Clear Sky Science · en
Mechanistic insights into PCBP1-driven unfolding of selected i-motif DNA at G1/S checkpoint
DNA Folds That Act Like Traffic Lights
Inside our cells, the genetic code is more than a simple straight ladder of DNA. Parts of it can fold into unusual shapes that act like tiny switches, helping to control when cells copy their DNA and divide. This study focuses on one such shape, called an i-motif, and on a protein named PCBP1 that can recognize and unfold these structures right as a cell prepares to replicate its DNA. Understanding this interaction sheds light on how cells keep their genomes stable and what might go wrong in cancer.
Strange DNA Knots in Cancer-Linked Regions
Most people learn that DNA forms the famous double helix, but certain stretches rich in the letter C (for cytosine) can fold into a four-stranded knot known as an i-motif. These structures tend to appear in control regions of genes that drive cell growth, such as cMYC and BCL2. For years, scientists debated whether i-motifs really form in living cells, because they are easier to see in acidic test-tube conditions than at the near-neutral conditions inside the body. Using specialized antibodies that recognize i-motifs, recent work, including the present study, has confirmed that they do appear in cell nuclei—and often cluster near key growth and cancer genes.
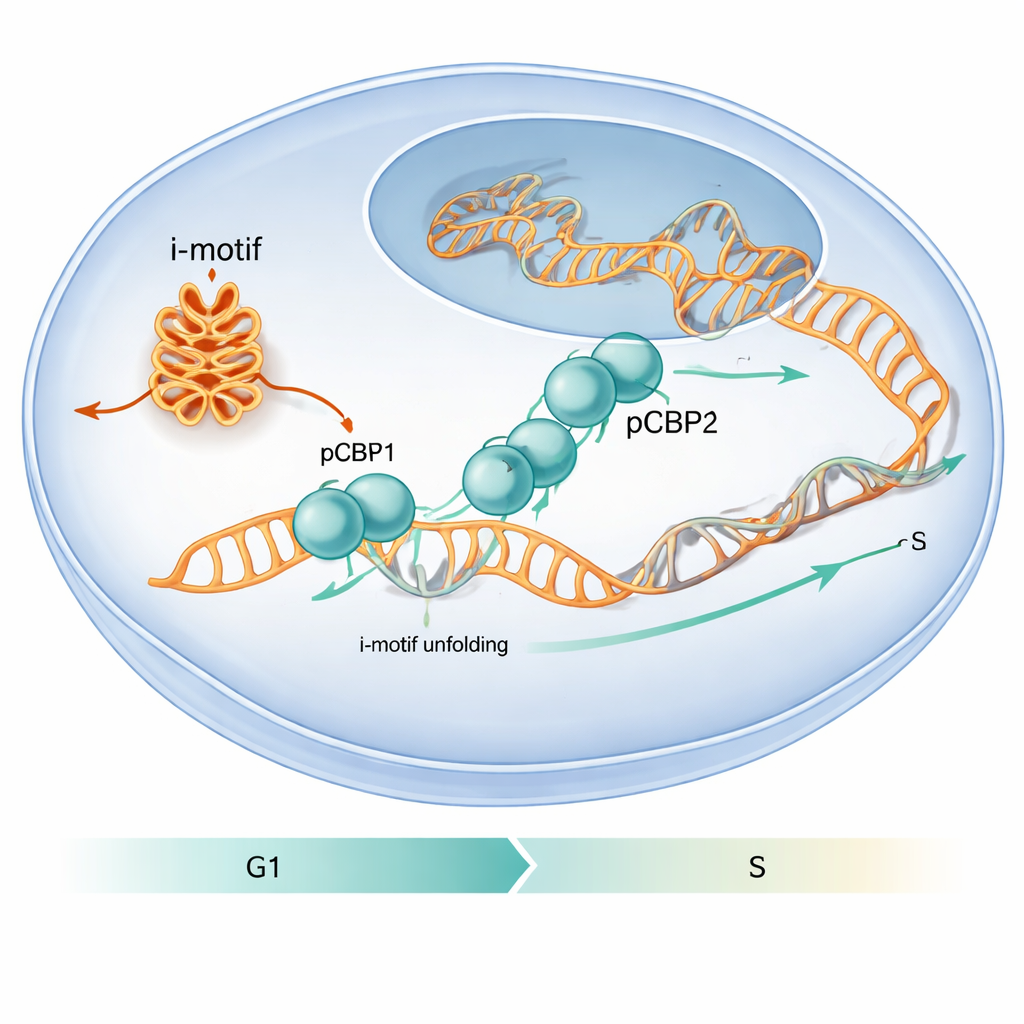
A Protein That Picks Out Special DNA Folds
The authors set out to discover how proteins in the cell handle these unusual DNA folds. They homed in on PCBP1, a protein already known to bind C-rich stretches of DNA and RNA and to influence the cell cycle. By analyzing existing genome-wide binding maps and performing targeted experiments, they found that PCBP1 frequently sits on C-rich regions that can form i-motifs, especially around gene start sites. In cell experiments using human cancer cell lines, regions from the promoters of cMYC, BCL2 and the insulin-linked ILPR sequence showed both strong i-motif signals and strong PCBP1 occupancy, suggesting that PCBP1 is a dedicated guardian of these structures.
How PCBP1 Grabs and Unwinds the Knot
In test-tube assays, the researchers compared how well PCBP1 binds folded i-motifs versus the same DNA in an unfolded form. They tuned the acidity so that the DNA either stayed folded or relaxed, while keeping the protein stable. PCBP1 always preferred the folded i-motif, binding it roughly twice as strongly as the same sequence when unfolded, and only weakly to unrelated DNA shapes. Once bound, PCBP1 could actively promote unfolding, allowing the i-motif strand to pair with its complementary partner. However, not all i-motifs behaved alike: some, such as the structure in the cMYC promoter, were unfolded quickly, whereas others, like those in BCL2, resisted and unfolded only slowly. Extra features such as hairpin loops in the DNA and the degree to which the cytosines were chemically protonated (carrying extra positive charge) could either help or hinder PCBP1’s unwinding activity.
