Clear Sky Science · en
The functional landscape of alternative splicing in hematopoietic lineage commitment
How tiny edits in genes shape our blood
Every second, your body makes millions of new blood cells. Behind this quiet miracle lies a molecular editing system that can cut and paste pieces of genetic messages in different ways, creating slightly different versions of the same protein. This study explores how that editing process, called alternative splicing, helps steer immature stem cells toward becoming red blood cells, white blood cells, or other blood cell types—and what happens when a single edited piece is missing.
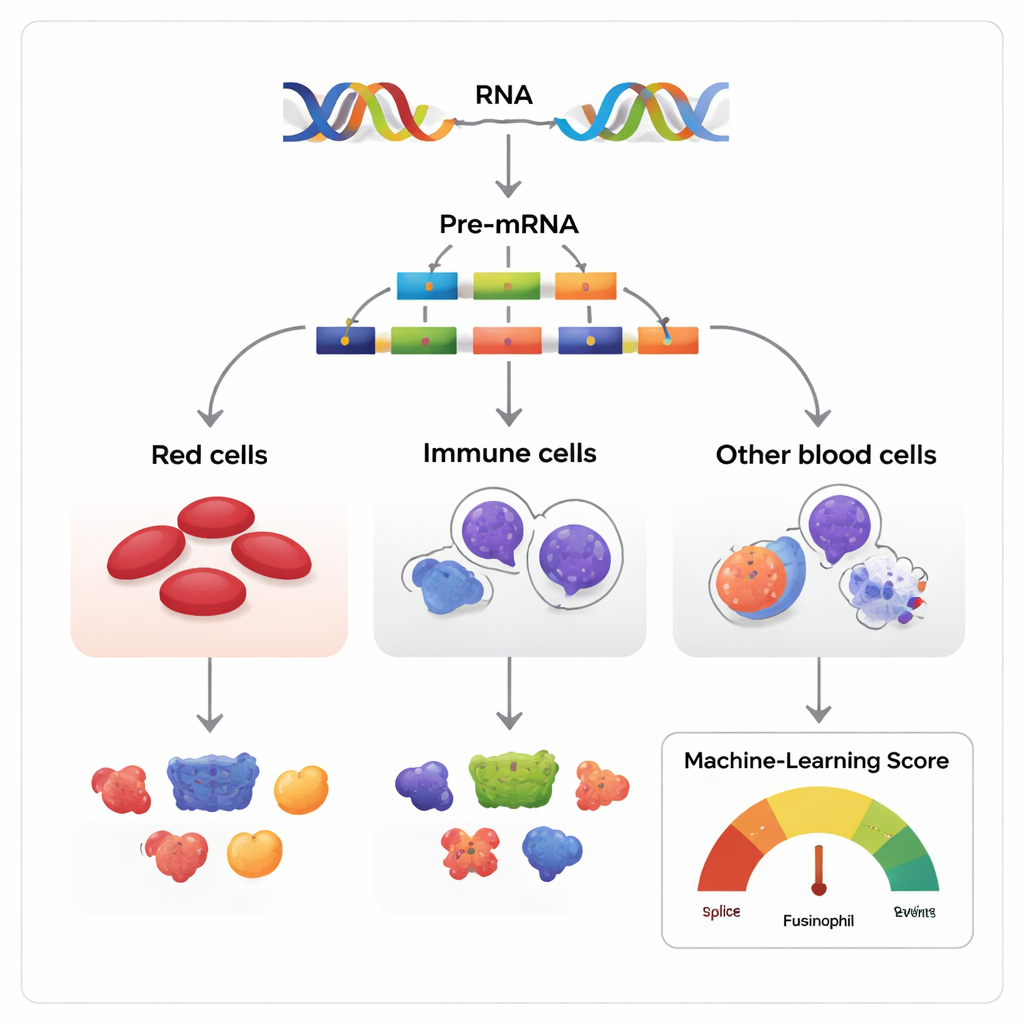
Many versions from the same genetic script
Genes are often described as blueprints, but in reality they are more like scripts that can be rearranged. When a gene is read, the initial RNA copy contains segments called exons that can be either kept or skipped before the final message is turned into protein. The authors looked at this process, known as exon skipping, across blood-forming tissues in humans, mice, and several other vertebrates. They assembled more than 270 RNA datasets that together follow blood stem and progenitor cells as they mature into three major families: red cell–producing erythroid cells, infection-fighting myeloid cells, and antibody-producing lymphoid cells.
Ranking which splice changes really matter
Because most multi-exon genes can be spliced in many ways, the central challenge is to distinguish harmless variations from those that truly influence cell fate. The researchers built a machine-learning model, called the Functional AS Score (FAScore), to tackle this problem. For each exon-skipping event, the model considers 19 pieces of information, such as how strongly its usage changes during cell development, how well the surrounding DNA is conserved across species, whether it alters known protein domains, and whether it contains sites for chemical modifications of the protein. The algorithm, trained using a positive–unlabeled strategy and a random forest classifier, outputs a score between 0 and 1 indicating how likely a given splice event is to have a functional impact.
Finding conserved, lineage-specific switches
Applying FAScore to tens of thousands of exon-skipping events, the team classified them into likely functional, non-functional, or uncertain groups. Events predicted to be functional were more often found in protein regions important for interactions, in evolutionarily conserved sequences, and in segments that host chemical tags like phosphorylation or SUMOylation. Many of these events were also active only in specific blood lineages or during fetal blood formation, suggesting they act as fine-tuned switches in particular developmental windows. The study further showed that some of the most ancient splice events—those shared across vertebrates for hundreds of millions of years—are especially likely to be functional, mirroring how older genes tend to carry core biological roles.
