Clear Sky Science · en
Lipid Nanoparticle Database towards structure-function modeling and data-driven design for nucleic acid delivery
Why tiny fat bubbles matter for future medicines
Lipid nanoparticles are microscopic fat-based bubbles that safely ferry genetic instructions—like mRNA vaccines—into our cells. They helped power COVID-19 vaccines, yet researchers still do not fully understand how their detailed chemical makeup controls how well they work. This article describes a new online resource, the Lipid Nanoparticle Database (LNPDB), created to pull scattered data into one place so scientists can systematically design better, safer gene-delivery medicines.
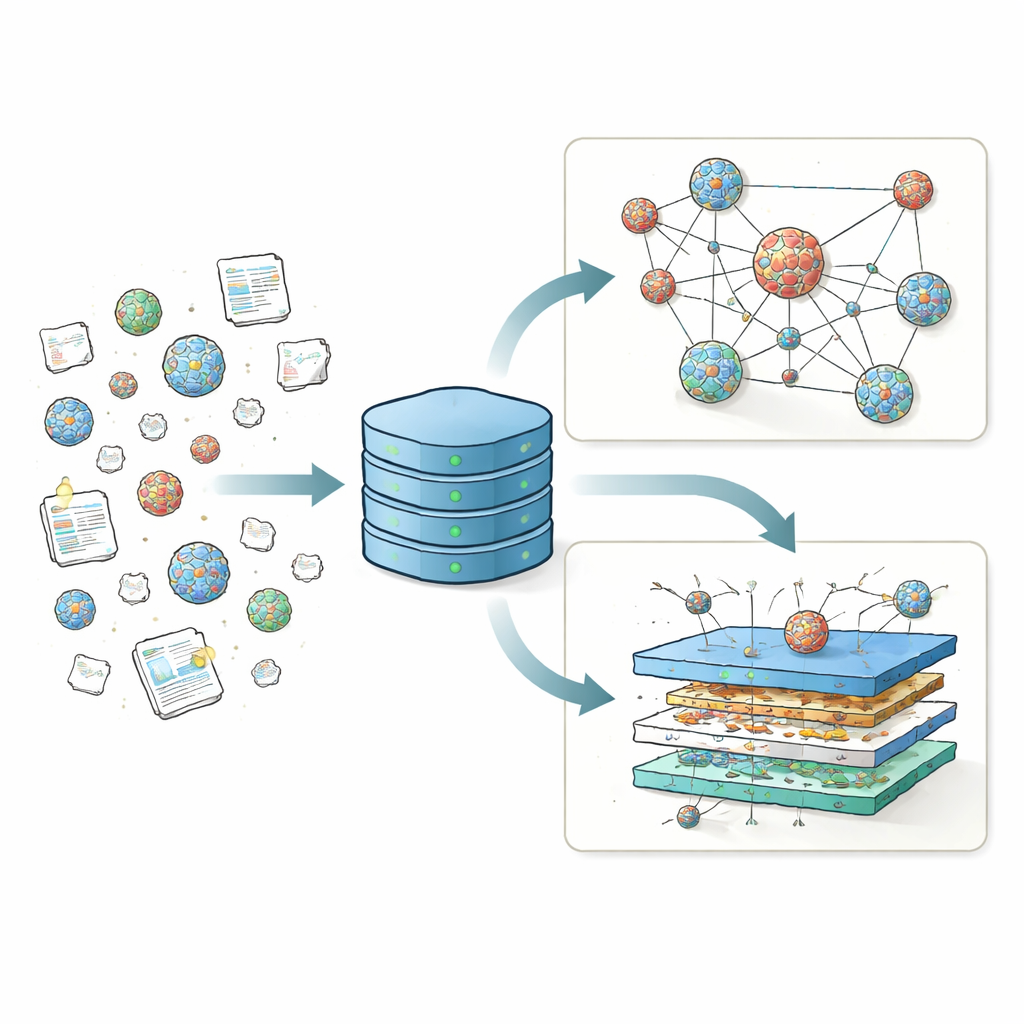
Bringing scattered results into one home
For years, different labs have tested thousands of lipid nanoparticle (LNP) recipes, changing the main charged lipid, helper lipids, cholesterol, and protective coating lipids to see which combinations deliver genetic material most effectively. But these results were reported in many formats across dozens of papers, making it difficult to compare studies or spot big-picture trends. Unlike protein science, which is anchored by a central Protein Data Bank that fueled tools like AlphaFold, the LNP field had no unified repository for its structure and performance data. LNPDB fills this gap by collecting detailed information for 19,528 LNP formulations drawn from 42 studies and a commercial supplier, and by standardizing how each particle’s ingredients, test conditions, and outcomes are encoded.
What lives inside the new database
Each LNP entry in LNPDB is described along three main axes: composition, experiment, and simulation. Composition fields record which lipids were used, how many nitrogen atoms the main charged lipid contains, and the exact mixing ratios between the four core components: ionizable lipid, helper lipid, cholesterol, and a polyethylene glycol (PEG)–lipid. Experimental fields capture what kind of genetic cargo was delivered—most often mRNA coding for a reporter protein—where it was sent (for example, cells in a dish, liver, lung, or muscle), how the particles were prepared, and how success was measured. Finally, simulation fields provide ready-to-use files that describe the physical behavior of each lipid molecule in enough detail to run atom-level computer simulations of lipid membranes. Together, these standardized descriptors turn a patchwork of individual screens into a coherent landscape that can be searched, filtered, and expanded by the community.