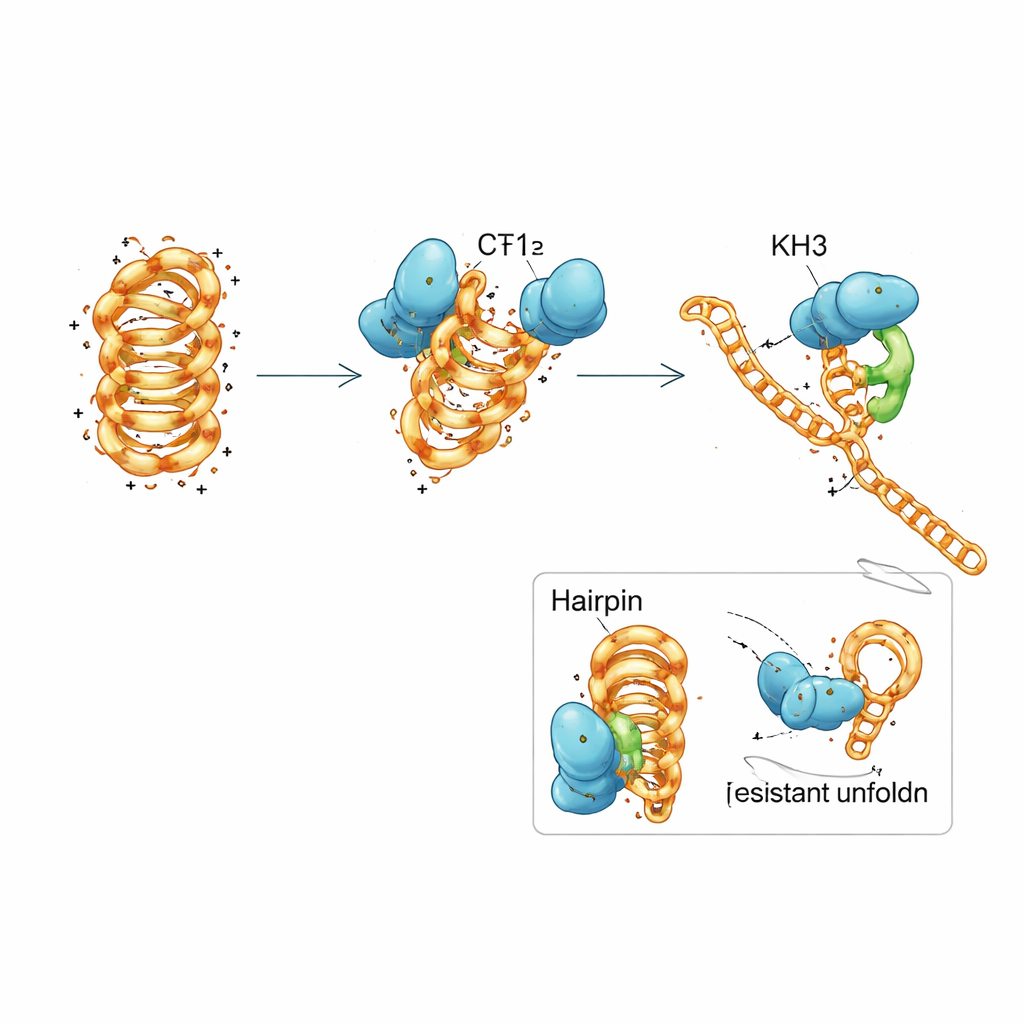
A Team Effort Within a Single Protein
PCBP1 is built from three repeated modules called KH domains, common motifs that grab short stretches of nucleic acids. The team dissected PCBP1 into pieces and discovered that no single KH domain could fully mimic the behavior of the entire protein. The first two domains together could latch onto both folded and unfolded DNA and nudge the i-motif toward a less stable form, but they only slowly promoted full unfolding. The third domain by itself barely bound at all. When all three domains were present and allowed to work together, the protein regained its strong preference for folded i-motifs and its efficient ability to unfold them. Detailed biophysical measurements and computer simulations suggested a stepwise mechanism: KH1 and KH2 first dock onto flexible loop regions of the i-motif and partially disrupt selected base pairs, which then allows KH3 to engage and drive the structure into an open, replication-ready state.
Keeping the Cell Cycle on Schedule
The work also shows that this molecular dance matters for cell behavior. When the researchers reduced PCBP1 levels in human cells, more i-motif structures appeared at specific gene promoters, DNA damage markers increased, and cells stalled at the critical G1/S checkpoint—the moment just before DNA replication begins. Under normal conditions, PCBP1’s presence at i-motif–forming regions peaks around this checkpoint, then declines as S phase starts and the i-motifs are resolved. This timing suggests that PCBP1 acts as a caretaker: it binds and unfolds particular i-motifs at just the right moment so DNA replication can proceed smoothly and the genome remains intact. To a lay reader, the message is that unusual DNA folds can function like temporary roadblocks, and PCBP1 is one of the specialized tools the cell uses to remove them, helping to prevent errors that might otherwise contribute to cancer.
Citation: Sengupta, P., Gillet, N., Obi, I. et al. Mechanistic insights into PCBP1-driven unfolding of selected i-motif DNA at G1/S checkpoint. Nat Commun 17, 1149 (2026). https://doi.org/10.1038/s41467-026-68822-5
Keywords: i-motif DNA, PCBP1 protein, cell cycle checkpoint, genome stability, DNA secondary structure