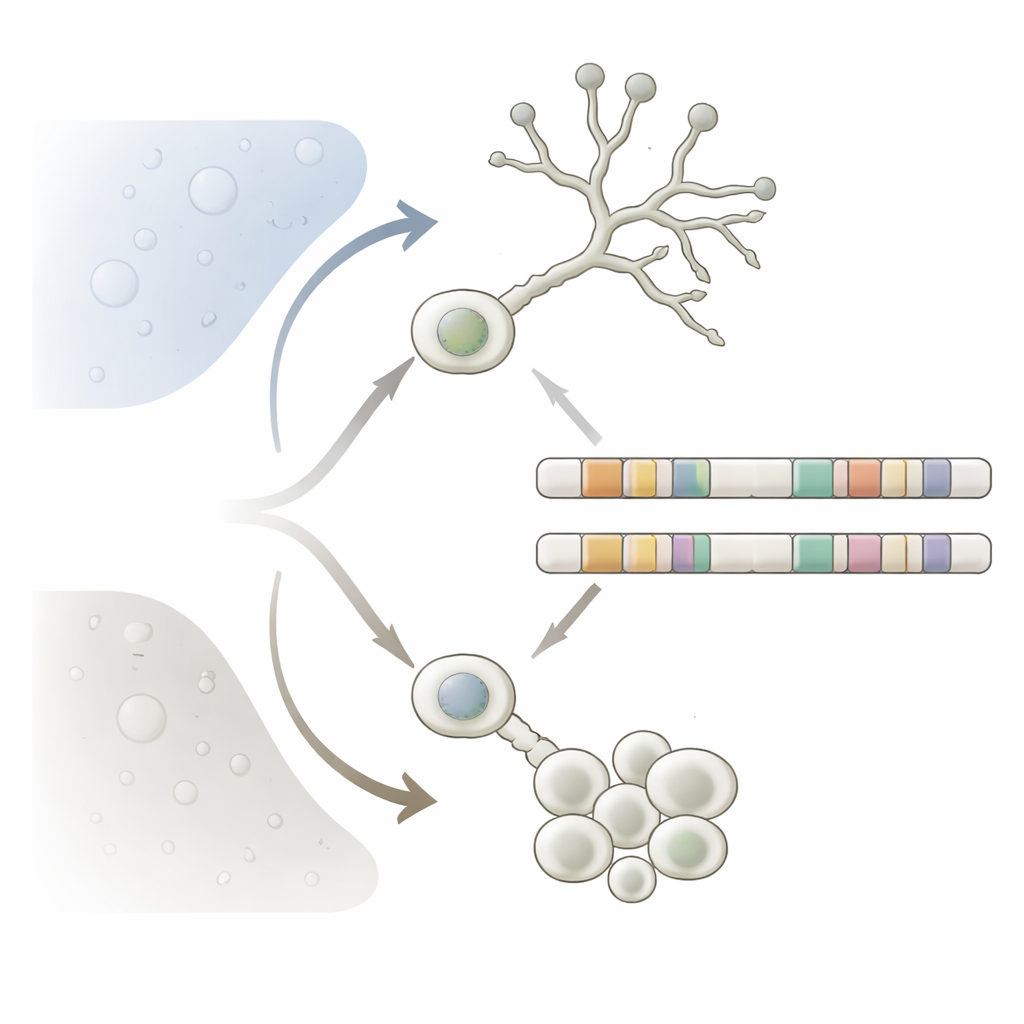
Dimorphic fungi are important not just for ecology but also for human health. In Mucorales, the yeast form is made of isolated round cells that favor low-oxygen, sugary conditions and grow by budding. The mycelial form forms long filaments that thrive in oxygen-rich environments and can invade tissues and surfaces. In several Mucorales species, only the filamentous form is strongly infectious, causing mucormycosis, a severe disease in people with weakened immune systems. Understanding how these fungi switch form can reveal why they are so adaptable, why they resist some drugs, and which genetic features distinguish dangerous dimorphic species from their harmless relatives.
A Genome Built for Switching
The authors focused on the model fungus Mucor lusitanicus and followed it through four stages: yeast, early mycelium, the reverse switch back toward yeast, and mature mycelium. By sequencing RNA, they measured which genes were active in each state. They found that about 70% of all genes change their activity during the shape shift, far more than in many other fungi. Yeast cells tend to turn on genes for basic metabolism and making cellular building blocks, while mycelia favor genes involved in internal signaling and the cell skeleton, which supports filament growth. This broad reprogramming shows that dimorphism is not a minor tweak but a whole-organism reconfiguration.
Duplicated Genes with Split Duties
A key discovery is that many genes come in pairs or small families whose copies have specialized for one form or the other. The team first revisited a known example: two ferroxidase genes and two iron-transporter genes that together import iron, a vital nutrient. One member of each pair is used in yeast, the other in mycelium. Knocking out the yeast-specific copies crippled yeast growth but left mycelial growth mostly intact, and the reverse was true for the mycelial copies. Extending this analysis genome-wide, the researchers identified 490 such “dimorphic families” in which at least one copy is yeast-specific and another is mycelium-specific. Altogether, roughly one in nine genes in the genome belongs to such families, covering many different cellular roles. This suggests that, instead of relying on one version of a protein to function everywhere, the fungus has evolved twin versions tuned to the very different conditions of liquid, low-oxygen yeast life versus solid, oxygen-rich mycelial life.