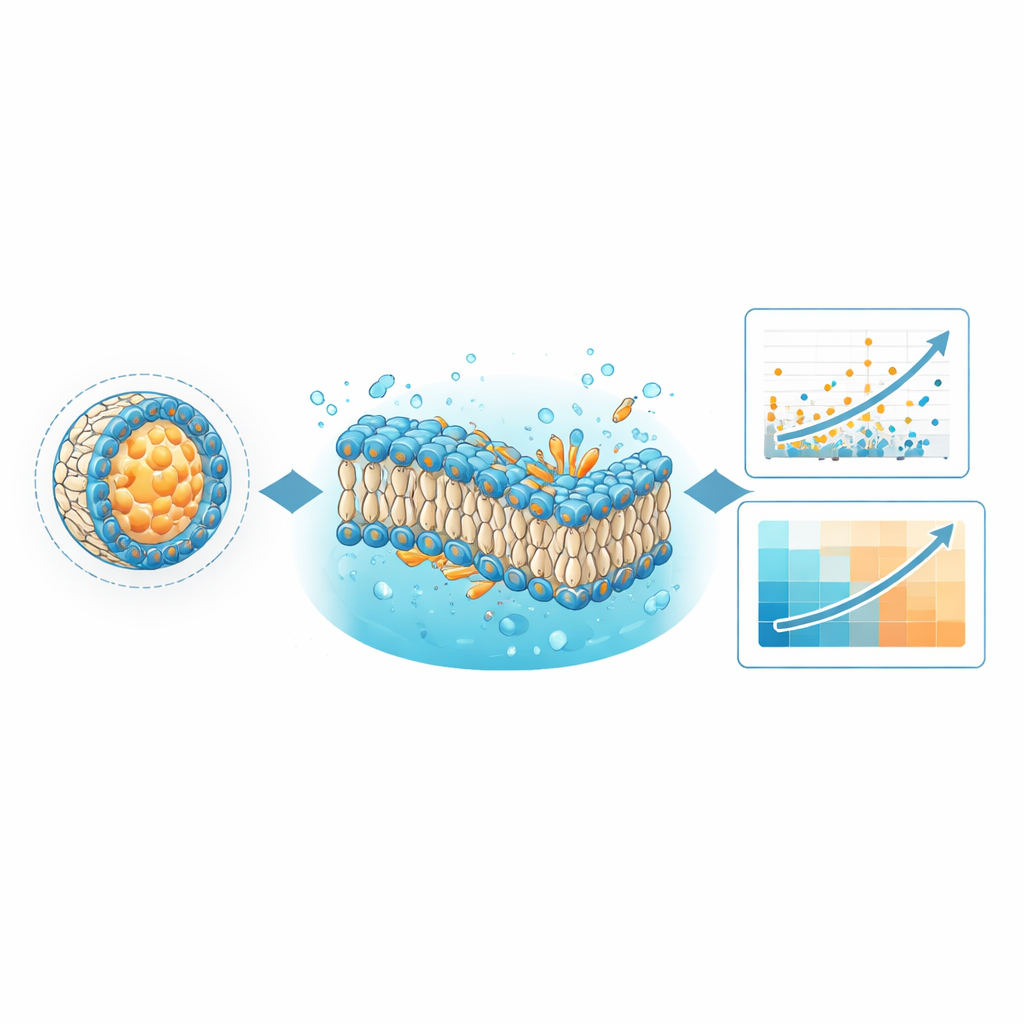
Teaching computers to spot better delivery recipes
One immediate use of LNPDB is to improve machine-learning models that predict which formulations will deliver genetic material most effectively. The authors retrained their existing deep learning model, called LiON, with the expanded LNPDB dataset, more than doubling the number of examples it had previously seen. LiON learns patterns that link the chemical structures of the ionizable lipids, the mix of helper components, and the testing context to how well each formulation performed. With the richer data, LiON’s predictions matched experimental results better for most test sets and outperformed a competing model called AGILE on several independent datasets. This suggests that a broad, diverse, and continuously growing training set is key for building general-purpose design tools for future LNP medicines.
Watching model membranes to uncover hidden rules
The database is also designed for a very different kind of computation: physics-based molecular dynamics simulations. Using the simulation files bundled with LNPDB, the team built simplified membranes representing selected LNP formulations and watched their behavior over microseconds of simulated time. They asked two questions: do the modeled lipid bilayers stay intact, and what overall shape do the key lipids adopt within the membrane? The simulations revealed that formulations whose membranes stayed stable were more likely to succeed experimentally. They also quantified a feature called the “critical packing parameter,” which reflects whether a lipid is more cone-shaped or inverted-cone-shaped in the membrane. In several tested libraries, lipids whose shapes favored negative curvature—thought to help particles fuse with and disrupt endosomal membranes—showed stronger delivery, sometimes correlating with performance better than the deep learning model itself.
A new foundation for smarter nanomedicine
To a non-specialist, the core message is that this work builds a shared, growing “map” of how the ingredients and structure of tiny fat bubbles relate to their ability to deliver genetic therapies. By pulling together tens of thousands of past experiments, enabling powerful prediction models, and providing tools to simulate how particles behave at the molecular level, LNPDB lays the groundwork for more rational design instead of trial-and-error tinkering. Over time, this kind of data-driven approach could speed the creation of more effective vaccines, gene-editing treatments, and other nucleic acid–based therapies, while helping researchers understand why certain nanoparticle recipes work—and others do not.
Citation: Collins, E., Ji, J., Kim, SG. et al. Lipid Nanoparticle Database towards structure-function modeling and data-driven design for nucleic acid delivery. Nat Commun 17, 2464 (2026). https://doi.org/10.1038/s41467-026-68818-1
Keywords: lipid nanoparticles, mRNA delivery, nanomedicine, machine learning, molecular dynamics