Clear Sky Science · en
Phased-assembly-driven pangenome graphs for structural variant genotyping and complex trait mapping in dairy cattle
Why cow genetics matter to your glass of milk
Dairy cows are the unseen engines behind milk, cheese, and yogurt. Yet even within a single breed like Holsteins, no two animals share exactly the same DNA. Much of that hidden variation comes not from tiny spelling changes in genes, but from larger additions, deletions, and rearrangements of DNA. This study shows how a new kind of cattle reference genome, called a pangenome graph, can capture that big, structural DNA diversity and link it to important traits such as milk yield, body size, fertility, and disease resilience.
Looking beyond one “standard” cow genome
For years, genetic studies in humans and livestock have leaned on a single reference genome as a map. That approach works reasonably well for single-letter DNA changes but misses many larger structural variants, which can span dozens to millions of DNA bases. These larger changes are especially common in regions that are hard to sequence, such as repetitive stretches near chromosome ends. In cattle, such structural variants are already known to affect milk production, growth, reproduction, and health, but traditional short-read sequencing and single-reference maps leave much of this variation invisible.
Building a richer DNA map for Holstein cattle
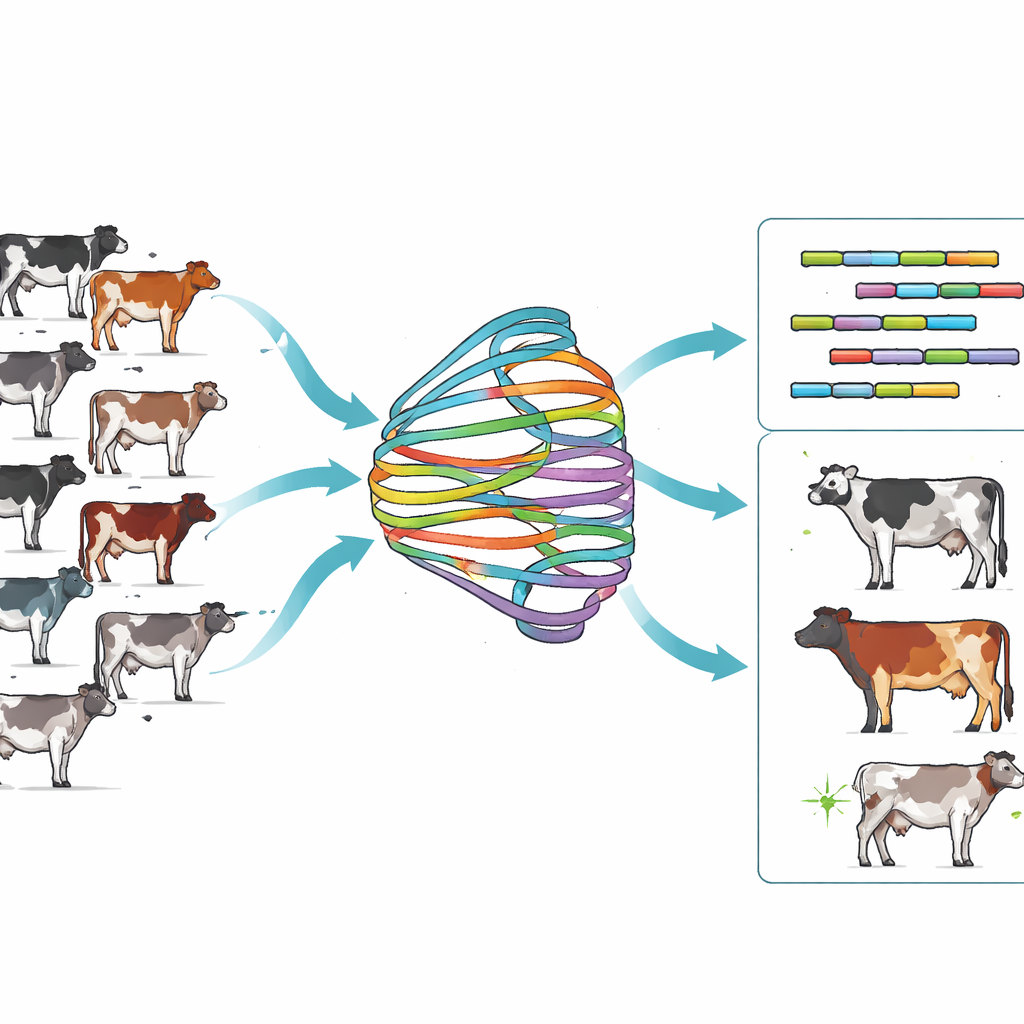
The researchers set out to build a much more complete genetic map for Holsteins, the world’s dominant dairy breed. They used long-read sequencing to generate 40 haploid genome assemblies from 20 Holstein cows and then combined them with a method called Minigraph-Cactus to construct a breed-specific pangenome graph named H20D. Instead of a single linear DNA sequence, this graph holds a shared “core” that most cows have plus many alternative branches that capture insertions, deletions, and complex rearrangements. About 95% of the sequence was shared across all animals, but the remaining 5% contained variable and even unique segments that would be overlooked in a single reference. When the team compared H20D to a cross-breed cattle graph built from 13 breeds, they found the Holstein-focused graph was less tangled yet still rich in breed-relevant variation, especially larger and more complex structural differences.