Back-to-Back Genes and New Control Switches
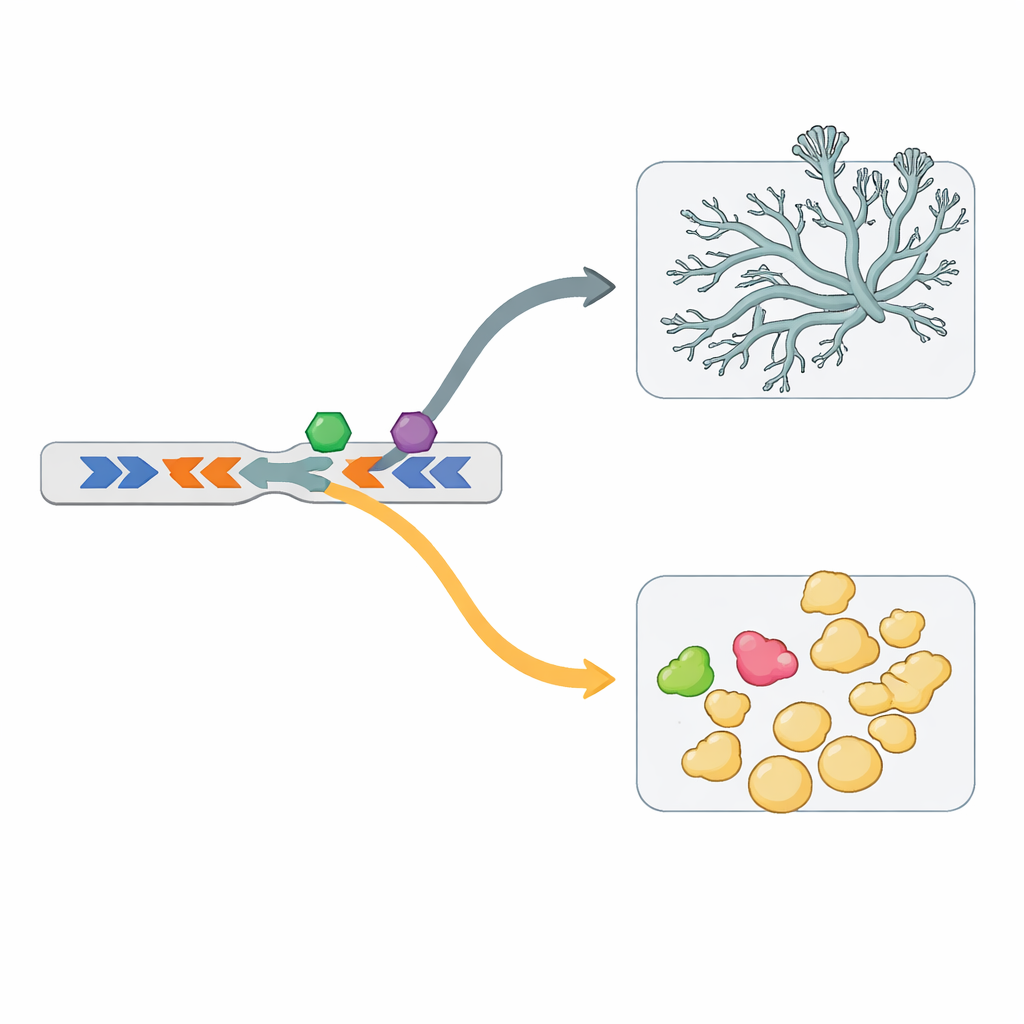
The genome not only duplicates genes but also arranges them in special layouts. Many of the iron-import genes, for example, sit in “head-to-head” pairs: two genes placed in opposite directions, sharing a central control region. One such pair is active in yeast, the neighboring pair in mycelium. Swapping these shared control regions experimentally flipped when each gene was turned on, proving that this arrangement works as a coordinated switch. A survey of the whole genome found over a thousand such head-to-head pairs, with hundreds involved in dimorphism. The shared control regions for yeast- and mycelium-linked pairs carry distinct DNA motifs, implying that they are read by different regulatory factors.
Master Regulators and Evolutionary Clues
To find those regulators, the researchers used the shared control DNA from iron-import genes as bait to fish out proteins that bind there. They identified two previously uncharacterized proteins, named DFL and DKL, and made mutant strains lacking each. These mutants showed strongly disturbed switching: DKL mutants could no longer form yeast at all, and both mutants lost the normal pattern of gene activation across thousands of dimorphism-related genes. Looking across related species, the team saw that dimorphic Mucorales tend to keep duplicated, form-specific gene families, head-to-head structures, and the dfl gene, whereas closely related fungi that do not switch forms often lack these features. This pattern suggests that these genomic traits evolved together as a toolkit for dimorphism and can serve as markers to predict which species are likely to be shape-shifters.
What This Means for Fungal Disease
In simple terms, this work shows that Mucorales fungi have rebuilt their genomes around the challenge of living two separate lives. They solve this by duplicating important genes, tuning one copy for yeast and the other for mycelium, wiring many of them into back-to-back control units, and using dedicated regulators to coordinate which version is used when. Because the invasive, disease-causing form is often the mycelium, and because some of the affected genes handle iron uptake and drug sensitivity, these insights point toward new ways to predict which fungi can become dangerous and to design treatments that disrupt their ability to switch shape.
Citation: Tahiri, G., Navarro-Mendoza, M.I., Lax, C. et al. Coordinated gene family evolution shapes the genome of dimorphic Mucorales. Nat Commun 17, 2148 (2026). https://doi.org/10.1038/s41467-026-68866-7
Keywords: fungal dimorphism, Mucorales, gene duplication, genome regulation, fungal pathogenesis