Clear Sky Science · en
HIV-seq reveals gene expression differences between HIV-transcribing cells from viremic and suppressed people with HIV
Why this research matters for people living with HIV
Modern HIV treatment can drive the virus in blood to undetectable levels, turning a once-fatal infection into a chronic, manageable condition. Yet if treatment stops, the virus almost always rebounds. This paper explores a hidden group of infected immune cells that keep HIV alive under the radar, and introduces a new technology, "HIV‑seq," that lets scientists finally see what these cells are doing one by one. Understanding these stealthy cells could inform future strategies aimed not just at controlling HIV, but possibly at curing it.
Finding the virus hiding in plain sight
HIV persists in the body by embedding its genetic material in long‑lived immune cells, especially CD4 T cells. Some of these infected cells stay silent, while others actively make bits of viral RNA or protein even when blood tests say the virus is undetectable. These “active reservoir” cells may drive chronic inflammation and spark rapid viral rebound when treatment is interrupted. However, standard single‑cell RNA sequencing often misses them, because HIV transcripts can be rare and frequently lack the poly‑A tail that the technology normally grabs onto. The authors set out to solve this detection problem so they could compare HIV‑producing cells in people with uncontrolled infection (viremia) versus those on successful antiretroviral therapy (ART).
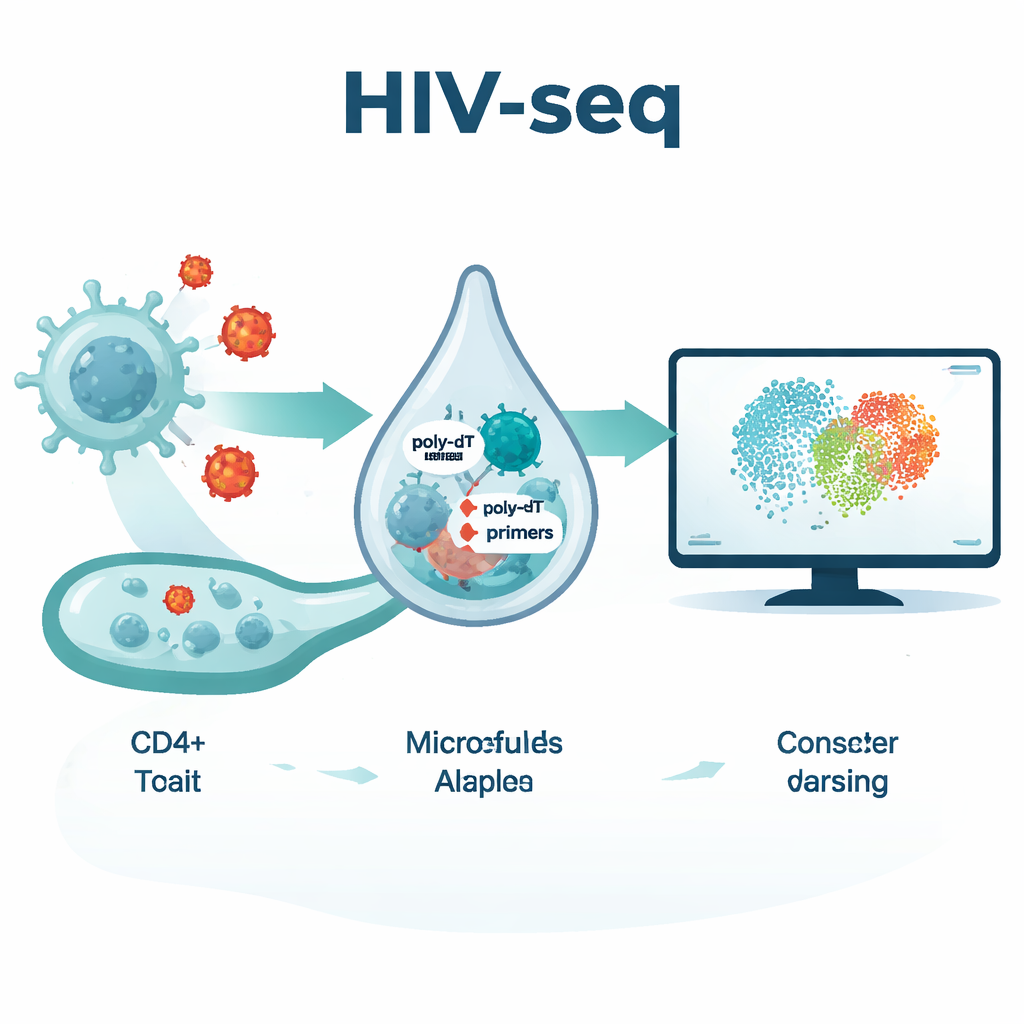
A new lens: the HIV‑seq method
The team developed HIV‑seq by adding several carefully designed HIV‑specific capture primers to the usual single‑cell sequencing chemistry. These extra primers latch onto conserved regions of the HIV genome, boosting recovery of viral RNA whether or not it carries a poly‑A tail. They combined this with DNA‑barcoded antibodies that simultaneously measure dozens of surface proteins, giving each sequenced cell a rich molecular profile. In head‑to‑head tests on blood cells from people with untreated HIV, HIV‑seq roughly doubled the number of viral reads per infected cell without distorting the host cell’s gene expression patterns or producing false signals in HIV‑negative donors. This allowed the researchers to define HIV‑RNA‑positive cells as those with at least one confidently detected viral transcript and to map where those cells sit within the broader T‑cell landscape.
What HIV‑producing cells look like during active infection
