Clear Sky Science · en
DNA diamond formulates a decomposable composite letter constellation model for DNA data storage
Why future data might live in DNA
Our phones, companies, and scientific instruments are generating data far faster than hard drives and magnetic tapes can grow. DNA—the same molecule that carries genetic information in living things—can also be used to store digital files in an incredibly compact, long‑lasting form. This paper introduces a new way to pack even more information into synthetic DNA strands while keeping it practical and reliable to read back, potentially making DNA storage cheaper and more scalable.
From four DNA letters to richer mixtures
Traditional DNA storage uses the four natural DNA bases—A, T, G, and C—to represent digital bits, much like zeros and ones on a disk. In that scheme, each position in a DNA strand can carry at most two bits of information, because it is limited to one of four choices. The authors build on an emerging idea: instead of placing a single base at each position, they create carefully controlled mixtures of bases, called composite letters. For example, a position might be made of a 50:50 mix of A and T, or a 25:25:25:25 mix of all four bases. When many copies of each strand are synthesized, sequencing these mixtures reveals the base proportions and, in turn, a digital symbol that can represent more than two bits.
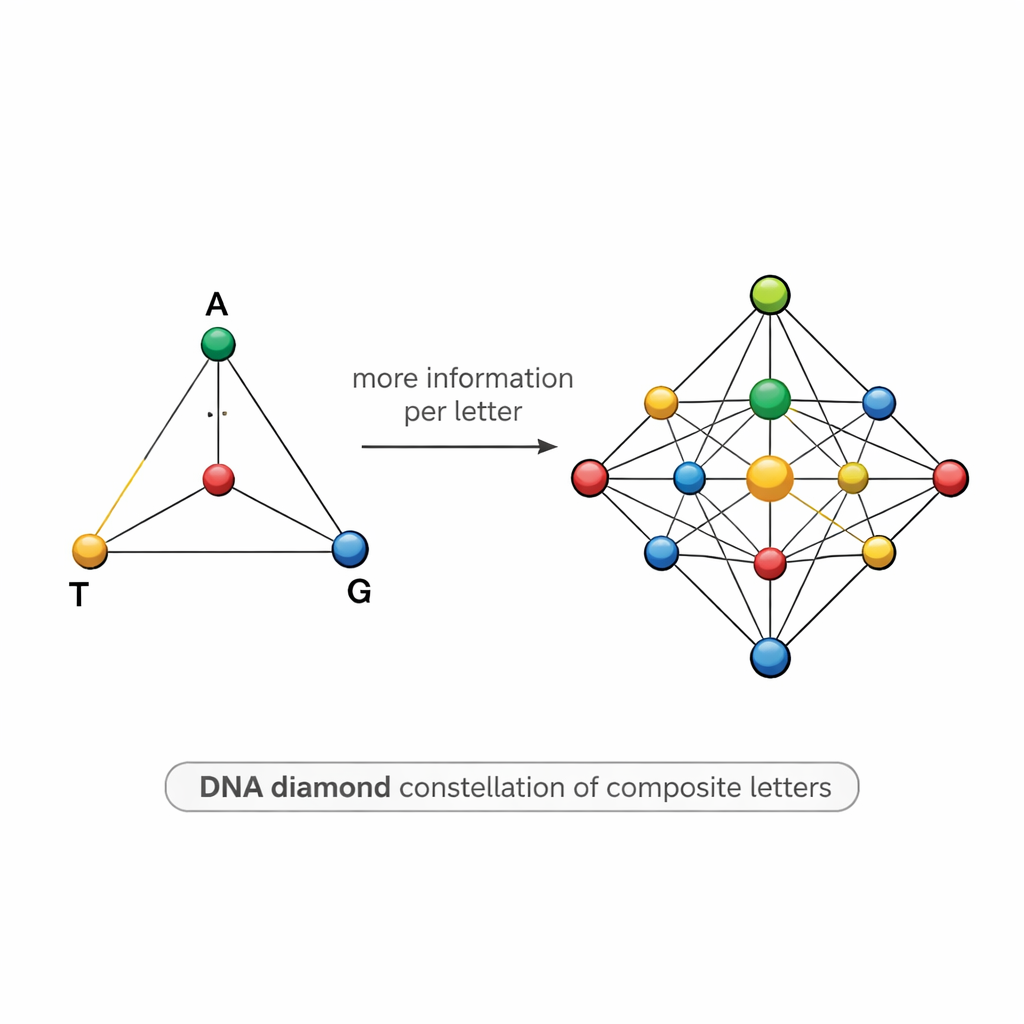
A diamond-shaped map of DNA symbols
Designing such mixtures is tricky. If two symbols are too similar—for instance, one is 50% A and 50% T and another is 55% A and 45% T—sequencing noise can blur them together, causing errors and forcing scientists to sequence far more copies than they would like. To tackle this, the team proposes a structured “DNA diamond” model: a set of 15 composite letters arranged like points on a tetrahedron whose corners are A, T, G, and C. The set includes pure bases at the corners, equal mixtures of two bases along the edges, mixtures of three bases on each face, and a perfectly even mixture of all four bases at the center. This carefully chosen constellation raises the theoretical information per position to about 3.9 bits, while keeping the symbols distinct enough to be told apart in practice.
