Clear Sky Science · en
Structural basis for transcriptional regulation by the cell division regulator MraZ in Mycoplasma genitalium
How a Tiny Bacterium Controls When It Divides
Every living cell must decide when to split in two, and that decision can mean the difference between healthy growth and runaway infection. This study looks inside one of the simplest known bacteria, Mycoplasma genitalium, to reveal in atomic detail how a single protein, called MraZ, latches onto DNA and switches key cell-division genes on or off. By understanding this minimalist control system, scientists hope to uncover general rules of bacterial growth that could one day inspire new kinds of antibiotics or synthetic “minimal cells.”
A Stripped-Down Cell with Big Lessons
Mycoplasma genitalium is famous for its tiny genome: just a fraction of the DNA found in common bacteria like E. coli. That small size makes it a powerful model for figuring out which genes and control systems are truly essential for life. Many bacteria keep their cell-division and cell-wall genes together in a block called the dcw cluster. In wall-less mycoplasmas, most of those genes are gone, but a handful remain, including mraZ at the very front of the cluster. MraZ acts as a traffic cop, controlling the activity of its neighboring genes, which in turn influence how and when the cell divides.
A Repeated DNA Pattern as a Control Switch
Just upstream of the mraZ gene, the researchers found a highly conserved stretch of DNA that acts as a docking station for the MraZ protein. This region contains four short repeated segments, or “boxes,” that have a nearly identical sequence across many bacterial species. By carefully mutating one, two, or more of these boxes and then measuring how tightly MraZ could still grab the DNA, the team showed that the protein binds cooperatively: each box helps strengthen the overall grip. Reporter experiments using a fluorescent marker confirmed that the more these boxes were disrupted, the less effectively MraZ could repress gene activity, underscoring their importance as a finely tuned control panel. 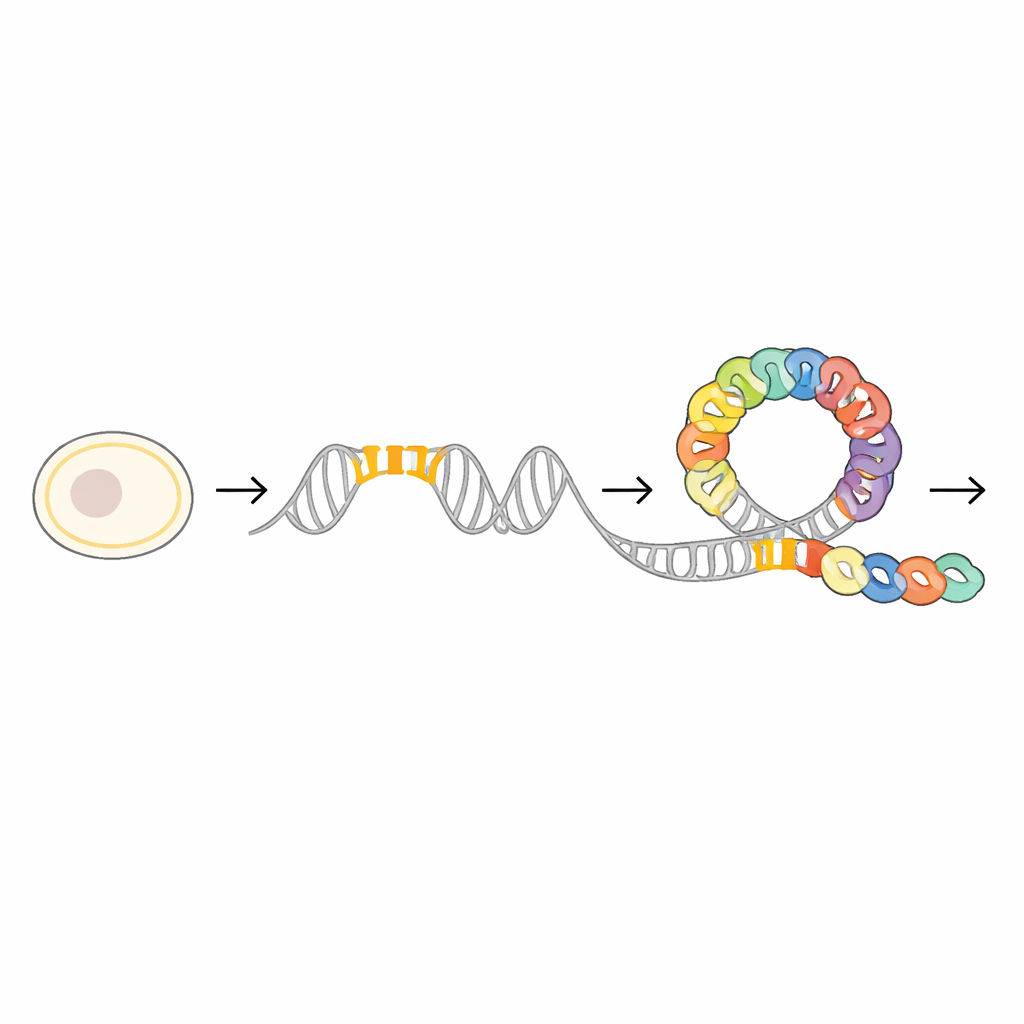
