Clear Sky Science · en
CRISPR-Cas9 trans-cleavage is hindered by a flanked R-loop, an elongated spacer, and an inactive HNH domain
Why tiny DNA cuts matter
CRISPR-Cas9 is famous as a molecular scalpel that can cut DNA at chosen sites, but this tool has a second, less appreciated behavior: once activated, it can also nibble on other bits of genetic material nearby. Understanding when this “collateral” cutting turns on or off is crucial for making safer gene-editing therapies and more sensitive diagnostic tests. This study dissects the physical features of the Cas9–DNA–RNA complex that determine whether Cas9 quietly makes its intended cut, or also starts chewing through stray single strands of DNA.
How the CRISPR scissors get started
To act, Cas9 binds a short piece of guide RNA that steers it to a matching DNA sequence in a genome. When Cas9 finds its target, the guide RNA pairs with one DNA strand, forcing the two strands apart and creating a DNA–RNA hybrid region called an R-loop. In its classic role, Cas9 then cuts both DNA strands at that spot. But recent work showed that once Cas9 is activated in this way, its RuvC cutting domain can also slice unrelated single-stranded DNA, such as poly(T) stretches, elsewhere in the solution. The authors set out to ask: which exact geometric and structural features of the target DNA and guide RNA make this collateral activity strong, weak, or absent?
Short versus long targets: giving Cas9 room to move
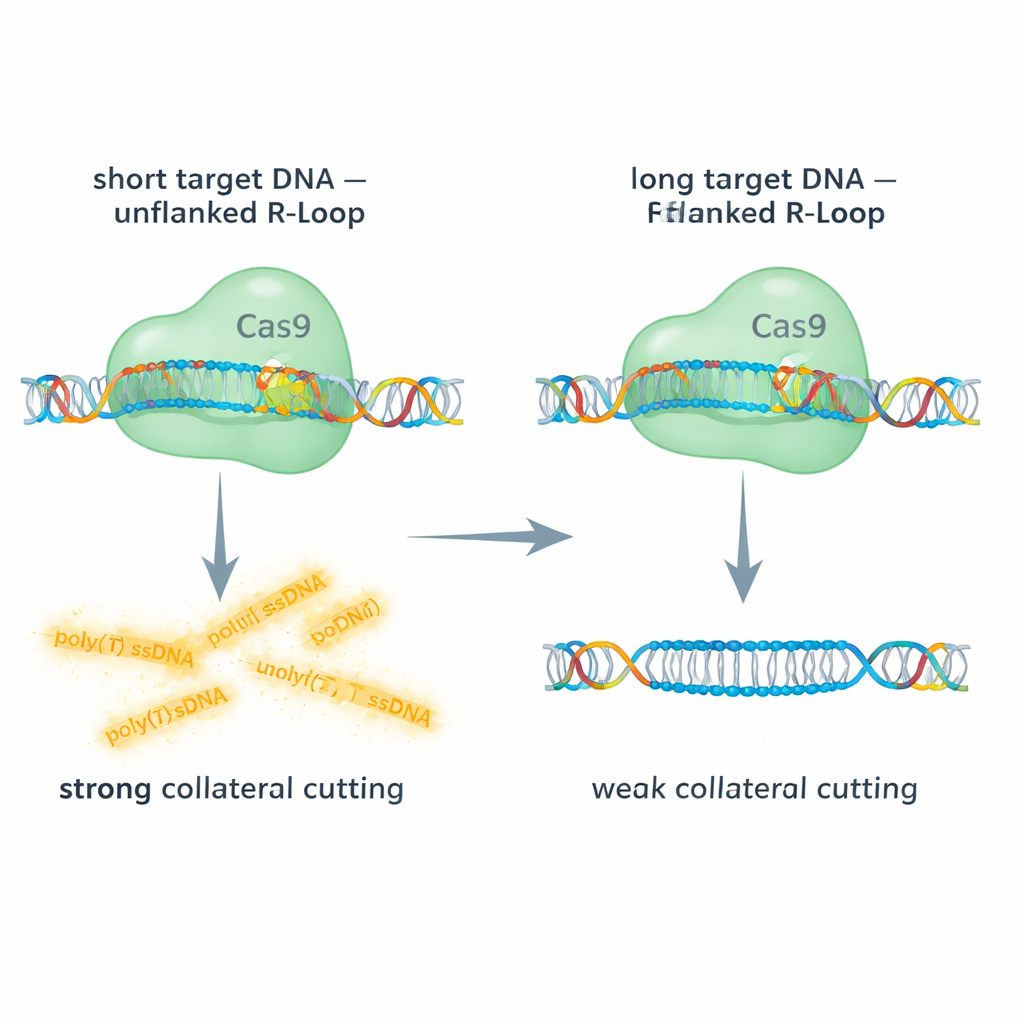
The team compared Cas9 acting on short and long double-stranded DNA targets, using fluorescent readouts to track both the on-target cut and collateral cutting of a single-stranded DNA probe. With short DNA targets, the R-loop at the guide RNA’s 5′ end is “unflanked” – there is no extra double-stranded DNA continuing beyond the hybrid region. Under these conditions, Cas9 showed robust collateral activity on