Experimental proof: tuning blood cell outcomes
To test the model’s predictions, the authors chose several high-scoring exons in genes active in different blood lineages and deleted those exons in mouse blood stem and progenitor cells using CRISPR. The results matched the model’s expectations: removing certain exons in the KLF6 and SSBP3 genes impaired the formation of myeloid colonies without harming red cell output, whereas deleting exons in EPB41L1 and TBC1D23 altered erythroid colony formation. In particular, skipping exon 15 of TBC1D23 reduced the production of red blood cell precursors in mice and zebrafish, leading to fewer circulating red cells and lower hemoglobin levels, while largely sparing white blood cells.
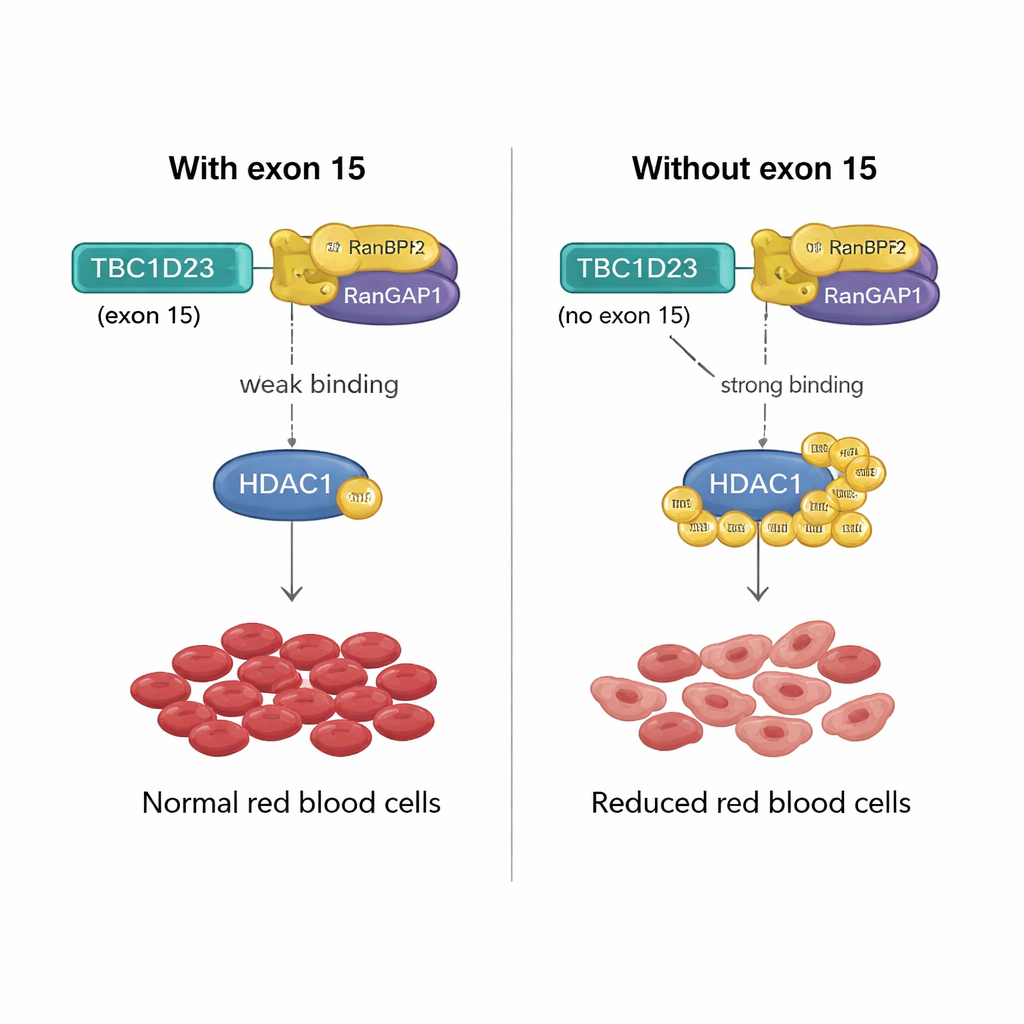
A molecular lever for red blood cell production
How can a 15–amino acid stretch encoded by exon 15 of TBC1D23 have such an effect? The team found that including this exon weakens TBC1D23’s binding to a protein pair called RANBP2/RANGAP1, which collaborates to attach SUMO tags to other proteins. Without exon 15, TBC1D23 binds this pair more strongly, boosting SUMOylation of a key enzyme, HDAC1. This heightened tagging changes the activity of many transcription factors—master regulators of gene expression—throwing off the gene programs needed for proper maturation of red blood cells. When the researchers engineered a version of HDAC1 that cannot be SUMO-tagged at two crucial positions, it rescued red cell formation in cells lacking TBC1D23 exon 15, confirming that this chemical tagging is the critical step.
Why this matters for health and future therapies
To a non-specialist, the message of this work is that not all genetic changes are created equal: sometimes, the difference between healthy blood and anemia lies in whether a tiny segment of a gene is kept or skipped in the final message. By combining large-scale RNA data with a sophisticated scoring system, the study provides a roadmap for spotting which splice variants are most likely to influence how stem cells choose their fates. This approach not only deepens our understanding of how blood cells are formed in health and disease, but also offers a general strategy for pinpointing important splice events in other organs, potentially guiding future gene-based therapies and precision treatments.
Citation: Hu, X., Wang, J., Chen, L. et al. The functional landscape of alternative splicing in hematopoietic lineage commitment. Nat Commun 17, 2047 (2026). https://doi.org/10.1038/s41467-026-68811-8
Keywords: alternative splicing, hematopoiesis, machine learning, red blood cells, gene regulation