A Ring-Shaped Protein That Opens to Embrace DNA
To see how this control panel works at the atomic level, the scientists used cryo-electron microscopy and X-ray crystallography to solve several three-dimensional structures of MraZ alone and bound to DNA. On its own, MraZ molecules join together into ring-like assemblies made of eight or nine identical subunits. These rings have a distinctive “cradle-like” surface formed by a small beta-sheet structure, unlike the spiral helices often seen in DNA-binding proteins. When MraZ encounters its four-box DNA segment, the ring does not simply sit on top of the helix; instead, it flexes open and rearranges so that four of its subunits line up along the DNA, each cradling one box in the major groove.
Key Contact Points That Read the DNA Code
The high-resolution structures revealed that each DNA-contacting subunit uses three positively charged side chains—specific points on the protein surface—to read the DNA sequence. These chemical “fingers” reach into the groove of the double helix and form precise hydrogen bonds with particular base pairs in the conserved boxes. When the researchers changed any one of these three residues, MraZ largely lost its ability to bind DNA and to shut down the test reporter gene. Additional contacts with the DNA backbone help stabilize the complex but are less sequence-specific. Together, these findings show how MraZ combines a highly specialized reading head with a flexible multi-subunit body to recognize its target region. 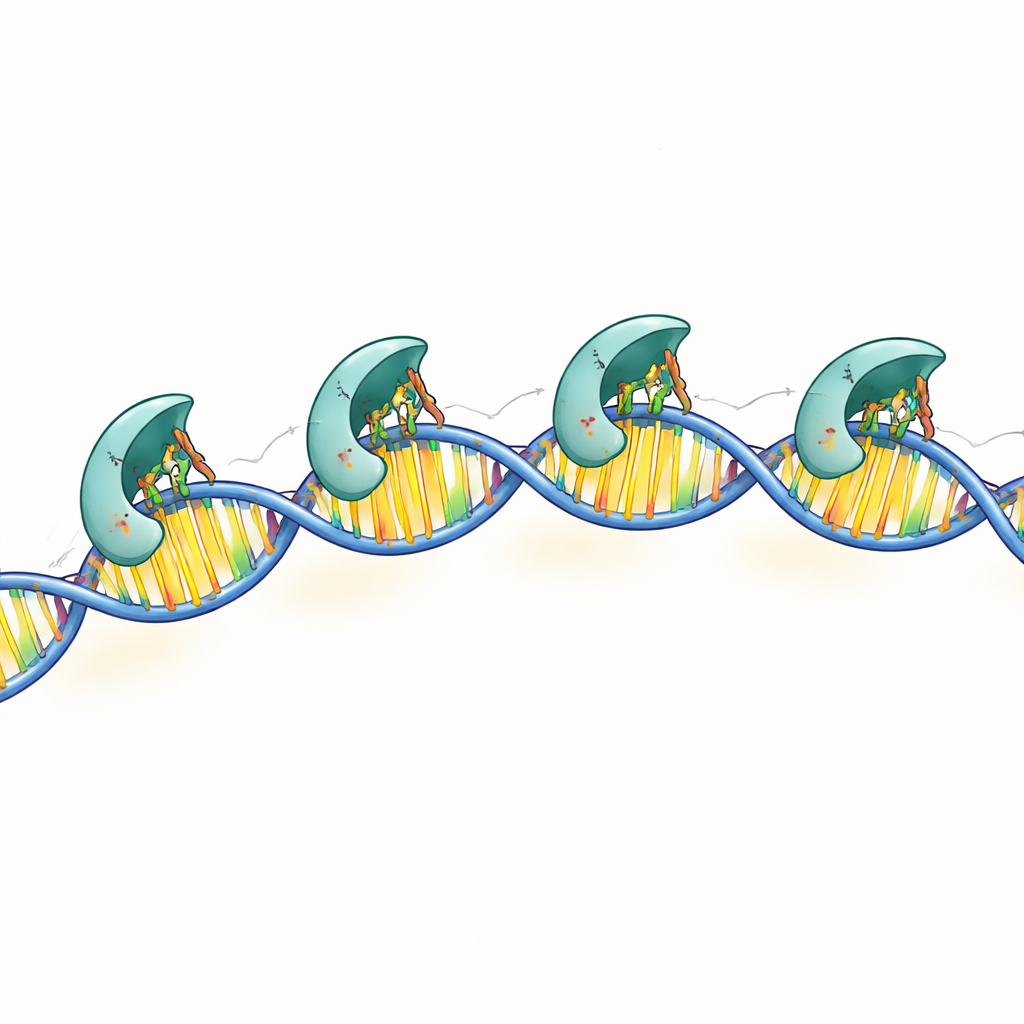
Oligomers That Tune the Strength of Control