Finding more meaningful variants, more accurately
To test whether this new map actually improves genetic analysis, the authors compared H20D-based structural variant calls to a suite of popular tools that work either from assembled genomes or directly from read alignments. Using the pangenome as a benchmark, the within-breed, fully phased graph consistently outperformed both long-read and short-read methods alone, identifying roughly ten thousand additional structural variants per animal. Diploid (two-copy) graphs built from phased assemblies captured many more variants and produced more accurate genotypes than graphs built from single, unphased assemblies. The advantages were strongest in problematic regions rich in repeats, where other methods often disagreed or failed. Crucially, when the team used the H20D graph as a reference for a short-read genotyping tool called PanGenie, they could recover a large fraction of the long-read discoveries—far more than with traditional short-read structural variant callers.
From DNA structures to real-world dairy traits
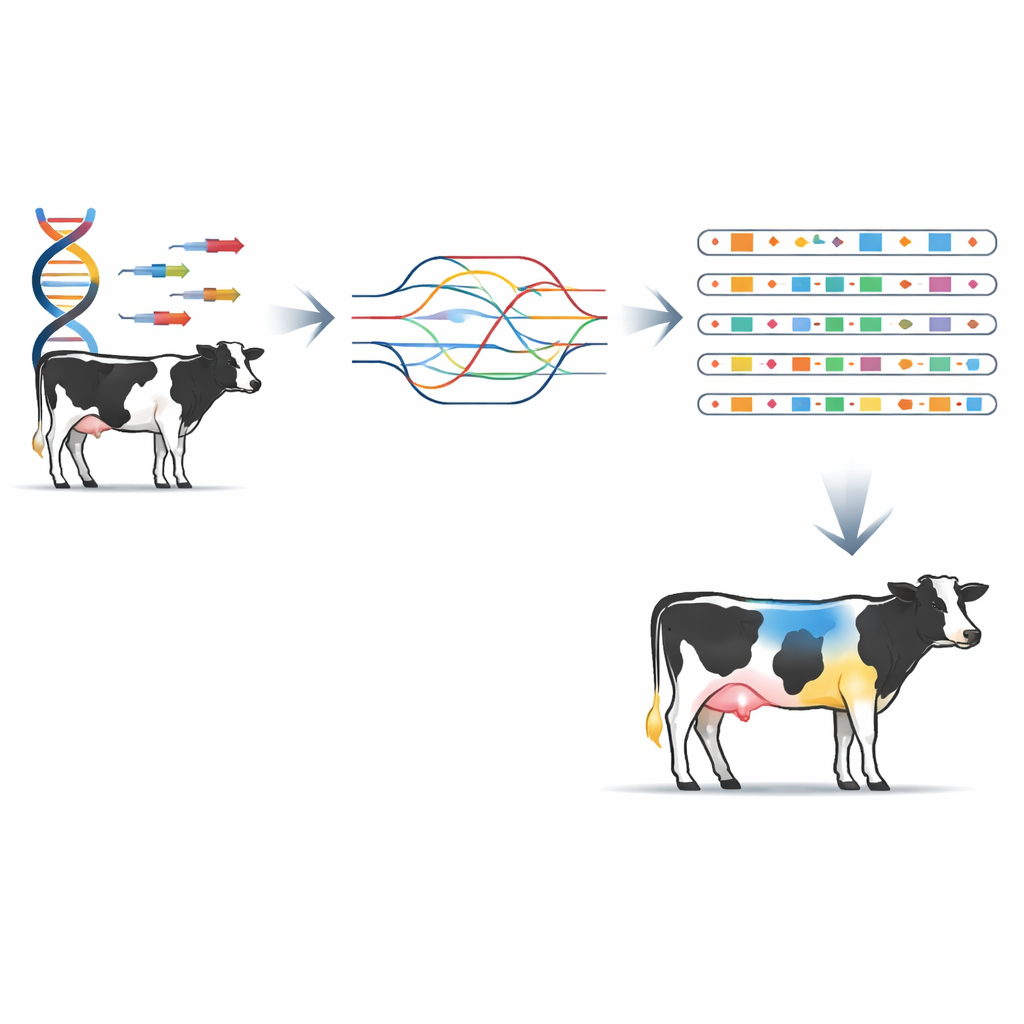
Armed with this detailed structural map, the researchers then turned to real animals and traits. They genotyped structural variants in 173 Holstein cattle with rich performance records and ran genome-wide association studies across 46 traits spanning milk production, body form, fertility, health, and longevity. They uncovered 196 significant associations, involving 135 structural variants tied to 42 traits. In many genomic regions, structural variants lined up with known single-letter signals but showed stronger statistical support, suggesting they may be closer to the actual biological causes. For example, a sizeable deletion overlapping a gene called MATN3 was linked to stature and may alter bone development, while an insertion near the EPPK1 gene in fat and brain tissues was associated with milk fat percentage, hinting at effects on fat metabolism or secretion.
What this means for future herds
This work shows that pangenome graphs built from phased assemblies within a single breed can greatly sharpen our view of cattle genetics. By capturing structural variants that standard references miss and tying them directly to economically important traits, these maps promise more precise breeding decisions. In practice, that could mean selecting bulls and cows not just on thousands of single-letter markers, but also on the larger DNA segments that influence milk yield, efficiency, health, and resilience. As long-read sequencing and pangenome tools become more accessible, similar approaches could accelerate genetic improvement in many livestock species, ultimately shaping healthier herds and more sustainable dairy production.
Citation: Yang, L., Gao, Y., Kuhn, K.L. et al. Phased-assembly-driven pangenome graphs for structural variant genotyping and complex trait mapping in dairy cattle. Nat Commun 17, 2186 (2026). https://doi.org/10.1038/s41467-026-68807-4
Keywords: cattle pangenome, structural variants, Holstein dairy, genome-wide association, precision breeding