single-stranded DNA. In contrast, when they used longer DNA segments that left extra double-stranded DNA flanking the R-loop, collateral cutting dropped dramatically, sometimes by about 90%, even though the main on-target cut still occurred. Targeting long single-stranded DNA, which removes the R-loop altogether, largely restored collateral activity. These comparisons reveal that a double-stranded “cap” sitting next to the R-loop stiffens the complex and physically hinders access or flexibility needed for Cas9’s RuvC domain to slice other strands.
Fine-tuning with guide length and mismatches
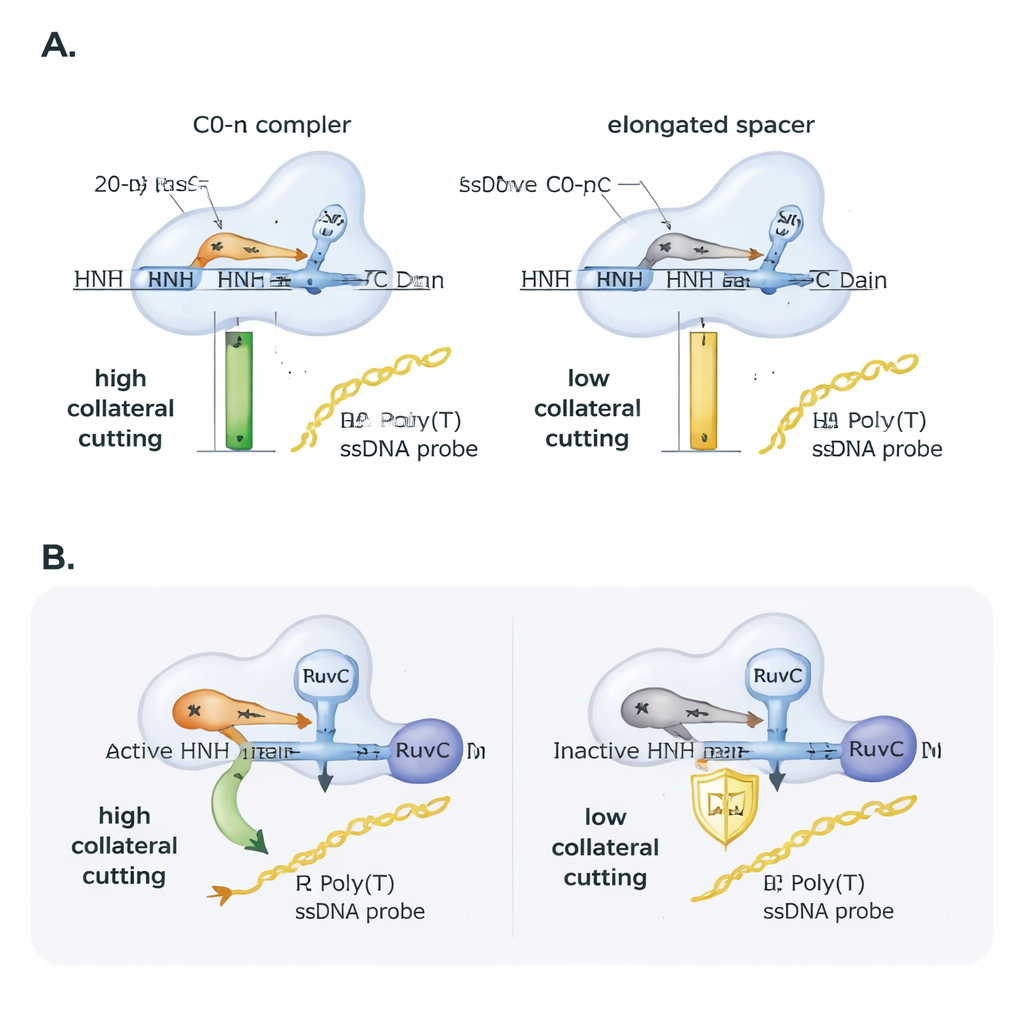
The researchers then explored how the guide RNA itself tunes this behavior. They introduced tiny mismatches between the guide and the target DNA and tracked how well Cas9 still cut. The main on-target cut tolerated many single-base mismatches, but collateral cutting was more fragile and strongly depended on exactly where the mismatch landed, underscoring its sensitivity. Next, they systematically lengthened the guide RNA spacer beyond the usual 20 building blocks. Even though Cas9 could still grab and cut the target DNA, collateral activity fell almost linearly as the spacer length increased: adding just two extra bases cut collateral activity roughly in half, and four extra bases reduced it even further. In practical tests using SARS-CoV-2 genetic material, only DNA amplicons arranged to produce an unflanked R-loop with a standard-length guide gave a strong collateral signal, highlighting how primer and guide design can make or break CRISPR-based detection assays.
An essential helper domain behind the scenes