Because MraZ forms rings and other multi-unit shapes, the team asked whether this clustering is required for DNA binding or simply fine-tunes it. By engineering a version of MraZ that could no longer assemble into rings, they found that the protein still bound the four-box DNA sequence, but with weaker affinity. This monomeric form struggled especially when the spacing between boxes was altered, suggesting that the full oligomer helps bridge and align distant boxes, raising the local concentration of binding motifs along the DNA. The authors propose a dynamic model in which MraZ toggles between a closed ring and an open, DNA-engaged form, using its assembly state as a dial to adjust how strongly it clamps down on the promoter.
What This Means for Bacteria and Beyond
In simple terms, this work explains how a small protein in a minimal bacterium grabs a repeated pattern on DNA and uses it as a master switch for cell-division genes. The combination of a cradle-like reading head and a flexible ring body allows MraZ to recognize its target with high precision while remaining adaptable to different DNA arrangements. Because similar proteins and DNA motifs appear across many bacteria, the mechanism uncovered here is likely a shared strategy for coordinating growth and division. Insights from this pared-down system may help researchers design streamlined genetic circuits in synthetic cells and, in the long term, could contribute to new ways of disrupting bacterial growth in disease.
Citation: Sánchez-Alba, L., Varejão, N., Durand, A. et al. Structural basis for transcriptional regulation by the cell division regulator MraZ in Mycoplasma genitalium. Nat Commun 17, 2132 (2026). https://doi.org/10.1038/s41467-026-68809-2
Keywords: bacterial cell division, DNA–protein interaction, transcriptional regulation, cryo-electron microscopy, Mycoplasma genitalium