Smarter decoding with entropy and indexing
Reading data back from DNA means inferring which composite letter was intended at each position from noisy measurements of base frequencies. The authors borrow a strategy from telecommunications called set partitioning. First, they look at how “mixed” a position appears, using a quantity called entropy that is low for pure bases and higher for complex mixtures. This quickly assigns each position to one of four groups: pure bases, two‑base mixes, three‑base mixes, or the four‑base mix. Then, within the chosen group, a more precise likelihood calculation picks the most probable letter. This two‑stage approach reduces confusion between symbols and cuts computation time compared with earlier methods. To further keep strands from being mistaken for one another, each DNA piece carries error‑protected index sequences at both ends, and reads of the wrong length—often caused by insertion or deletion errors—are filtered out before decoding.
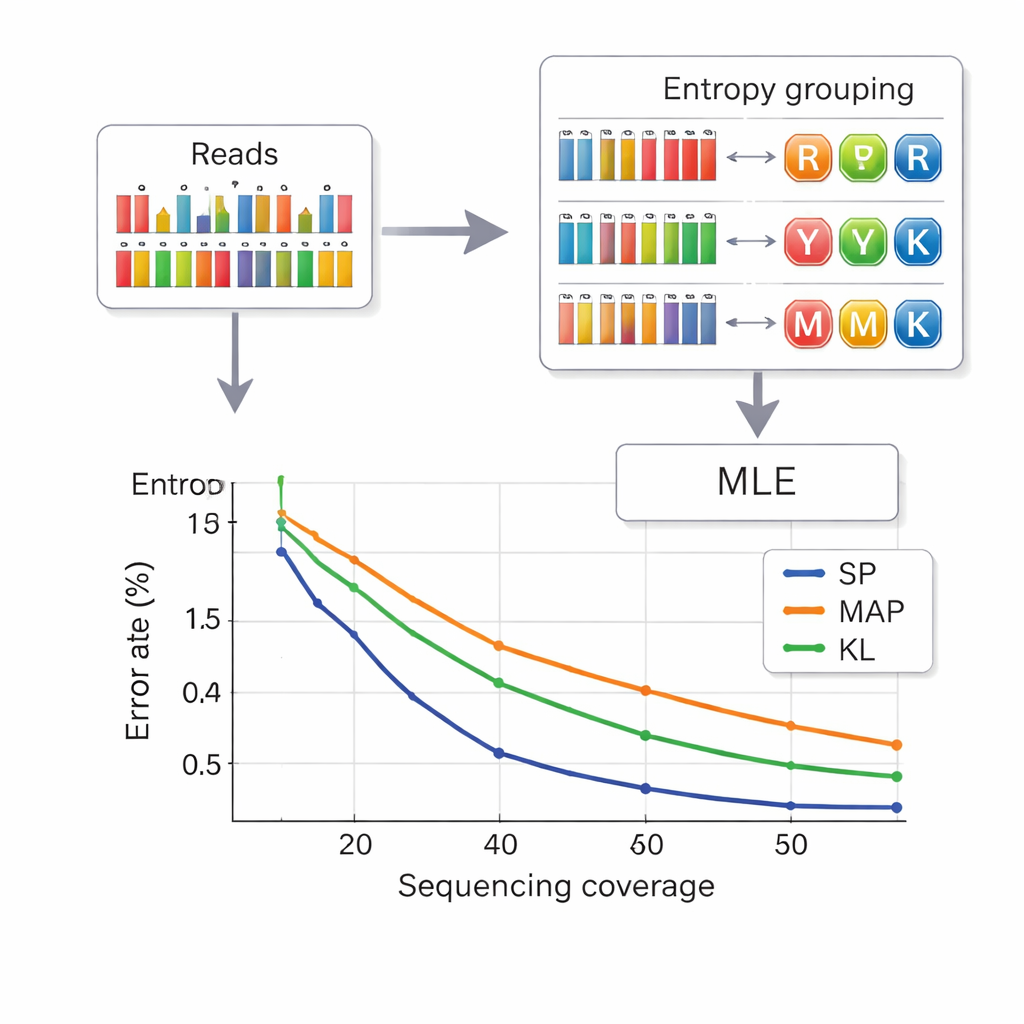
Packing more data with fewer reads
The researchers tested their system in both small and large DNA pools, using commercial synthesis platforms. With an eight‑letter composite alphabet, they reached a payload density of 2.5 bits per DNA position and could recover files perfectly with an average of 14 sequencing reads per strand—better density than earlier six‑letter schemes while needing fewer reads. With the full 15‑letter DNA diamond alphabet, they achieved 3.125 bits per position for the main data and still recovered everything without errors at 33‑fold coverage. Simulations and experiments also showed that their entropy‑based method performs nearly as well as the most accurate, but slower, decoding approach, and clearly better than older techniques, especially at lower sequencing depths.
What this means for future memory
To a lay reader, the key message is that the authors have found a way to teach DNA “new tricks” without inventing new chemistry: by cleverly mixing the existing four bases and decoding them more intelligently, they can store more bits of data per molecule while controlling costs. Their diamond‑shaped alphabet, combined with robust indexing and error correction, shows that high‑capacity DNA data storage is possible with relatively modest sequencing effort. As DNA synthesis and sequencing continue to get cheaper, such designs could help turn DNA from a laboratory curiosity into a realistic medium for archiving the world’s digital memories.
Citation: Ge, Q., Ren, M., Qi, T. et al. DNA diamond formulates a decomposable composite letter constellation model for DNA data storage. Nat Commun 17, 1704 (2026). https://doi.org/10.1038/s41467-026-68861-y
Keywords: DNA data storage, composite letters, information density, error correction, digital archiving