Cas9 has two cutting domains, RuvC and HNH. Prior work linked collateral cutting directly to RuvC, but this study shows that HNH still matters. When the authors used a Cas9 variant with an inactive HNH domain, collateral activity plunged, even though binding to the target and nicking behavior remained. Interestingly, if they provided Cas9 with a DNA target that was already nicked in one strand, the inactive-HNH version regained collateral cutting similar to the normal enzyme. This suggests that HNH’s role is partly mechanical: by cutting or loosening the targeted strand, it helps the protein settle into a shape that exposes RuvC to nearby single-stranded DNA. Structural analyses of existing 3D models supported this view, showing that unflanked R-loops and standard-length guides let the RNA’s 5′ end “cap” against Cas9 and position the catalytic regions favorably, while flanked R-loops and elongated guides pack the protein more tightly and likely shield the RuvC site from stray strands.
What this means for future tools
For non-specialists, the main message is that Cas9’s behavior is not all-or-nothing: tiny geometric details—how far the DNA extends, how long the guide is, and whether a helper domain can complete its cut—decide whether the enzyme sticks to its main job or also shreds nearby single strands. Short targets that leave the R-loop unflanked, standard 20-base guides, and an active HNH domain jointly promote strong collateral cutting; long flanking DNA, stretched guides, or an inactive HNH domain suppress it. These insights give researchers a more precise dial to tune Cas9, helping them design safer gene-editing systems that avoid unwanted collateral damage, or more powerful diagnostic assays that deliberately harness this collateral activity to detect tiny amounts of viral or genetic material.
Citation: Montagud-Martínez, R., Ruiz, R., Baldanta, S. et al. CRISPR-Cas9 trans-cleavage is hindered by a flanked R-loop, an elongated spacer, and an inactive HNH domain. Nat Commun 17, 1998 (2026). https://doi.org/10.1038/s41467-026-68789-3
Keywords: CRISPR-Cas9, collateral cleavage, R-loop, guide RNA spacer, nucleic acid diagnostics