Applying HIV‑seq to four people with untreated HIV, the authors analyzed more than 85,000 CD4 T cells and identified 1,072 cells actively transcribing HIV. These infected cells were rarely naïve cells; instead, they clustered mainly among effector memory T cells that have already been activated by prior immune encounters. A striking subset showed a cytotoxic or “killer” profile, expressing genes for granzymes, perforin, and other molecules typically associated with cells that destroy infected targets. At the same time, HIV‑positive cells had reduced levels of several natural antiviral defenses and restriction factors, suggesting an internal environment more favorable to viral replication. Pathway analyses showed heightened activity of signaling circuits known to drive HIV gene expression, including NFAT and protein kinase C, as well as chemokine pathways that could influence where these cells travel in the body.
How HIV‑producing cells change under successful treatment
The researchers then examined three of the same individuals after at least six months of effective ART, when virus in the blood was suppressed. As expected, HIV‑producing cells were much rarer, but HIV‑seq could still detect 25 such cells among over 75,000 CD4 T cells. These cells were again enriched among effector memory T cells, but their character was different: they no longer clustered in the highly cytotoxic group. Instead, they were concentrated in memory T‑cell subsets marked by the IL‑7 receptor and other features of long‑lived, self‑renewing cells. Many expressed the survival protein BCL‑2, and their gene signatures pointed to activation of TGF‑β–related pathways that are known to dampen inflammation and may help keep HIV in a low‑activity, harder‑to‑detect state. Compared to the vigorous antiviral and inflammatory responses seen during viremia, HIV‑positive cells under ART showed an anti‑inflammatory, more “tolerant” profile.
Implications for future HIV cure strategies
To a lay observer, these results paint a picture of HIV as a shape‑shifter: during untreated infection, infected cells look like short‑lived fighters that both spread the virus and fan the flames of inflammation; once ART quiets active replication, the remaining HIV‑producing cells resemble long‑lived survivors that hide behind calming, anti‑inflammatory signals and strong survival programs. HIV‑seq provides a powerful new way to track these elusive cells at single‑cell resolution, helping researchers pinpoint which cellular pathways sustain them and how they might be exposed or eliminated. By clarifying how HIV‑transcribing cells differ between active infection and drug‑suppressed states, this work offers concrete clues for designing “shock‑and‑kill” or “block‑and‑lock” approaches that could one day reduce, or even clear, the viral reservoir.
Citation: Frouard, J., Telwatte, S., Luo, X. et al. HIV-seq reveals gene expression differences between HIV-transcribing cells from viremic and suppressed people with HIV. Nat Commun 17, 1540 (2026). https://doi.org/10.1038/s41467-026-68797-3
Keywords: HIV reservoir, single-cell sequencing, antiretroviral therapy, viral latency, immune memory T cells